Search
Search Results
-
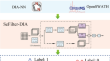
SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics
Mass spectrometry is crucial in proteomics analysis, particularly using Data Independent Acquisition (DIA) for reliable and reproducible mass...
-
Data-Independent Acquisition Peptidomics
In recent years, data-independent acquisition (DIA) has emerged as a powerful analysis method in biological mass spectrometry (MS). Compared to the...
-
Ultra-fast label-free quantification and comprehensive proteome coverage with narrow-window data-independent acquisition
Mass spectrometry (MS)-based proteomics aims to characterize comprehensive proteomes in a fast and reproducible manner. Here we present the...

-
Quantitative proteomics analysis based on data-independent acquisition reveals the effect of Shenling Baizhu powder (SLP) on protein expression in MAFLD rat liver tissue
BackgroundMetabolic associated fatty liver disease (MAFLD) has become the most common chronic liver disease worldwide, and it is also a high-risk...

-
MaxDIA enables library-based and library-free data-independent acquisition proteomics
MaxDIA is a software platform for analyzing data-independent acquisition (DIA) proteomics data within the MaxQuant software environment. Using...

-
Data-independent acquisition boosts quantitative metaproteomics for deep characterization of gut microbiota
Metaproteomics can provide valuable insights into the functions of human gut microbiota (GM), but is challenging due to the extreme complexity and...

-
Data-independent acquisition-based mass spectrometry(DIA-MS) for quantitative analysis of patients with chronic hepatitis B
Chronic hepatitis B is a significant public health problem and complex pathologic process, and unraveling the underlying mechanisms and...

-
An MSstats workflow for detecting differentially abundant proteins in large-scale data-independent acquisition mass spectrometry experiments with FragPipe processing
Technological advances in mass spectrometry and proteomics have made it possible to perform larger-scale and more-complex experiments. The volume and...

-
Quantitative proteomics analysis of human vitreous in rhegmatogenous retinal detachment associated with choroidal detachment by data-independent acquisition mass spectrometry
The prognosis of rhegmatogenous retinal detachment (RRD) with choroidal detachment (RRDCD) is often poor and complicated. This study focused on the...

-
Optimized data-independent acquisition approach for proteomic analysis at single-cell level
BackgroundSingle-cell proteomic analysis provides valuable insights into cellular heterogeneity allowing the characterization of the cellular...

-
Introducing untargeted data-independent acquisition for metaproteomics of complex microbial samples
Mass spectrometry-based metaproteomics is a relatively new field of research that enables the characterization of the functionality of microbiota....

-
Deep representation features from DreamDIAXMBD improve the analysis of data-independent acquisition proteomics
We developed DreamDIA XMBD (denoted as DreamDIA), a software suite based on a deep representation model for data-independent acquisition (DIA) data...

-
Prioritize biologically relevant ions for data-independent acquisition (BRI-DIA) in LC–MS/MS-based lipidomics analysis
IntroductionData-dependent acquisition (DDA) is the most commonly used MS/MS scan method for lipidomics analysis on orbitrap-based instrument....

-
quantms: a cloud-based pipeline for quantitative proteomics enables the reanalysis of public proteomics data
The volume of public proteomics data is rapidly increasing, causing a computational challenge for large-scale reanalysis. Here, we introduce quantms (

-
Proteomics for Tea Plant
Plant proteome is highly dynamic and is essential for the investigation of protein expression, interactions, and modifications to better understand...
-
Label-Free Proteomics of Quantity-Limited Samples Using Ion Mobility-Assisted Data-Independent Acquisition Mass Spectrometry
Over the past two decades, unbiased data-independent acquisition (DIA) approachesProteomicsapproaches have gained increasing popularity in the...
-
Novel FFPE proteomics method suggests prolactin induced protein as hormone induced cytoskeleton remodeling spatial biomarker
Robotically assisted proteomics provides insights into the regulation of multiple proteins achieving excellent spatial resolution. However,...

-
Proteomic signature associated with chronic kidney disease (CKD) progression identified by data-independent acquisition mass spectrometry
BackgroundHalting progression of chronic kidney disease (CKD) to established end stage kidney disease is a major goal of global health research. The...

-
Generalized precursor prediction boosts identification rates and accuracy in mass spectrometry based proteomics
Data independent acquisition mass spectrometry (DIA-MS) has recently emerged as an important method for the identification of blood-based biomarkers....

