Abstract
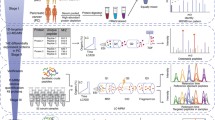
Nowadays, the ideal biomarker for colorectal cancer (CRC) has not been found. Two-dimensional electrophoresis (2-DE) and mass spectrometry (MS) are suitable techniques for searching new biomarkers. In this chapter, we describe methodology for biomarker discovery based on a proteomic approach. In addition, special attention is given to the sample preparation, including protein extraction, fractionation, and cleanup, as we consider this a critical step. Comparing the proteomic profile of tumor and mucosa, we identified the nucleoside diphosphate kinase A (NDKA) protein as a candidate biomarker for CRC. Finally, we validated NDKA with an ELISA kit using serum samples from individuals of a screening cohort. Our results suggest that serum NDKA is a potential biomarker for screening of CRC and premalignant advanced adenomas (AA).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Duffy MJ, Lamerz R, Haglund C et al (2014) Tumor markers in colorectal cancer, gastric cancer and gastrointestinal stromal cancers: European group on tumor markers 2014 guidelines update. Int J Cancer 134:2513–2522
Alvarez-Chaver P, Rodríguez-Piñeiro AM, Rodríguez-Berrocal FJ et al (2011) Selection of putative colorectal cancer markers by applying PCA on the soluble proteome of tumors: NDK A as a promising candidate. J Proteome 74:874–886
Görg A, Weiss W, Dunn MJ (2004) Current two-dimensional electrophoresis technology for proteomics. Proteomics 4:3665–3685
Otero-Estévez O, De Chiara L, Barcia-Castro L et al (2016) Evaluation of serum nucleoside diphosphate kinase A for the detection of colorectal cancer. Sci Rep 6:26703. https://doi.org/10.1038/srep26703
Alvarez-Chaver P, Otero-Estévez O, Páez de la Cadena M et al (2014) Proteomics for discovery of candidate colorectal cancer biomarkers. World J Gastroenterol 20:3804–3824
Bordier C (1981) Phase separation of integral membrane proteins in Triton X-114 solution. J Biol Chem 256:1604–1607
Santoni V, Kieffer S, Desclaux D et al (2000) Membrane proteomics: Use of additive main effects with multiplicative interaction model to classify plasma membrane proteins according to their solubility and electrophoretic properties. Electrophoresis 21:3329–3344
Heukeshoven J, Dernick R (1985) Simplified method for silver staining of proteins in polyacrylamide gels and the mechanism of silver staining. Electrophoresis 6:103–112
Shevchenko A, Wilm M, Vorm O et al (1996) Mass spectrometric sequencing of proteins silver-stained polyacrylamide gels. Anal Chem 68:850–858
Towbin H, Staehelin T, Gordon J (1979) Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A 76:4350–4354
Crowther JR (2009) The Elisa Guidebook. Series title: Methods in Molecular Biology, vol Vol. 515, 2nd edn. Humana Press, New York
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2018 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Álvarez-Chaver, P., De Chiara, L., Martínez-Zorzano, V.S. (2018). Proteomic Profiling for Colorectal Cancer Biomarker Discovery. In: Beaulieu, JF. (eds) Colorectal Cancer. Methods in Molecular Biology, vol 1765. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-7765-9_16
Download citation
DOI: https://doi.org/10.1007/978-1-4939-7765-9_16
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-7764-2
Online ISBN: 978-1-4939-7765-9
eBook Packages: Springer Protocols