Abstract
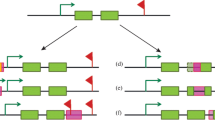
Transposable elements, or transposons, are DNA segments that are repeated within the same genome and are an important component of the genomes of most species. It is generally believed that they play an important role in evolution and genome restructuring. In this work we built a network based on a score which represents the transposon identity. This score is calculated by comparing all currently known D. melanogaster transposons to each other using a Neddleman-Wunsch alignment algorithm. We then use this score to build networks with transposons having a minimal value of identity. We start with networks of transposons with total identity (all have score one) to networks where they may have any identity score (all have non-negative score). The number of successful comparisons as a function of the minimal score shows an abrupt transition for minimal scores at 0.25 which can be associated to general properties of repetitions usually found in genomes. We also show that this score leads to a transition in the topology of transposon networks from scale-free to almost fully-connected.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Preview
Unable to display preview. Download preview PDF.
Similar content being viewed by others
References
Erdős, P., Rényi, A.: On the evolution of random graphs. Publ. Math. Inst. Hung. Acad. Sci. 5, 17–61 (1960)
Barabási, A., Albert, R.: Emergence of scaling in random networks. Science 286(5439), 509 (1999)
Barabasi, A., Oltvai, Z.: Network biology: understanding the cell’s functional organization. Nature Reviews Genetics 5(2), 101–113 (2004)
Jeong, H., Tombor, B., Albert, R., Oltvai, Z., Barabasi, A., et al.: The large-scale organization of metabolic networks. Nature 407(6804), 651–654 (2000)
Bartolome, C., Maside, X., Charlesworth, B.: On the abundance and distribution of transposable elements in the genome of Drosophila melanogaster. Molecular Biology and Evolution 19(6), 926–937 (2002)
Rizzon, C., Marais, G., Gouy, M., Biémont, C.: Recombination rate and the distribution of transposable elements in the Drosophila melanogaster genome. Genome Research 12, 400–407 (2002)
Durbin, R., Eddy, S.R., Krogh, A., Mitchison, G.: Biological sequence analysis: probabilistic models of proteins and nucleic acids. Cambridge Univ. Press, Cambridge (2000)
Whiteford, N., Haslam, N., Weber, G., Prügel-Bennett, A., Essex, J.W., Roach, P.L., Bradley, M., Neylon, C.: An analysis of the feasibility of short read sequencing. Nucl. Acids. Res. 33(19), e171 (2005)
Author information
Authors and Affiliations
Editor information
Rights and permissions
Copyright information
© 2008 Springer-Verlag Berlin Heidelberg
About this paper
Cite this paper
Castro-e-Silva, A., Weber, G., Machado, R.F., Wanner, E.F., Guerra-Sá, R. (2008). Identity Transposon Networks in D. melanogaster . In: Bazzan, A.L.C., Craven, M., Martins, N.F. (eds) Advances in Bioinformatics and Computational Biology. BSB 2008. Lecture Notes in Computer Science(), vol 5167. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-540-85557-6_15
Download citation
DOI: https://doi.org/10.1007/978-3-540-85557-6_15
Publisher Name: Springer, Berlin, Heidelberg
Print ISBN: 978-3-540-85556-9
Online ISBN: 978-3-540-85557-6
eBook Packages: Computer ScienceComputer Science (R0)