Abstract
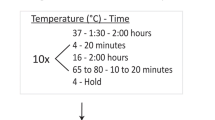
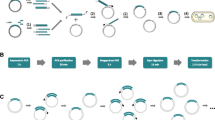
We developed a simple method (simple cloning) for subcloning DNA fragments into any location of a targeted vector without the need of restriction enzyme, ligase, exonuclease, or recombinase in Escherichia coli. This technology can be applied to common E. coli hosts (e.g., DH5α, JM109, TOP10, BL21(DE3)). The protocol includes three steps: (1) generate DNA insert and linear vector backbone by regular high-fidelity PCR, where these two DNA fragments contain 3′ and 5′ overlap** termini; (2) generate DNA multimers based on these two DNA fragments by using prolonged overlap extension-PCR (POE-PCR) without primers added; and (3) transform POE-PCR product to competent Escherichia coli cells directly, yielding the desired plasmid. Simple cloning provides a new cloning method with great simplicity and flexibility. Furthermore, this new method can be modified for the preparation of a large-size mutant library for directed evolution in E. coli. Using this method, it is very easy to generate a mutant library with a size of more than 107 per 50 μL of the POE-PCR product within 1 day.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Moxon ER, Higgins CF (1997) E. coli gene sequence - a blueprint for life. Nature 389:120–121
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, New York
Lund AH, Duch M, Pedersen FS (1996) Increased cloning efficiency by temperature-cycle ligation. Nucleic Acids Res 24:800–801
Young TS, Schultz PG (2010) Beyond the canonical 20 amino acids: expanding the genetic lexicon. J Biol Chem 285:11039–11044
Li CA, Evans RM (1997) Ligation independent cloning irrespective of restriction site compatibility. Nucleic Acids Res 25:4165–4166
Gillam EMJ, Copp JN, Ackerley DF et al (2014) Directed evolution library creation: methods and protocols, 2nd edn. Springer, New York
Abou-Nader M, Benedik MJ (2010) Rapid generation of random mutant libraries. Bioeng Bugs 1:337–340
Pai JC, Entzminger KC, Maynard JA (2012) Restriction enzyme-free construction of random gene mutagenesis libraries in Escherichia coli. Anal Biochem 421:640–648
You C, Zhang X-Z, Zhang YHP (2012) Simple cloning via direct transformation of PCR product (DNA Multimer) to Escherichia coli and Bacillus subtilis. Appl Environ Microbiol 78:1593–1595
Zhang X-Z, Zhang YHP (2011) Simple, fast and high-efficiency transformation system for directed evolution of cellulase in Bacillus subtilis. Microb Biotechnol 4:98–105
You C, Zhang YHP (2012) Easy preparation of a large-size random gene mutagenesis library in Escherichia coli. Anal Biochem 428:7–12
Shafikhani S, Siegel RA, Ferrari E et al (1997) Generation of large libraries of random mutants in Bacillus subtilis by PCR-based plasmid multimerization. Biotechniques 23:304–310
Acknowledgement
This work could not be finished without the support of the Nan**g Tech University and Virginia Tech. In addition, funding to YPZ for this work was provided in part by DOE EERE award (DE-EE0006968) and by the Virginia Agricultural Experiment Station and the Hatch Program of the National Institute of Food and Agriculture, US Department of Agriculture.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Science+Business Media New York
About this protocol
Cite this protocol
Zhong, C., You, C., Wei, P., Zhang, YH.P. (2017). Simple Cloning by Prolonged Overlap Extension-PCR with Application to the Preparation of Large-Size Random Gene Mutagenesis Library in Escherichia coli . In: Hughes, R. (eds) Synthetic DNA. Methods in Molecular Biology, vol 1472. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-6343-0_4
Download citation
DOI: https://doi.org/10.1007/978-1-4939-6343-0_4
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-6341-6
Online ISBN: 978-1-4939-6343-0
eBook Packages: Springer Protocols