Abstract
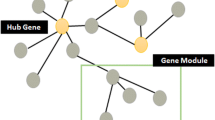
Regulatory network is often characterized by complex interactions between transcription factors (TFs) and target genes. The synergistic regulations among multiple TFs may co-induce/co-suppress the expressions of the similar target genes. Such information is important for understanding stress response signaling pathways in plants. In this chapter, we present a computational tool, CoReg, for mining co-regulatory gene modules from network topology. The analysis results can be used to interpret co-regulation effects in regulatory networks generated by high-through TF–DNA interaction screenings such as yeast-one-hybrid, ChIP-seq, and DAP-seq.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Reece-Hoyes JS, Diallo A, Lajoie B, Kent A, Shrestha S, Kadreppa S et al (2011) Enhanced yeast one-hybrid assays for high-throughput gene-centered regulatory network map**. Nat Methods 8:1059–1064
Bulyk ML (2006) Protein binding microarrays for the characterization of DNA-protein interactions. Adv Biochem Eng Biotechnol 104:65–68
Johnson DS, Mortazavi A, Myers RM, Wold B (2007) Genome-wide map** of in vivo protein-DNA interactions. Science 316:1497–1502
O’Malley RC, Huang SSC, Song L, Lewsey MG, Bartlett A, Nery JR et al (2016) Cistrome and epicistrome features shape the regulatory DNA landscape. Cell 165:1280–1292
Liu W, Stewart CN (2016) Plant synthetic promoters and transcription factors. Curr Opin Biotechnol 27:36–44
Wu WS, Lai FJ (2015) Functional redundancy of transcription factors explains why most binding targets of a transcription factor are not affected when the transcription factor is knocked out. BMC Syst Biol 9(Suppl 6): S2
Song Q, Grene R, Heath LS, Li S (2017) Identification of regulatory modules in genome scale transcription regulatory networks. BMC Syst Biol 11:140
Taylor-Teeples M, Lin L, De Lucas M, Turco G, Toal TW, Gaudinier A et al (2015) An Arabidopsis gene regulatory network for secondary cell wall synthesis. Nature 517: 571–575
Acknowledgments
We thank Virginia Agricultural Experiment Station (Blacksburg), USADA National Institute of Food and Agriculture, US Department of Agriculture (Washington, DC) for supporting the development of CoReg.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Song, Q., Li, S. (2023). Identification of Plant Co-regulatory Modules Using CoReg. In: Song, Q., Tao, Z. (eds) Transcription Factor Regulatory Networks. Methods in Molecular Biology, vol 2594. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-2815-7_16
Download citation
DOI: https://doi.org/10.1007/978-1-0716-2815-7_16
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-2814-0
Online ISBN: 978-1-0716-2815-7
eBook Packages: Springer Protocols