Abstract
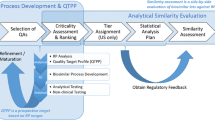
FDA recommends a stepwise approach for obtaining the totality-of-the-evidence for assessing biosimilarity between a proposed biosimilar product and its corresponding reference biologic product being considered (US Food and Drug Administration.: Guidance for industry: scientific considerations in demonstrating biosimilarity to a reference product. US Food and Drug Administration, Silver Spring, 2015 [6]). The stepwise approach starts with analytical studies for assessing similarity in critical quality attributes (CQAs), which are relevant to clinical outcomes. For critical quality attributes that are most relevant to clinical outcomes (Tier 1 CQAs), FDA requires equivalence testing to be performed for similarity assessment, based on an equivalence acceptance criteria. In practice, the number of Tier 1 CQAs might be greater than one, and should be no more than four. The number of biosimilar lots is often recommended to be no less than 10, and the ratio between the reference product sample size and biosimilar product sample size is recommended within the range from \( 2/3 \) to \( 3/2 \) (US Food and Drug Administration.: Guidance for industry: Statistical Approaches to Evaluate Analytical Similarity. US Food and Drug Administration, Silver Spring, 2017 [7]). Accordingly, we derive the formulas for the power calculation for the sample size for analytical similarity assessment based on the equivalence testing currently used in analytical biosimilar assessment (Tsong et al. J Biopharm Stat 27:197–205, (2017)[10]).
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Burdick, R.K., Thomas, N., Cheng, A.: Statistical considerations in demonstrating cmc analytical similarity for a biosimilar product. Stat. Biopharm. Res. 9(3), 249–257 (2017)
Chen, Y.M., Weng, Y.T., Dong, X., Tsong, Y.: Wald tests for variance-adjusted equivalence assessment with normal endpoints. J. Biopharm. Stat. 27(2), 308–316 (2017)
Chow, S.C., Song, F., Bai, H.: Sample size requirement in analytical studies for similarity assessment. J. Biopharm. Stat. 27(2), 233–238 (2017)
Dong, X., Bian, Y., Tsong, Y., Wang, T.: Exact test-based approach for equivalence test with parameter margin. J. Biopharm. Stat. 27(2), 317–330 (2017)
Dong, X., Weng, Y.T., Tsong, Y.: Adjustment for unbalanced sample size for analytical biosimilar equivalence assessment. J. Biopharm. Stat. 27(2), 220–232 (2017)
Joarder, A.H.: Moments of the product and ratio of two correlated chi-square variables. Stat. Pap. 50(3), 581–592 (2009)
Smolyak, S.: Quadrature and interpolation formulas for tensor products of certain classes of functions. Soviet Math. Dokl. 4, 240–243. American Mathematical Society (1963)
Tsong, Y., Dong, X., Shen, M.: Development of statistical methods for analytical similarity assessment. J. Biopharm. Stat. 27(2), 197–205 (2017)
US Food and Drug Administration.: Guidance for industry: scientific considerations in demonstrating biosimilarity to a reference product. Silver Spring, MD., US Food and Drug Administration (2015)
US Food and Drug Administration.: Guidance for industry: Statistical Approaches to Evaluate Analytical Similarity. Silver Spring, MD., US Food and Drug Administration (2017)
Weng, Y.T., Tsong, Y., Shen, M., and Wang, C.: Improved Wald test for equivalence assessment of analytical biosimilarity. Int. J. of Clinical Biostat. And Biometrics. (2018) Submitted
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Additional information
Disclaimer: This article reflects the views of the authors and should not be construed to represent FDA’s views or policies.
Appendix
Appendix
-
1.
The proof of Theorem 1.
Proof: Because the power function is
given \( \upmu_{\text{T}} -\upmu_{\text{R}} =\uptheta \in \left( { - 1.5\upsigma_{\text{R}} , + 1.5\upsigma_{\text{R}} } \right),\upsigma_{\text{T}} ,\upsigma_{\text{R}} \). The inequailities in (A.1) is equivalent to
Because \( {\text{Z}} = \frac{{\left( {\overline{\text{X}}_{\text{T}} - \overline{\text{X}}_{\text{R}} -\uptheta} \right)}}{{\sqrt {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}} }}\sim {\text{N}}\left( {0, 1} \right) \), the inequailities in (A.2) is equivalent to
Let \( {\text{x}}_{1} = \frac{{\left( {{\text{n}}_{\text{T}} - 1} \right){\text{S}}_{\text{T}}^{2} }}{{\upsigma_{\text{T}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{T}} - 1}}^{2} \) and \( {\text{x}}_{2} = \frac{{\left( {{\text{n}}_{\text{R}} - 1} \right){\text{S}}_{\text{R}}^{2} }}{{\upsigma_{\text{R}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{R}} - 1}}^{2} \), then \( {\text{S}}_{\text{T}}^{2} = \frac{{{\text{x}}_{1}\upsigma_{\text{T}}^{2} }}{{\left( {{\text{n}}_{\text{T}} - 1} \right)}} \) and \( {\text{S}}_{\text{R}}^{2} = \frac{{{\text{x}}_{2}\upsigma_{\text{R}}^{2} }}{{\left( {{\text{n}}_{\text{R}} - 1} \right)}} \), the inequalities in (A.3) is
Let \( {\text{A}}_{1} = \frac{{1.5\upsigma_{\text{R}} -\uptheta}}{{\sqrt {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}} }} - {\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}} \sqrt {\frac{{\frac{{{\text{x}}_{1}\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} \left( {{\text{n}}_{\text{T}} - 1} \right)}} + \frac{{{\text{x}}_{2}\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} \left( {{\text{n}}_{\text{R}} - 1} \right)}}}}{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}}}} \) and \( {\text{B}}_{1} = \frac{{ - 1.5\upsigma_{\text{R}} -\uptheta}}{{\sqrt {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}} }} + {\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}} \sqrt {\frac{{\frac{{{\text{x}}_{1}\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} \left( {{\text{n}}_{\text{T}} - 1} \right)}} + \frac{{{\text{x}}_{2}\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} \left( {{\text{n}}_{\text{R}} - 1} \right)}}}}{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}}}} \),
From (A.4) since \( {\text{B}}_{1} < {\text{A}}_{1} \), we can get that \( \frac{{{\text{x}}_{1}\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} \left( {{\text{n}}_{\text{T}} - 1} \right)}} + \frac{{{\text{x}}_{2}\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} \left( {{\text{n}}_{\text{R}} - 1} \right)}} < \frac{{9\upsigma_{\text{R}}^{2} }}{{4{\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}}^{2} }} \), then the power function (A.1) is
Given \( {\text{x}}_{1} \sim\upchi_{{{\text{n}}_{\text{T}} - 1}}^{2} \) and \( {\text{x}}_{2} \sim\upchi_{{{\text{n}}_{\text{R}} - 1}}^{2} \), the power function is
where \( {\text{f}}\left( {{\text{x}}_{1} , {\text{x}}_{2} } \right) = {\text{f}}\left( {{\text{x}}_{1} } \right){\text{f}}\left( {{\text{x}}_{2} } \right) \) is the density function of two independent Chi-square distributions with \( {\text{x}}_{1} \sim\upchi_{{{\text{n}}_{\text{T}} - 1}}^{2} \) and \( {\text{x}}_{2} \sim\upchi_{{{\text{n}}_{\text{R}} - 1}}^{2} \). \( {\mathbf{I}}\{ .\} \) is the indication function to restrict the triangle area formed by \( {\text{x}}_{1} \) and \( {\text{x}}_{2} \).
-
2.
The proof of Theorem 2.
Proof: By using the Satterthwaite approximation
Where \( {\text{df}}^{ *} = \left( {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}} \right)^{2} /\left( {\frac{{\upsigma_{\text{T}}^{4} }}{{\left( {{\text{n}}_{\text{T}}^{ *} } \right)^{2} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{\text{R}}^{4} }}{{\left( {{\text{n}}_{\text{R}}^{ *} } \right)^{2} ({\text{n}}_{\text{R}} - 1)}}} \right) \), let \( {\text{S}}^{2} = \frac{{{\text{S}}_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{{\text{S}}_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }} \) and \( {\text{x}} = \frac{{{\text{df}}^{ *} {\text{S}}^{2} }}{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}}}\sim\upchi_{{{\text{df}}^{ *} }}^{2} \), then the inequalities (A.3) is
Let \( {\text{A}}_{1}^{ '} = \frac{{1.5\upsigma_{\text{R}} -\uptheta}}{{\sqrt {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}} }} - {\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}} \sqrt {\frac{\text{x}}{{{\text{df}}^{ *} }}} \sqrt {\frac{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}}}{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}}}} \) and \( {\text{B}}_{1}^{ '} = \frac{{ - 1.5\upsigma_{\text{R}} -\uptheta}}{{\sqrt {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}} }} + {\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}} \sqrt {\frac{\text{x}}{{{\text{df}}^{ *} }}} \sqrt {\frac{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}}}{{\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}} }}}}} \), using \( {\text{B}}_{1}^{ '} < {\text{A}}_{1}^{ '} \), we can get \( {\text{x}} < L = \frac{{9\upsigma_{\text{R}}^{2} {\text{df}}^{ *} }}{{4{\text{t}}_{{{\text{df}}^{ *} , 1 -\upalpha}}^{2} \left( {\frac{{\upsigma_{\text{T}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{\text{R}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}} \right)}} \), the power function (A.1) is \( {\text{E}}_{\text{x }} \left\{ {{\text{P}}({\text{B}}_{1}^{ '} < Z < {\text{A}}_{1}^{ '} )} \right\}, \) which is equivalent to
-
3.
The proof of Theorem 4.
Proof: If there are 2 quality attributes that are correlated, assume the 2 quality attributes in the reference product \( {\text{X}}_{{1,{\text{R}}}} \) and \( {\text{X}}_{{2, {\text{R}}}} \) have the correlation value \( \uprho \); the same as the 2 quality attributes in the test product \( {\text{X}}_{{1,{\text{T}}}} \) and \( {\text{X}}_{{2, {\text{T}}}} \)
We further assume the sample size of either quality attribute is the same in the reference product or in the test product, that is \( {\text{n}}_{{1,{\text{R}}}} = {\text{n}}_{{2,{\text{R}}}} = {\text{n}}_{\text{R}} \) and \( {\text{n}}_{{1,{\text{T}}}} = {\text{n}}_{{2,{\text{T}}}} = {\text{n}}_{\text{T}} \), given the hypotheses in (6) and test statistics in (7), the power of passing both equivalence testings is equivalent to calculate
The degrees of freedom \( {\text{df}}_{1}^{ *} = \left( {\frac{{\upsigma_{{1, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{{1, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}} \right)^{2} /\left( {\frac{{\upsigma_{{1, {\text{T}}}}^{4} }}{{\left( {{\text{n}}_{\text{T}}^{ *} } \right)^{2} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{1, {\text{R}}}}^{4} }}{{\left( {{\text{n}}_{\text{R}}^{ *} } \right)^{2} ({\text{n}}_{\text{R}} - 1)}}} \right) \) and \( {\text{df}}_{2}^{ *} = \left( {\frac{{\upsigma_{{2, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}}^{ *} }} + \frac{{\upsigma_{{2, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}}^{ *} }}} \right)^{2} /\left( {\frac{{\upsigma_{{1, {\text{T}}}}^{4} }}{{\left( {{\text{n}}_{\text{T}}^{ *} } \right)^{2} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{1, {\text{R}}}}^{4} }}{{\left( {{\text{n}}_{\text{R}}^{ *} } \right)^{2} ({\text{n}}_{\text{R}} - 1)}}} \right) \). Let \( {\text{u}}_{\text{T}} = \frac{{({\text{n}}_{\text{T}} - 1){\text{S}}_{{1, {\text{T}}}}^{2} }}{{\upsigma_{{1, {\text{T}}}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{T}} - 1}}^{2} \), \( {\text{v}}_{\text{T}} = \frac{{({\text{n}}_{\text{T}} - 1){\text{S}}_{{2, {\text{T}}}}^{2} }}{{\upsigma_{{2, {\text{T}}}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{T}} - 1}}^{2} \) and \( {\text{u}}_{\text{R}} = \frac{{({\text{n}}_{\text{R}} - 1){\text{S}}_{{1, {\text{R}}}}^{2} }}{{\upsigma_{{1, {\text{R}}}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{R}} - 1}}^{2} \), \( {\text{v}}_{\text{R}} = \frac{{({\text{n}}_{\text{R}} - 1){\text{S}}_{{2, {\text{R}}}}^{2} }}{{\upsigma_{{2, {\text{R}}}}^{2} }}\sim\upchi_{{{\text{n}}_{\text{R}} - 1}}^{2} \), and \( \uptheta_{1} =\upmu_{{1,{\text{T}}}} -\upmu_{{1,{\text{R}}}} \), \( \uptheta_{2} =\upmu_{{2,{\text{T}}}} -\upmu_{{2,{\text{R}}}} \). The inequailities in (A.9) is equivalent to
where \( {\mathbf{B}} \) is a bi-variate normal distribution with mean \( 0 \) and variance-covariance matrix
let \( \uptau_{1} = \sqrt {\frac{{\upsigma_{{1, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{{1, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}} }}} \); \( \uptau_{2} = \sqrt {\frac{{\upsigma_{{2, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{{2, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}} }}} \); \( \uprho^{ *} = \frac{{\uprho \upsigma _{{1,{\text{T}}}}\upsigma_{{2, {\text{T}}}} }}{{{\text{n}}_{\text{T}} }} + \frac{{\uprho \upsigma _{{1,{\text{R}}}}\upsigma_{{2, {\text{R}}}} }}{{{\text{n}}_{\text{R}} }}/\sqrt {\left( {\frac{{\upsigma_{{1, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{{1, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}} }}} \right)\left( {\frac{{\upsigma_{{2, {\text{T}}}}^{2} }}{{{\text{n}}_{\text{T}} }} + \frac{{\upsigma_{{2, {\text{R}}}}^{2} }}{{{\text{n}}_{\text{R}} }}} \right)} \), then \( {\mathbf{B}} \) is a bi-variate normal distribution with mean \( 0 \) and variance-covariance matrix \( \left[ {\begin{array}{*{20}c} {\uptau_{1}^{2} } & {\uprho^{ *}\uptau_{1}\uptau_{2} } \\ {\uprho^{ *}\uptau_{1}\uptau_{2} } & {\uptau_{2}^{2} } \\ \end{array} } \right] \). The power function would be
Let \( {\text{U}} = \frac{{\upsigma_{{1, {\text{T}}}}^{2} {\text{u}}_{\text{T}} }}{{{\text{n}}_{\text{T}}^{ *} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{1, {\text{R}}}}^{2} {\text{u}}_{\text{R}} }}{{{\text{n}}_{\text{R}}^{ *} ({\text{n}}_{\text{R}} - 1)}} \le \frac{{9\upsigma_{{1.{\text{R}}}}^{2} }}{{4{\text{t}}_{{{\text{df}}_{1}^{ *} , 1 -\upalpha}}^{2} }} \) and \( {\text{V}} = \frac{{\upsigma_{{2, {\text{T}}}}^{2} {\text{v}}_{\text{T}} }}{{{\text{n}}_{\text{T}}^{ *} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{2, {\text{R}}}}^{2} {\text{v}}_{\text{R}} }}{{{\text{n}}_{\text{R}}^{ *} ({\text{n}}_{\text{R}} - 1)}} \le \frac{{9\upsigma_{{2.{\text{R}}}}^{2} }}{{4{\text{t}}_{{{\text{df}}_{2}^{ *} , 1 -\upalpha}}^{2} }} \) by the inequalities (A.10), the limits \( {\text{LL}}_{1} = - 1.5\upsigma_{{1, {\text{R}}}} -\uptheta_{1} + {\text{t}}_{{{\text{df}}_{1}^{ *} , 1 -\upalpha}} \sqrt {\text{U}} \), \( {\text{LL}}_{2} = - 1.5\upsigma_{{2, {\text{R}}}} -\uptheta_{2} + {\text{t}}_{{{\text{df}}_{2}^{ *} , 1 -\upalpha}} \sqrt {\text{V}} \) and \( {\text{UL}}_{1} = 1.5\upsigma_{{1, {\text{R}}}} -\uptheta_{1} - {\text{t}}_{{{\text{df}}_{1}^{ *} , 1 -\upalpha}} \sqrt {\text{U}} \), \( {\text{UL}}_{2} = 1.5\upsigma_{{2, {\text{R}}}} -\uptheta_{2} - {\text{t}}_{{{\text{df}}_{2}^{ *} , 1 -\upalpha}} \sqrt {\text{V}} \); then
The problem remains to find the joint distribution of U and V. Please note that from theorem 4, \( {\text{F}}\left( { {\text{u}}_{\text{T}} , {\text{u}}_{\text{R}} ,{\text{v}}_{\text{T}} , {\text{v}}_{\text{R}} } \right) = {\text{f}}_{2} \left( {{\text{u}}_{\text{T}} , {\text{v}}_{\text{T}} } \right) * {\text{f}}_{2} \left( {{\text{u}}_{\text{R}} , {\text{v}}_{\text{R}} } \right) \) where \( {\text{f}}_{2} \) is the joint density of a bivariate chi-square distribution. Let \( \frac{{\upsigma_{{1, {\text{T}}}}^{2} {\text{u}}_{\text{T}} }}{{{\text{n}}_{\text{T}}^{ *} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{1, {\text{R}}}}^{2} {\text{u}}_{\text{R}} }}{{{\text{n}}_{\text{R}}^{ *} ({\text{n}}_{\text{R}} - 1)}} =\uplambda_{1} {\text{u}}_{\text{T}} +\upmu_{1} {\text{u}}_{\text{R}} = {\text{U}} \) and \( \frac{{\upsigma_{{2, {\text{T}}}}^{2} {\text{v}}_{\text{T}} }}{{{\text{n}}_{\text{T}}^{ *} ({\text{n}}_{\text{T}} - 1)}} + \frac{{\upsigma_{{2, {\text{R}}}}^{2} {\text{v}}_{\text{R}} }}{{{\text{n}}_{\text{R}}^{ *} ({\text{n}}_{\text{R}} - 1)}} =\uplambda_{2} {\text{v}}_{\text{T}} +\upmu_{2} {\text{v}}_{\text{R}} = {\text{V}} \), and we want to get the joint distribution of \( {\text{U }} \) and \( {\text{V}} \). The Jacobian transformation matrix is
Thus
We then integrate out \( {\text{u}}_{\text{R}} , {\text{v}}_{\text{T}} \), we can get the joint distribution of \( {\text{U}} \) and \( {\text{V}} \)
We can get the power function as
where \( {\text{f}}_{1} \left( {{\text{x}}_{1} , {\text{x}}_{2} } \right) = \frac{1}{{2\uppi \uptau _{1}\uptau_{2} \sqrt {1 -\uprho^{ * 2} } }}{ \exp }\left\{ { - \frac{{{\text{x}}_{1}^{2} }}{{2\uptau_{1}^{2} \left( {1 -\uprho^{ * 2} } \right)}} + \frac{{\uprho^{ *} {\text{x}}_{1} {\text{x}}_{2} }}{{\uptau_{1}\uptau_{2} \left( {1 -\uprho^{ * 2} } \right)}} - \frac{{{\text{x}}_{2}^{2} }}{{2\uptau_{2}^{2} \left( {1 -\uprho^{ * 2} } \right)}}} \right\} \) be the joint distribution of the bi-variate normal distribution with mean zero and variance-covariance matrix \( \left[ {\begin{array}{*{20}c} {\uptau_{1}^{2} } & {\uprho^{ *}\uptau_{1}\uptau_{2} } \\ {\uprho^{ *}\uptau_{1}\uptau_{2} } & {\uptau_{2}^{2} } \\ \end{array} } \right] \).
Rights and permissions
Copyright information
© 2019 This is a U.S. government work and not under copyright protection in the U.S.; foreign copyright protection may apply
About this paper
Cite this paper
Wang, T., Tsong, Y., Shen, M. (2019). Sample Size Consideration for Equivalent Test of Tier-1 Quality Attributes for Analytical Biosimilarity Assessment. In: Liu, R., Tsong, Y. (eds) Pharmaceutical Statistics. MBSW 2016. Springer Proceedings in Mathematics & Statistics, vol 218. Springer, Cham. https://doi.org/10.1007/978-3-319-67386-8_3
Download citation
DOI: https://doi.org/10.1007/978-3-319-67386-8_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-67385-1
Online ISBN: 978-3-319-67386-8
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)