Abstract
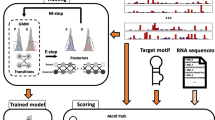
RNA transcripts can form a variety of higher-order structures. We developed a large-scale affinity analysis system, FOREST (Folded RNA Element Profiling with Structure Library), to investigate the function of these RNA structures on transcriptome-wide scale. Here we describe a protocol to analyze RNA–protein interactions using FOREST . Users of the protocol prepare an RNA structure library comprised of diverse species of transcripts and perform high-throughput characterization of the RNA–protein interactions to obtain quantitative and comprehensive information on the binding affinities and specificities. Moreover, we demonstrate how FOREST can be used to analyze a non-canonical structure, the RNA G-quadruplex, without sequencing bias, because the quantification is performed directly on a microarray without sequence amplification. FOREST will contribute to the discovery of RNA structure motifs that determine RNA–protein interactions.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Yao Z, Weinberg Z, Ruzzo WL (2006) CMfinder—a covariance model based RNA motif finding algorithm. Bioinformatics 22:445–452
Nawrocki EP, Eddy SR (2013) Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29:2933–2935
Siegfried N, Busan S, Rice G et al (2014) RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat Methods 11:959–965
Lu Z et al (2016) RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165:1267–1279
Mustoe AM et al (2018) Pervasive regulatory functions of mRNA structure revealed by high-resolution SHAPE probing. Cell 173:181–195
Zhang Z, **ng Y (2017) CLIP-seq analysis of multi-mapped reads discovers novel functional RNA regulatory sites in the human transcriptome. Nucleic Acids Res 45:9260–9271
Hafner M, Katsantoni M, Köster T et al (2021) CLIP and complementary methods. Nat Rev Methods Primers 1:20
Dominguez D et al (2018) Sequence, structure, and context preferences of human RNA binding proteins. Mol Cell 70:854–867
Komatsu KR, Taya T, Matsumoto S et al (2020) RNA structure-wide discovery of functional interactions with multiplexed RNA motif library. Nat Commun 11:6275
Ray D et al (2009) Rapid and systematic analysis of the RNA recognition specificities of RNA-binding proteins. Nat Biotechnol 27:667–670
Triboulet R, Pirouz M, Gregory RI (2015) A single Let-7 microRNA bypasses LIN28-mediated repression. Cell Rep 13:260–266
Ustianenko D et al (2018) LIN28 selectively modulates a subclass of Let-7 microRNAs. Mol Cell 71:271–283
Huang Z-L et al (2018) Identification of G-quadruplex-binding protein from the exploration of RGG motif/G-quadruplex interactions. J Am Chem Soc 140:17945–17955
Tippana R, Chen MC, Demeshkina NA, Ferré-D’Amaré AR, Myong S (2019) RNA G-quadruplex is resolved by repetitive and ATP-dependent mechanism of DHX36. Nat Commun 10:1855
Acknowledgments
We would like to thank P. Karagiannis for critical reading of the paper. This work was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI No. 20H05626, 15H05722, to H.S.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Miyashita, E., Komatsu, K.R., Saito, H. (2022). Large-Scale Analysis of RNA–Protein Interactions for Functional RNA Motif Discovery Using FOREST. In: Parrish, N.F., Iwasaki, Y.W. (eds) piRNA. Methods in Molecular Biology, vol 2509. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2380-0_17
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2380-0_17
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2379-4
Online ISBN: 978-1-0716-2380-0
eBook Packages: Springer Protocols