Abstract
Drosophila melanogaster offers a powerful model system for interrogating interactions between host cells and human bacterial pathogens. Brucella, a gram-negative, facultative intracellular bacterium is the causative agent of brucellosis, a zoonotic disease of global consequence. Over the past several decades, pathogen factors that mediate Brucella infection have been identified. However, host factors that mediate infection have remained obscure. We have used the power of the Drosophila S2 cell system to identify and characterize host factors that support infection by Brucella melitensis. Host protein inositol-requiring enzyme 1 (IRE1α), a transmembrane kinase and master regulator of the eukaryotic unfolded protein response, was shown to play an important role in regulating Brucella infection, thereby providing the first glimpse of host mechanisms that are subverted by the pathogen to support its intracellular lifestyle. Furthermore, our study also established the Drosophila S2 cell as a powerful system for elucidating Brucella host factors. Here, we describe a protocol for using the Drosophila S2 cell system for studying the Brucella–host interaction.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Dorer MS, Kirton D, Bader JS, Isberg RR (2006) RNA interference analysis of Legionella in Drosophila cells: Exploitation of early secretory apparatus dynamics. PLoS Pathog 2:315–327
Ramet M, Manfruelli P, Pearson A, Mathey-Prevot B, Ezekowitz RAB (2002) Functional genomic analysis of phagocytosis and identification of a Drosophila receptor for E-coli. Nature 416:644–648
Agaisse H, Burrack LS, Philips JA, Rubin EJ, Perrimon N, Higgins DE (2005) Genome-wide RNAi screen for host factors required for intracellular bacterial infection. Science 309:1248–1251
Pielage JF, Powell KR, Kalman D, Engel JN (2008) RNAi screen reveals an AbI kinase-dependent host cell pathway involved in Pseudomonas aeruginosa internalization. PLoS Pathog 4(3):e1000031. doi:10.1371/journal.ppat.1000031
Cherry S (2008) Genomic RNAi screening in Drosophila S2 cells: what have we learned about host-pathogen interactions? Curr Opin Microbiol 11:262–270
Cheng LW, Viala JPM, Stuurman N, Wiedemann U, Vale RD, Portnoy DA (2005) Use of RNA interference in Drosophila S2 cells to identify host pathways controlling compartmentalization of an intracellular pathogen. Proc Natl Acad Sci U S A 102:13646–13651
Philips JA, Rubin EJ, Perrimon N (2005) Drosophila RNAi screen reveals CD36 family member required for mycobacterial infection. Science 309:1251–1253
Derre I, Pypaert M, Dautry-Varsat A, Agaisse H (2007) RNAi screen in Drosophila cells reveals the involvement of the tom complex in Chlamydia infection. PLoS Pathog 3:1446–1458
Elwell CA, Ceesay A, Kim JH, Kalman D, Engel JN (2008) RNA interference screen identifies AbI kinase and PDGFR signaling in Chlamydia trachomatis entry. PLoS Pathog. doi:10.1371/journal.ppat.1000021
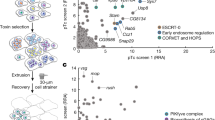
Qin QM, Pei J, Ancona V, Shaw BD, Ficht TA, de Figueiredo P (2008) RNAi screen of endoplasmic reticulum-associated host factors reveals a role for IRE1 alpha in supporting Brucella replication. PLoS Pathog 4
Gibbs EPJ (2005) Emerging zoonotic epidemics in the interconnected global community. Vet Rec 157:673–679
Sarinas PS, Chitkara RK (2003) Brucellosis. Semin Respir Infect 18:168–182
Godfroid J, Cloeckaert A, Liautard JP, Kohler S, Fretin D, Walravens K, Garin-Bastuji B, Letesson JJ (2005) From the discovery of the Malta fever’s agent to the discovery of a marine mammal reservoir, brucellosis has continuously been a re-emerging zoonosis. Vet Res 36:313–326
Sauret JM, Vilissova N (2002) Human brucellosis. J Am Board Fam Pract 15:401–406
Ariza J, Bosilkovski M, Cascio A, Colmenero JD, Corbel MJ, Falagas ME, Memish ZA, Roushan MRH, Rubinstein E, Sipsas NV, Solera J, Young EJ, Pappas G (2007) Perspectives for the treatment of brucellosis in the 21st century: the Ioannina recommendations. PLoS Med 4:1872–1878
Schurig GG, Sriranganathan N, Corbel MJ (2002) Brucellosis vaccines: past, present and future. Vet Microbiol 90:479–496
Rittig MG, Kaufmann A, Robins A, Shaw B, Sprenger H, Gemsa D, Foulongne V, Rouot B, Dornand J (2003) Smooth and rough lipopolysaccharide phenotypes of Brucella induce different intracellular trafficking and cytokine/chemokine release in human monocytes. J Leukoc Biol 74:1045–1055
Monreal D, Grillo MJ, Gonzalez D, Marin CM, De Miguel MJ, Lopez-Goni I, Blasco JM, Cloeckaert A, Moriyon I (2003) Characterization of Brucella abortus O-polysaccharide and core lipopolysaccharide mutants and demonstration that a complete core is required for rough vaccines to be efficient against Brucella abortus and Brucella ovis in the mouse model. Infect Immun 71:3261–3271
O’Callaghan D, Cazevieille C, Allardet-Servent A, Boschiroli ML, Bourg G, Foulongne V, Frutos P, Kulakov Y, Ramuz M (1999) A homologue of the Agrobacterium tumefaciens VirB and Bordetella pertussis Ptl type IV secretion systems is essential for intracellular survival of Brucella suis. Mol Microbiol 33:1210–1220
Celli J, de Chastellier C, Franchini DM, Pizarro-Cerda J, Moreno E, Gorvel AP (2003) Brucella evades macrophage killing via VirB-dependent sustained interactions with the endoplasmic reticulum. J Exp Med 198:545–556
Pei JW, Turse JE, Wu QM, Ficht TA (2006) Brucella abortus rough mutants induce macrophage oncosis that requires bacterial protein synthesis and direct interaction with the macrophage. Infect Immun 74:2667–2675
Guzman-Verri C, Chaves-Olarte E, von Eichel-Streiber C, Lopez-Goni I, Thelestam M, Arvidson S, Gorvel JP, Moreno E (2001) GTPases of the Rho subfamily are required for Brucella abortus internalization in nonprofessional phagocytes—direct activation of Cdc42. J Biol Chem 276:44435–44443
Celli J, Salcedo SP, Gorvel JP (2005) Brucella coopts the small GTPase Sar1 for intracellular replication. Proc Natl Acad Sci U S A 102:1673–1678
Kahl-McDonagh MM, Ficht TA (2006) Evaluation of protection afforded by Brucella abortus and Brucella melitensis unmarked deletion mutants exhibiting different rates of clearance in BALB/c mice. Infect Immun 74:4048–4057
Pei J, Ficht TA (2004) Brucella abortus rough mutants are cytopathic for macrophages in culture. Infect Immun 72(1):440–450
Hong PC, Tsolis RM, Ficht TA (2000) Identification of genes required for chronic persistence of Brucella abortus in mice. Infect Immun 68:4102–4107
Allen CA, Adams LG, Ficht TA (1998) Transposon-derived Brucella abortus rough mutants are attenuated and exhibit reduced intracellular survival. Infect Immun 66:1008–1016
Wu Q, Pei J, Turse C, Ficht TA (2006) Mariner mutagenesis of Brucella melitensis reveals genes with previously uncharacterized roles in virulence and survival. BMC Microbiol 6:102
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2014 Springer Science+Business Media New York
About this protocol
Cite this protocol
Pandey, A., Ding, S.L., Ficht, T.A., de Figueiredo, P. (2014). siRNA Screens Using Drosophila Cells to Identify Host Factors Required for Infection. In: Vergunst, A., O'Callaghan, D. (eds) Host-Bacteria Interactions. Methods in Molecular Biology, vol 1197. Humana Press, New York, NY. https://doi.org/10.1007/978-1-4939-1261-2_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-1261-2_13
Published:
Publisher Name: Humana Press, New York, NY
Print ISBN: 978-1-4939-1260-5
Online ISBN: 978-1-4939-1261-2
eBook Packages: Springer Protocols