Search
Search Results
-
TFscope: systematic analysis of the sequence features involved in the binding preferences of transcription factors
Characterizing the binding preferences of transcription factors (TFs) in different cell types and conditions is key to understand how they...

-

-

-
Circ-IP6K2 suppresses tumor progression by modulating the miR-1292-5p/CAMK2N1 signal in clear cell renal cell carcinoma
Renal cell carcinoma (RCC) is a malignant tumor originating from the epithelial cells of the renal tubules. The clear cell RCC subtype is closely...

-
gcplyr: an R package for microbial growth curve data analysis
BackgroundCharacterization of microbial growth is of both fundamental and applied interest. Modern platforms can automate collection of...

-
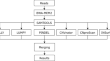
ProcaryaSV: structural variation detection pipeline for bacterial genomes using short-read sequencing
BackgroundStructural variations play an important role in bacterial genomes. They can mediate genome adaptation quickly in response to the external...
-
Purifying selection drove the adaptation of mitochondrial genes along with correlation of gene rearrangements and evolutionary rates in two subfamilies of Whitefly (Insecta: Hemiptera)
Insect mitochondrial genomes (mitogenomes) are usually represented by a conserved gene order. Whiteflies exhibit gene rearrangement in their...

-
Massively integrated coexpression analysis reveals transcriptional regulation, evolution and cellular implications of the yeast noncanonical translatome
BackgroundRecent studies uncovered pervasive transcription and translation of thousands of noncanonical open reading frames (nORFs) outside of...

-

-
Complete substitution with modified nucleotides in self-amplifying RNA suppresses the interferon response and increases potency
The use of modified nucleotides to suppress the interferon response and maintain translation of self-amplifying RNA (saRNA), which has been achieved...

-
The contribution of silencer variants to human diseases
BackgroundAlthough disease-causal genetic variants have been found within silencer sequences, we still lack a comprehensive analysis of the...

-
Panpipes: a pipeline for multiomic single-cell and spatial transcriptomic data analysis
Single-cell multiomic analysis of the epigenome, transcriptome, and proteome allows for comprehensive characterization of the molecular circuitry...

-
Computational gastronomy: capturing culinary creativity by making food computable
Cooking, a quintessential creative pursuit, holds profound significance for individuals, communities, and civilizations. Food and cooking transcend...

-
Detection of allele-specific expression in spatial transcriptomics with spASE
Spatial transcriptomics technologies permit the study of the spatial distribution of RNA at near-single-cell resolution genome-wide. However, the...

-
Ensemble Machine Learning and Predicted Properties Promote Antimicrobial Peptide Identification
AbstractThe emergence of antibiotic-resistant microbes raises a pressing demand for novel alternative treatments. One promising alternative is the...

-
TaqTth-hpRNA: a novel compact RNA-targeting tool for specific silencing of pathogenic mRNA
Pathogenic allele silencing is a promising treatment for genetic hereditary diseases. Here, we develop an RNA-cleaving tool, TaqTth-hpRNA, consisting...

-
Comparative Functional Characterization of nst1, nst2, and nst3 in Arabidopsis thaliana Uncovers Previously Unknown Functions in Diverse Developmental Pathways Beyond Secondary Wall Formation
The regulation of secondary cell wall formation in Arabidopsis thaliana has been extensively studied with NST1 , NST2 , and NST3 playing key roles in...

