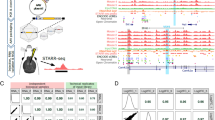
Abstract
Recombinant adeno-associated viruses can be coupled with short regulatory elements to restrict viral expression to specific cellular populations. These viral vectors can be used as tools for basic research to dissect many aspects of the biology of specific cellular subtypes in health and disease, and across species. A handful of enhancers have already been described in the nervous system, and recent studies suggest that transcriptomic and epigenetic data can be leveraged to systematize the discovery of novel elements to restrict viral expression to any cell type. However, a thorough characterization of the expression profile conferred by these short sequences is required to demonstrate their utility in the experimental context in which they will be ultimately used. Here we describe a complete guide to select, screen, and validate the expression profile of enhancers to target specific subtypes of neurons.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dimidschstein J et al (2016) A viral strategy for targeting and manipulating interneurons across vertebrate species. Nat Neurosci 19:1743–1749
Vormstein-Schneider D, Lin JD et al (2020) Viral manipulation of functionally distinct interneurons in mice, non-human primates and humans. Nat Neurosci 23:1629–1636
Bedbrook CN, Deverman BE, Gradinaru V (2018) Viral strategies for targeting the central and peripheral nervous systems. Annu Rev Neurosci 41:323–348
Hrvatin S et al (2019) A scalable platform for the development of cell-type-specific viral drivers. elife 8:e48089
Mich JK et al (2021) Functional enhancer elements drive subclass-selective expression from mouse to primate neocortex. Cell Rep 34(13):108754
Mingozzi F et al (2003) Induction of immune tolerance to coagulation factor IX antigen by in vivo hepatic gene transfer. J Clin Invest 11(9):1347–1356
**ong W et al (2019) AAV cis-regulatory sequences are correlated with ocular toxicity. Proc Natl Acad Sci U S A 116(12):5785–5794
Vesuna S et al (2020) Deep posteromedial cortical rhythm in dissociation. Nature 586:87–94
Mo A et al (2015) Epigenomic signatures of neuronal diversity in the mammalian brain. Neuron 86:1369–1384
Gray SJ et al (2011) Production of recombinant adeno-associated viral vectors and use in in vitro and in vivo administration. Curr Protoc Neurosci 4:4.17
Fulco CP et al (2016) Systematic map** of functional enhancer-promoter connections with CRISPR interference. Science 354(6313):769–773
Visel A et al (2007) VISTA enhancer browser–a database of tissue-specific human enhancers. Nucleic Acids Res 35:D88–D92
Bejerano G et al (2004) Ultraconserved elements in the human genome. Science 304:1321–1325
Dimitrieva S, Bucher P (2013) UCNEbase—a database of ultraconserved non-coding elements and genomic regulatory blocks. Nucleic Acids Res 41:D101–D109
Andersson R et al (2014) An atlas of active enhancers across human cell types and tissues. Nature 507:455–461
Dousse A, Junier T, Zdobnov EM (2016) CEGA—a catalog of conserved elements from genomic alignments. Nucleic Acids Res 44:D96–D100
Dickel DE et al (2018) Ultraconserved enhancers are required for normal development. Cell 172:491–499
Ron G et al (2017) Promoter-enhancer interactions identified from Hi-C data using probabilistic models and hierarchical topological domains. Nat Commun 8:2237
Huang J et al (2018) Dissecting super-enhancer hierarchy based on chromatin interactions. Nat Commun 9(1):943
Tasic B et al (2016) Adult mouse cortical cell taxonomy revealed by single cell transcriptomics. Nat Neurosci 19:335–346
Saunders A et al (2018) Molecular diversity and specializations among the cells of the adult mouse brain. Cell 174:1015–1030.e16
Tremblay R, Lee S, Rudy B (2016) GABAergic interneurons in the neocortex: from cellular properties to circuits. Neuron 91:260–292
Siepel A et al (2005) Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res 15(8):1034–1050
Kent WJ et al (2002) The human genome browser at UCSC. Genome Res 12(6):996–1006
Buenrostro J et al (2013) Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat Methods 10:1213–1218
Barski A et al (2007) High-resolution profiling of histone methylations in the human genome. Cell 129:823–837
Johnson DS et al (2007) Genome-wide map** of in vivo protein-DNA interactions. Science 316:1497–1502
Mikkelsen T et al (2007) Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448:553–560
Spitz F, Furlong E (2012) Transcription factors: from enhancer binding to developmental control. Nat Rev Genet 13:613–626
Tuteja G et al (2014) Automated discovery of tissue-targeting enhancers and transcription factors from binding motif and gene function data. PLoS Comput Biol 10(1):e1003449
He HH et al (2010) Nucleosome dynamics define transcriptional enhancers. Nat Genet 42(4):343–347
Fang R et al (2021) Comprehensive analysis of single cell ATAC-seq data with SnapATAC. Nat Commun 12:1337
Feng J et al (2012) Identifying ChIP-seq enrichment using MACS. Nat Protoc 7(9):1728–1740
Freese NH et al (2016) Integrated genome browser: visual analytics platform for genomics. Bioinformatics 32(14):2089–2095
Jüttner J et al (2019) Targeting neuronal and glial cell types with synthetic promoter AAVs in mice, non-human primates and humans. Nat Neurosci 22:1345–1356
Blankvoort S, Witter MP, Noonan J, Cotney J, Kentros C (2018) Marked diversity of unique cortical enhancers enables neuron-specific tools by enhancer-driven gene expression. Curr Biol 28(13):2103–2114
Wu Z, Yang H, Colosi P (2010) Effect of genome size on AAV vector packaging. Mol Ther 18(1):80–86
Powell SK, Rivera-Soto R, Gray SJ (2015) Viral expression cassette elements to enhance transgene target specificity and expression in gene therapy. Discov Med 19(102):49–57
Chalfie M et al (1994) Green fluorescent protein as a marker for gene-expression. Science 263:802–805
Heim R, Tsien RY (1996) Engineering green fluorescent protein for improved brightness, longer wavelengths and fluorescence resonance energy transfer. Curr Biol 6:178–182
Matz MV et al (1999) Fluorescent proteins from nonbioluminescent Anthozoa species. Nat Biotechnol 17:969–973
Shaner NC et al (2004) Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat Biotechnol 22:1567–1572
Strack RL et al (2009) A rapidly maturing far-red derivative of DsRed-Express2 for whole-cell labeling. Biochemistry 48:8279–8281
Griesbeck O et al (2001) Reducing the environmental sensitivity of yellow fluorescent protein. J Biol Chem 276:29188–29194
Kalderon D, Roberts BL, Richardson WD, Smith AE (1984) A short amino acid sequence able to specify nuclear location. Cell 39:499–509
Robbins J, Dilworth SM, Laskey RA, Dingwall C (1991) Two interdependent basic domains in nucleoplasmin nuclear targeting sequence: identification of a class of bipartite nuclear targeting sequence. Cell 64:615–623
Fowler DK et al (2016) A MultiSite gateway toolkit for rapid cloning of vertebrate expression constructs with diverse research applications. PloS one 11(8):e0159277
Chen TW et al (2013) Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499(7458):295–300
Piatkevich KD et al (2018) A robotic multidimensional directed evolution approach applied to fluorescent voltage reporters. Nat Chem Biol 14(4):352–360
Marvin JS et al (2019) A genetically encoded fluorescent sensor for in vivo imaging of GABA. Nat Methods 16:763–770
Magnus CJ et al (2019) Ultrapotent chemogenetics for research and potential clinical applications. Science 364:6436
Boyden ES et al (2005) Millisecond-timescale, genetically targeted optical control of neural activity. Nat Neurosci 8:1263–1268
Yizhar O et al (2011) Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature 477(7363):171–178
Graybuck LT et al (2021) Enhancer viruses for combinatorial cell-subclass-specific labeling. Neuron 109(9):1449–1464.e13
Chen Q et al (2020) Dysfunction of cortical GABAergic neurons leads to sensory hyper-reactivity in a Shank3 mouse model of ASD. Nat Neurosci 23:520–532
Watakabe A et al (2015) Comparative analyses of adeno-associated viral vector serotypes 1, 2, 5, 8 and 9 in marmoset, mouse and macaque cerebral cortex. Neurosci Res 93:144–157
Chan KY et al (2017) Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat Neurosci 20:1172–1179
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 This is a U.S. government work and not under copyright protection in the U.S.; foreign copyright protection may apply
About this protocol
Cite this protocol
Lin, J., Dimidschstein, J. (2023). Enhancers for Selective Targeting. In: Eldridge, M.A., Galvan, A. (eds) Vectorology for Optogenetics and Chemogenetics. Neuromethods, vol 195. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2918-5_9
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2918-5_9
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2917-8
Online ISBN: 978-1-0716-2918-5
eBook Packages: Springer Protocols