Abstract
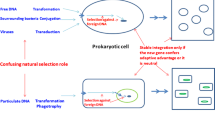
Horizontal gene transfer (HGT) between closely and distantly related lineages is a recurrent event in the Tree of Life (ToL) and provides genetic novelty that the recipient can utilize. Often, HGT is considered as an impediment in the reconstruction of life’s genealogy, and hence the concept of the ToL as a representation of evolutionary events is considered by some as erroneous. However, biased gene transfer, or the transfer between closely related lineages, can actually reinforce the familiar tree-like pattern by making organisms in close phylogenetic proximity appear even more closely related. The consequence of this bias is that the ToL remains tree-like when viewed from a distance, but looking closely, branches are fuzzy and boundaries between lineages may become indistinct as a result of recurrent HGT.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Hilario E, Gogarten JP (1993) Horizontal transfer of ATPase genes—the tree of life becomes a net of life. Biosystems 31(2–3):111–119
Huang J, Gogarten JP (2006) Ancient horizontal gene transfer can benefit phylogenetic reconstruction. Trends Genet 22(7):361–366
Huang J, Gogarten JP (2009) Ancient gene transfer as a tool in phylogenetic reconstruction. Methods Mol Biol 532:127–139
Lawrence JG, Retchless AC (2009) The interplay of homologous recombination and horizontal gene transfer in bacterial speciation. Methods Mol Biol 532:29–53
Fraser C, Alm EJ, Polz MF, Spratt BG, Hanage WP (2009) The bacterial species challenge: making sense of genetic and ecological diversity. Science 323(5915):741–746
Doolittle WF (1999) Phylogenetic classification and the universal tree. Science 284(5423):2124–2129
Doolittle WF, Bapteste E (2007) Pattern pluralism and the Tree of Life hypothesis. Proc Natl Acad Sci U S A 104(7):2043–2049
Bapteste E, O’Malley MA, Beiko RG et al (2009) Prokaryotic evolution and the tree of life are two different things. Biol Direct 4:34
Koonin EV, Wolf YI, Puigbò P (2009) The phylogenetic forest and the quest for the elusive tree of life. Cold Spring Harb Symp Quant Biol 74:205–213
Andam CP, Gogarten JP (2011) Biased gene transfer in microbial evolution. Nat Rev Microbiol 9(7):543–555
Lapierre P, Gogarten JP (2009) Estimating the size of the bacterial pan-genome. Trends Genet 25(3):107–110
Gogarten JP, Townsend JP (2005) Horizontal gene transfer, genome innovation and evolution. Nat Rev Microbiol 3(9):679–687
Abby SS, Tannier E, Gouy M, Daubin V (2012) Lateral gene transfer as a support for the tree of life. Proc Natl Acad Sci U S A 109(13):4962–4967
Christin P-A, Edwards EJ, Besnard G et al (2012) Adaptive evolution of C(4) photosynthesis through recurrent lateral gene transfer. Curr Biol 22(5):445–449
Acuña R, Padilla BE, Flórez-Ramos CP et al (2012) Adaptive horizontal transfer of a bacterial gene to an invasive insect pest of coffee. Proc Natl Acad Sci U S A 109(11):4197–4202
Danchin EGJ, Rosso M-N, Vieira P et al (2010) Multiple lateral gene transfers and duplications have promoted plant parasitism ability in nematodes. Proc Natl Acad Sci U S A 107(41):17651–17656
Hehemann J-H, Correc G, Barbeyron T, Helbert W, Czjzek M, Michel G (2010) Transfer of carbohydrate-active enzymes from marine bacteria to Japanese gut microbiota. Nature 464(7290):908–912
Thomas F, Barbeyron T, Tonon T, Génicot S, Czjzek M, Michel G (2012) Characterization of the first alginolytic operons in a marine bacterium: from their emergence in marine Flavobacteriia to their independent transfers to marine Proteobacteria and human gut Bacteroides. Environ Microbiol 14(9):2379–2394
Lurie-Weinberger MN, Peeri M, Gophna U (2012) Contribution of lateral gene transfer to the gene repertoire of a gut-adapted methanogen. Genomics 99(1):52–58
Raz Y, Tannenbaum E (2010) The influence of horizontal gene transfer on the mean fitness of unicellular populations in static environments. Genetics 185(1):327–337
Swithers KS, Soucy SM, Gogarten JP (2012) The role of reticulate evolution in creating innovation and complexity. Int J Evol Biol 2012:1–10
Schjørring S, Krogfelt KA (2011) Assessment of bacterial antibiotic resistance transfer in the gut. Int J Microbiol 2011:312956
Nelson KE, Clayton RA, Gill SR et al (1999) Evidence for lateral gene transfer between archaea and bacteria from genome sequence of Thermotoga maritima. Nature 399(6734):323–329
Zhaxybayeva O, Swithers K, Lapierre P et al (2009) On the chimeric nature, thermophilic origin, and phylogenetic placement of the Thermotogales. Proc Natl Acad Sci U S A 106:5865–5870
Boussau B, Gueguen L, Gouy M (2008) Accounting for horizontal gene transfers explains conflicting hypotheses regarding the position of aquificales in the phylogeny of Bacteria. BMC Evol Biol 8:272
Le Fourn C, Brasseur G, Brochier-Armanet C et al (2011) An oxygen reduction chain in the hyperthermophilic anaerobe Thermotoga maritima highlights horizontal gene transfer between Thermococcales and Thermotogales. Environ Microbiol 13(8):2132–2145
Beiko RG, Harlow TJ, Ragan MA (2005 Oct 4) Highways of gene sharing in prokaryotes. Proc Natl Acad Sci U S A 102(40):14332–7
Smillie CS, Smith MB, Friedman J, Cordero OX, David LA, Alm EJ (2011) Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480(7376):241–244
McNulty SN, Foster JM, Mitreva M et al (2010) Endosymbiont DNA in endobacteria-free filarial nematodes indicates ancient horizontal genetic transfer. PLoS ONE 5(6):e11029
Madsen JS, Burmølle M, Hansen LH, Sørensen SJ (2012) The interconnection between biofilm formation and horizontal gene transfer. FEMS Immunol Med Microbiol 65(2):183–195
Andam CP, Williams D, Gogarten JP (2010) Biased gene transfer mimics patterns created through shared ancestry. Proc Natl Acad Sci U S A 107(23):10679–10684
Tamminen M, Virta M, Fani R, Fondi M (2012) Large-scale analysis of plasmid relationships through gene-sharing networks. Mol Biol Evol 29(4):1225–1240
Zhaxybayeva O, Gogarten JP, Charlebois RL, Doolittle WF, Papke RT (2006) Phylogenetic analyses of cyanobacterial genomes: quantification of horizontal gene transfer events. Genome Res 16(9):1099–1108
Eppley JM, Tyson GW, Getz WM, Banfield JF (2007) Genetic exchange across a species boundary in the archaeal genus ferroplasma. Genetics 177(1):407–416
Popa O, Hazkani-Covo E, Landan G, Martin W, Dagan T (2011) Directed networks reveal genomic barriers and DNA repair bypasses to lateral gene transfer among prokaryotes. Genome Res 21(4):599–609
Gogarten JP, Doolittle WF, Lawrence JG (2002) Prokaryotic evolution in light of gene transfer. Mol Biol Evol 19(12):2226–2238
Puigbò P, Wolf YI, Koonin EV (2010) The tree and net components of prokaryote evolution. Genome Biol Evol 2:745–756
Andam CP, Williams D, Gogarten JP (2010) Natural taxonomy in light of horizontal gene transfer. Biol Philos 25(4):589–602
Andam CP, Gogarten JP (2011) Biased gene transfer and its implications for the concept of lineage. Biol Direct 6:47
Omelchenko MV, Galperin MY, Wolf YI, Koonin EV (2010) Non-homologous isofunctional enzymes: a systematic analysis of alternative solutions in enzyme evolution. Biol Direct 5:31
Zhang R-G, Andersson CE, Savchenko A et al (2003) Structure of Escherichia coli ribose-5-phosphate isomerase: a ubiquitous enzyme of the pentose phosphate pathway and the Calvin cycle. Structure 11(1):31–42
Zhang R-G, Andersson CE, Skarina T et al (2003) The 2.2 A resolution structure of RpiB/AlsB from Escherichia coli illustrates a new approach to the ribose-5-phosphate isomerase reaction. J Mol Biol 332(5):1083–1094
Roos AK, Andersson CE, Bergfors T et al (2004) Mycobacterium tuberculosis ribose-5-phosphate isomerase has a known fold, but a novel active site. J Mol Biol 335(3):799–809
Marchler-Bauer A, Lu S, Anderson JB et al (2011) CDD: a Conserved Domain Database for the functional annotation of proteins. Nucleic Acids Res 39:D225–D229 (Database issue)
Roos AK, Mariano S, Kowalinski E, Salmon L, Mowbray SL (2008) D-ribose-5-phosphate isomerase B from Escherichia coli is also a functional D-allose-6-phosphate isomerase, while the Mycobacterium tuberculosis enzyme is not. J Mol Biol 382(3):667–679
Stern A, Mayrose I, Penn O, Shaul S, Gophna U, Pupko T (2010) An evolutionary analysis of lateral gene transfer in thymidylate synthase enzymes. Syst Biol 59(2):212–225
Grassi L, Caselle M, Lercher MJ, Lagomarsino MC (2012) Horizontal gene transfers as metagenomic gene duplications. Mol Biosyst 8(3):790–795
Karberg KA, Olsen GJ, Davis JJ (2011) Similarity of genes horizontally acquired by Escherichia coli and Salmonella enterica is evidence of a supraspecies pangenome. Proc Natl Acad Sci U S A 108(50):20154–20159
Skip**ton E, Ragan MA (2011) Lateral genetic transfer and the construction of genetic exchange communities. FEMS Microbiol Rev 35(5):707–735
Beauregard-Racine J, Bicep C, Schliep K, Lopez P, Lapointe F-J, Bapteste E (2011) Of woods and webs: possible alternatives to the tree of life for studying genomic fluidity in E. coli. Biol Direct 6:39
Sharon I, Battchikova N, Aro E-M et al (2011) Comparative metagenomics of microbial traits within oceanic viral communities. ISME J 5(7):1178–1190
Lang AS, Zhaxybayeva O, Beatty JT (2012) Gene transfer agents: phage-like elements of genetic exchange. Nat Rev Microbiol 10(7):472–482
Ronning C, Losada L, Brinkac L et al (2010) Genetic and phenotypic diversity in Burkholderia: contributions by prophage and phage-like elements. BMC Microbiol 10(1):202
Naor A, Lapierre P, Mevarech M, Papke RT, Gophna U (2012) Low species barriers in halophilic archaea and the formation of recombinant hybrids. Curr Biol 22(15):1444–1448
Frost LS, Leplae R, Summers AO, Toussaint A (2005) Mobile genetic elements: the agents of open source evolution. Nat Rev Microbiol 3(9):722–732
Bapteste E, Bouchard F, Burian RM (2012) Philosophy and evolution: minding the gap between evolutionary patterns and tree-like patterns. Methods Mol Biol 856:81–110
Lukjancenko O, Wassenaar TM, Ussery DW (2010) Comparison of 61 sequenced Escherichia coli genomes. Microb Ecol 60(4):708–720
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32(5):1792–1797
Guindon S, Gascuel O (2003) A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52(5):696–704
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25(7):1307–1320
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media New York
About this chapter
Cite this chapter
Andam, C., Gogarten, J.P. (2013). Biased Gene Transfer Contributes to Maintaining the Tree of Life. In: Gophna, U. (eds) Lateral Gene Transfer in Evolution. Springer, New York, NY. https://doi.org/10.1007/978-1-4614-7780-8_14
Download citation
DOI: https://doi.org/10.1007/978-1-4614-7780-8_14
Published:
Publisher Name: Springer, New York, NY
Print ISBN: 978-1-4614-7779-2
Online ISBN: 978-1-4614-7780-8
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)