Abstract
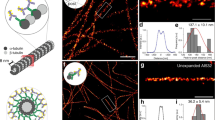
Multiple expansion microscopy approaches have been successfully used in the analysis of centrioles, centrosomes, and cilia, hel** to reveal the localization of numerous centrosomal and ciliary proteins at nanoscale resolution. In this chapter, we describe the use of two stable STED dyes in combination with expansion microscopy, which allows the robust detection by conventional and STED microscopy of proteins immunolabeled prior to sample expansion. We demonstrate the stability of these dyes during the crosslinking, polymerization, and denaturation steps of an expansion protocol thereby allowing their use in an immunolabel-first-expand-later approach. Our protocol overcomes the frequent technical limitation of poor, unreproducible binding of primary antibodies to proteins after denaturation. We demonstrate the applicability of this approach by analyzing both a centriole appendage protein Cep164 and a ciliary protein ARL13B.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Bornens M (2021) Centrosome organization and functions. Curr Opin Struct Biol 66:199–206
Nigg EA, Raff JW (2009) Centrioles, centrosomes, and cilia in health and disease. Cell 139(4):663–678
Arquint C, Gabryjonczyk AM, Nigg EA (2014) Centrosomes as signalling centres. Philos Trans R Soc Lond Ser B Biol Sci 369(1650):20130464
Gonczy P, Hatzopoulos GN (2019) Centriole assembly at a glance. J Cell Sci 132(4):jcs228833
Sullenberger C et al (2020) With age comes maturity: biochemical and structural transformation of a human centriole in the making. Cell 9(6):1429
Tanos BE et al (2013) Centriole distal appendages promote membrane docking, leading to cilia initiation. Genes Dev 27(2):163–168
Delgehyr N, Sillibourne J, Bornens M (2005) Microtubule nucleation and anchoring at the centrosome are independent processes linked by ninein function. J Cell Sci 118(Pt 8):1565–1575
Bowler M et al (2019) High-resolution characterization of centriole distal appendage morphology and dynamics by correlative STORM and electron microscopy. Nat Commun 10(1):993
Vorobjev IA, Chentsov Yu S (1982) Centrioles in the cell cycle. I. Epithelial cells. J Cell Biol 93(3):938–949
Gaudin N et al (2022) Evolutionary conservation of centriole rotational asymmetry in the human centrosome. eLife 11:e72382
van den Hoek H et al (2022) In situ architecture of the ciliary base reveals the stepwise assembly of intraflagellar transport trains. Science 377(6605):543–548
Le Guennec M et al (2020) A helical inner scaffold provides a structural basis for centriole cohesion. Sci Adv 6(7):eaaz4137
Schweizer N et al (2021) Sub-centrosomal map** identifies augmin-γTuRC as part of a centriole-stabilizing scaffold. Nat Commun 12(1):6042
Vásquez-Limeta A et al (2022) CPAP insufficiency leads to incomplete centrioles that duplicate but fragment. J Cell Biol 221(5):e202108018
Tian Y et al (2021) Superresolution characterization of core centriole architecture. J Cell Biol 220(4):e202005103
Gambarotto D et al (2019) Imaging cellular ultrastructures using expansion microscopy (U-ExM). Nat Methods 16(1):71–74
Sahabandu N et al (2019) Expansion microscopy for the analysis of centrioles and cilia. J Microsc 276(3):145–159
Ku T et al (2016) Multiplexed and scalable super-resolution imaging of three-dimensional protein localization in size-adjustable tissues. Nat Biotechnol 34(9):973–981
Graser S et al (2007) Cep164, a novel centriole appendage protein required for primary cilium formation. J Cell Biol 179(2):321–330
Cajanek L, Nigg EA (2014) Cep164 triggers ciliogenesis by recruiting Tau tubulin kinase 2 to the mother centriole. Proc Natl Acad Sci U S A 111(28):E2841–E2850
Caspary T, Larkins CE, Anderson KV (2007) The graded response to Sonic Hedgehog depends on cilia architecture. Dev Cell 12(5):767–778
Fisher S et al (2020) ARF family GTPases with links to cilia. Am J Physiol Cell Physiol 319(2):C404–c418
Kong D, Loncarek J (2021) Analyzing centrioles and cilia by expansion microscopy. Methods Mol Biol 2329:249–263
Acknowledgments
We thank Dr. Meredith Metzger and Dr. Catherine Sullenberger for critical reading of the manuscript. This work is supported by the Intramural Research Program of the National Institutes of Health (NIH), National Cancer Institute to JL.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2024 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Kong, D., Luvsanjav, D., Loncarek, J. (2024). Immunolabel-First-Expand-Later Expansion Microscopy Approach Using Stable STED Dyes. In: Mennella, V. (eds) Cilia. Methods in Molecular Biology, vol 2725. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-3507-0_5
Download citation
DOI: https://doi.org/10.1007/978-1-0716-3507-0_5
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-3506-3
Online ISBN: 978-1-0716-3507-0
eBook Packages: Springer Protocols