Abstract
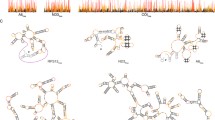
Mitochondrial pre-mRNAs in African trypanosomes adopt intricately folded, highly stable 2D and 3D structures. The RNA molecules are substrates of a U-nucleotide-specific insertion/deletion-type RNA editing reaction, which is catalyzed by a 0.8 MDa protein complex known as the editosome. RNA binding to the editosome is followed by a chaperone-mediated RNA remodeling reaction. The reaction increases the dynamic of specifically U-nucleotides to lower their base-pairing probability and as a consequence generates a simplified RNA folding landscape that is critical for the progression of the editing reaction cycle. Here we describe a chemical map** method to quantitatively monitor the chaperone-driven structural changes of pre-edited mRNAs upon editosome binding. The method is known as selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE). SHAPE is based on the differential electrophilic modification of ribose 2′-hydroxyl groups in structurally constraint (double-stranded) versus structurally unconstrained (single-stranded) nucleotides. Electrophilic anhydrides such as 1-methyl-7-nitroisatoic anhydride are used as probing reagents, and the ribose 2′-modified nucleotides are mapped as abortive cDNA synthesis products. As a result, SHAPE allows the identification of all single-stranded and base-paired regions in a given RNA, and the data are used to compute experimentally derived RNA 2D structures. A side-by-side comparison of the RNA 2D folds in the pre- and post-chaperone states finally maps the chaperone-induced dynamic of the different pre-mRNAs with single-nucleotide resolution.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Göringer HU (2012) RNA editing in African trypanosomes: a U-ser’s G-U-ide. In: Bindereif A (ed) RNA metabolism in trypanosomes, Nucleic acids and molecular biology, vol 28. Springer, Heidelberg, pp 149–165
Aphasizhev R, Aphasizheva I (2014) Mitochondrial RNA editing in trypanosomes: small RNAs in control. Biochimie 100:125–131. https://doi.org/10.1016/j.biochi.2014.01.003
Golas MM, Böhm C, Sander B, Effenberger K, Brecht M, Stark H, Göringer HU (2009) Snapshots of the RNA editing machine in trypanosomes captured at different assembly stages in vivo. EMBO J 28:766–778. https://doi.org/10.1038/emboj.2009.19
Göringer HU (2012) ‘Gestalt,’ composition and function of the Trypanosoma brucei editosome. Annu Rev Microbiol 66:65–82. https://doi.org/10.1146/annurev-micro-092611-150150
Leeder WM, Hummel NF, Göringer HU (2016) Multiple G-quartet structures in pre-edited mRNAs suggest evolutionary driving force for RNA editing in trypanosomes. Sci Rep 6:29810. https://doi.org/10.1038/srep29810
Müller UF, Göringer HU (2002) Mechanism of the gBP21-mediated RNA/RNA annealing reaction: matchmaking and charge reduction. Nucleic Acids Res 30:447–455
Kruse E, Voigt C, Leeder WM, Göringer HU (2013) RNA helicases involved in U-insertion/deletion-type RNA editing. Biochim Biophys Acta 1829:835–841. https://doi.org/10.1016/j.bbagrm.2013.04.003
Leeder WM, Voigt C, Brecht M, Göringer HU (2016) The RNA chaperone activity of the Trypanosoma brucei editosome raises the dynamic of bound pre-mRNAs. Sci Rep 6:19309. https://doi.org/10.1038/srep19309
Voigt C, Dobrychlop M, Kruse E, Czerwoniec A, Kasprzak JM, Bytner P, Campo CD, Leeder WM, Bujnicki JM, Göringer HU (2018) The OB-fold proteins of the Trypanosoma brucei editosome execute RNA-chaperone activity. Nucleic Acids Res 46:10353–10367. https://doi.org/10.1093/nar/gky668
Merino EJ, Wilkinson KA, Coughlan JL, Weeks KM (2005) RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J Am Chem Soc 127:4223–4231
Wilkinson KA, Merino EJ, Weeks KM (2006) Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat Protoc 1:1610–1616
Low JT, Weeks KM (2010) SHAPE-directed RNA secondary structure prediction. Methods 52:150–158. https://doi.org/10.1016/j.ymeth.2010.06.007
McGinnis JL, Dunkle JA, Cate JH, Weeks KM (2012) The mechanisms of RNA SHAPE chemistry. J Am Chem Soc 134:6617–1624. https://doi.org/10.1021/ja2104075
Lee B, Flynn RA, Kadina A, Guo JK, Kool ET, Chang HY (2017) Comparison of SHAPE reagents for map** RNA structures inside living cells. RNA 23:169–174. https://doi.org/10.1261/rna.058784.116
Deigan KE, Li TW, Mathews DH, Weeks KM (2009) Accurate SHAPE-directed RNA structure determination. Proc Natl Acad Sci U S A 106:97–102. https://doi.org/10.1073/pnas.0806929106
Lorenz R, Luntzer D, Hofacker IL, Stadler PF, Wolfinger MT (2016) SHAPE directed RNA folding. Bioinformatics 32:145–147. https://doi.org/10.1093/bioinformatics/btv523
Wang B, Wilkinson KA, Weeks KM (2008) Complex ligand-induced conformational changes in tRNA(Asp) revealed by single-nucleotide resolution SHAPE chemistry. Biochemistry 47:3454–3461. https://doi.org/10.1021/bi702372x
Watts JM, Dang KK, Gorelick RJ, Leonard CW, Bess JW Jr, Swanstrom R, Burch CL, Weeks KM (2009) Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460:711–716. https://doi.org/10.1038/nature08237
Weeks KM, Mauger DM (2011) Exploring RNA structural codes with SHAPE chemistry. Acc Chem Res 44:1280–1291. https://doi.org/10.1021/ar200051h
Turner R, Shefer K, Ares M Jr (2013) Safer one-pot synthesis of the ‘SHAPE’ reagent 1-methyl-7-nitroisatoic anhydride (1m7). RNA 19:1857–1863. https://doi.org/10.1261/rna.042374.113
Vasa SM, Guex N, Wilkinson KA, Weeks KM, Giddings MC (2008) ShapeFinder: a software system for high-throughput quantitative analysis of nucleic acid reactivity information resolved by capillary electrophoresis. RNA 14:1979–1990. https://doi.org/10.1261/rna.1166808
Karabiber F, McGinnis JL, Favorov OV, Weeks KM (2013) QuShape: rapid, accurate, and best-practices quantification of nucleic acid probing information, resolved by capillary electrophoresis. RNA 19:63–73. https://doi.org/10.1261/rna.036327.112
Cantara WA, Hatterschide J, Wu W, Musier-Forsyth K (2017) RiboCAT: a new capillary electrophoresis data analysis tool for nucleic acid probing. RNA 23:240–249. https://doi.org/10.1261/rna.058404.116
Yoon S, Kim J, Hum J, Kim H, Park S, Kladwang W, Das R (2011) HiTRACE: high-throughput robust analysis for capillary electrophoresis. Bioinformatics 27:1798–1805. https://doi.org/10.1093/bioinformatics/btr277
Kim H, Cordero P, Das R, Yoon S (2013) HiTRACE-Web: an online tool for robust analysis of high-throughput capillary electrophoresis. Nucleic Acids Res 41:W492–W498. https://doi.org/10.1093/nar/gkt501
Melton DA, Krieg PA, Rebagliati MR, Maniatis T, Zinn K, Green MR (1984) Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res 12:7035–7056. https://doi.org/10.1093/nar/12.18.7035
Cantor CR, Warshaw MM, Shapiro H (1970) Oligonucleotide interactions. III. Circular dichroism studies of the conformation of deoxyoligonucleotides. Biopolymers 9:1059–1077. https://doi.org/10.1002/bip.1970.360090909
Fasman GD (ed) (1975) Handbook of biochemistry and molecular biology, Nucleic acids, vol 1, 3rd edn. CRC Press, Boca Raton, FL, p 589
Rigaut G, Shevchenko A, Rutz B, Wilm M, Mann M, Séraphin B (1999) A generic protein purification method for protein complex characterization and proteome exploration. Nat Biotechnol 17:1030–1032. https://doi.org/10.1038/13732
Igo RP Jr, Palazzo SS, Burgess ML, Panigrahi AK, Stuart K (2000) Uridylate addition and RNA ligation contribute to the specificity of kinetoplastid insertion RNA editing. Mol Cell Biol 20:8447–8457. https://doi.org/10.1128/MCB.20.22.8447-8457.2000
Igo RP Jr, Weston DS, Ernst NL, Panigrahi AK, Salavati R, Stuart K (2002) Role of uridylate specific exoribonuclease activity in Trypanosoma brucei RNA editing. Eukaryot Cell 1:112–118. https://doi.org/10.1128/EC.1.1.112-118.2002
Böhm C, Katari VS, Brecht M, Göringer HU (2012) Trypanosoma brucei 20S editosomes have one RNA substrate-binding site and execute RNA unwinding activity. J Biol Chem 287:26268–26277. https://doi.org/10.1074/jbc.M112.365916
Rio DC (2012) Filter-binding assay for analysis of RNA-protein interactions. Cold Spring Harb Protoc 2012:1078–1081. https://doi.org/10.1101/pdb.prot071449
Katari VS, van Esdonk L, Göringer HU (2013) Molecular crowding inhibits U-insertion/deletion RNA editing in vitro: consequences for the in vivo reaction. PLoS One 8:e83796. https://doi.org/10.1371/journal.pone.0083796
Lusvarghi S, Sztuba-Solinska J, Purzycka KJ, Rausch JW, Le Grice SF (2013) RNA secondary structure prediction using high-throughput SHAPE. J Vis Exp 75:e50243. https://doi.org/10.3791/50243.
Bhattacharyya D, Mirihana Arachchilage G, Basu S (2016) Metal cations in G-quadruplex folding and stability. Front Chem 4(38). https://doi.org/10.3389/fchem.2016.00038
Acknowledgments
We acknowledge funding by the German Research Foundation (DFG) as part of the Collaborative Research Center (CRC)902 (Molecular Principles of RNA-based Regulation).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2020 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Leeder, WM., Göringer, H.U. (2020). Map** the RNA Chaperone Activity of the T. brucei Editosome Using SHAPE Chemical Probing. In: Heise, T. (eds) RNA Chaperones. Methods in Molecular Biology, vol 2106. Humana, New York, NY. https://doi.org/10.1007/978-1-0716-0231-7_10
Download citation
DOI: https://doi.org/10.1007/978-1-0716-0231-7_10
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-0716-0230-0
Online ISBN: 978-1-0716-0231-7
eBook Packages: Springer Protocols