Abstract
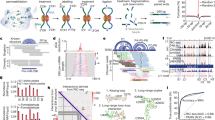
Mammalian genomes encode large amounts of noncoding RNAs (ncRNAs), which tend to form intricate structures and interact with their target RNA molecules through complementary base pairing with the help of proteins. Map** of intra- and inter-molecular RNA–RNA interactions (RRIs) is required to unravel the structure and targets of ncRNAs which are two essential aspects for understanding the molecular mechanisms of ncRNAs in various biological processes. At this frontiers, we recently invented RNA in situ conformation sequencing (RIC-seq) technology to profile protein-mediated RNA–RNA spatial interactions at single-nucleotide resolution in an unbiased manner. We have demonstrated that RIC-seq-identified RRIs are helpful for simultaneously deducing ncRNA structures and targets. Here, we summarize methods for probing RRIs and describe a step-by-step protocol for generating a successful RIC-seq library.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
Abbreviations
- ncRNA:
-
Noncoding RNA
- rRNA:
-
Ribosomal RNA
- tRNA:
-
Transfer RNA
- piRNA:
-
Piwi-interacting RNA
- lncRNA:
-
Long noncoding RNA
- circRNA:
-
Circular RNA
- eRNA:
-
Enhancer RNA
- AMT:
-
4′-Aminomethyltrioxsalen
- RBP:
-
RNA-binding protein
- EMSA:
-
Electrophoretic mobility shift assay
- SPR:
-
Surface plasmon resonance
- FRET:
-
Fluorescence resonance energy transfer
- CLASH:
-
Crosslinking, ligation, and sequencing of hybrids
- RAP:
-
RNA antisense purification
- hiCLIP:
-
RNA hybrid and individual-nucleotide resolution ultraviolet crosslinking and immunoprecipitation
- RPL:
-
RNA proximity ligation
- MARIO:
-
Map** RNA interactome in vivo
- PARIS:
-
Psoralen analysis of RNA interactions and structures
- SPLASH:
-
Sequencing of psoralen crosslinked, ligated, and selected hybrids
- LIGR-seq:
-
Ligation of interacting RNA followed by high-throughput sequencing
- COMRADES:
-
Crosslinking of matched RNAs and deep sequencing
- RIC-seq:
-
RNA in situ conformation sequencing
- RRI:
-
RNA–RNA interaction
- MNase:
-
Micrococcal nuclease
- pCp-biotin:
-
Biotinylated cytidine (bis) phosphate
- vRIC-seq:
-
Virion RIC-seq
- SHARC:
-
Spatial 2′-hydroxyl acylation reversible crosslinking
- ConA:
-
Concanavalin A
References
Adiconis X, Borges-Rivera D, Satija R et al (2013) Comparative analysis of RNA sequencing methods for degraded or low-input samples. Nat Methods 10:623–629
Aw JG, Shen Y, Wilm A et al (2016) In vivo map** of eukaryotic RNA interactomes reveals principles of higher-order organization and regulation. Mol Cell 62(4):603–617
Bak G, Han K, Kim K-S et al (2015) Electrophoretic mobility shift assay of RNA–RNA complexes. In: Schmidt FJ (ed) RNA-RNA interactions: methods and protocols. Springer New York, New York, NY, pp 153–163
Bartel DP (2004) MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 116:281–297
Cai Z, Cao C, Ji L et al (2020) RIC-seq for global in situ profiling of RNA-RNA spatial interactions. Nature 582:432–437
Cao C, Cai Z, **ao X et al (2021) The architecture of the SARS-CoV-2 RNA genome inside virion. Nat Commun 12:3917
Chen J, Xue Y (2016) Emerging roles of non-coding RNAs in epigenetic regulation. Sci China Life Sci 59:227–235
Djebali S, Davis CA, Merkel A et al (2012) Landscape of transcription in human cells. Nature 489:101–108
Engreitz JM, Sirokman K, McDonel P et al (2014) RNA-RNA interactions enable specific targeting of noncoding RNAs to nascent Pre-mRNAs and chromatin sites. Cell 159:188–199
Esteller M (2011) Non-coding RNAs in human disease. Nat Rev Genet 12:861–874
Gerstberger S, Hafner M, Tuschl T (2014) A census of human RNA-binding proteins. Nat Rev Genet 15:829–845
Gong C, Maquat LE (2011) lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470:284–288
Haas U, Sczakiel G, Laufer SD (2012) MicroRNA-mediated regulation of gene expression is affected by disease-associated SNPs within the 3′-UTR via altered RNA structure. RNA Biol 9:924–937
Halvorsen M, Martin JS, Broadaway S et al (2010) Disease-associated mutations that alter the RNA structural ensemble. PLoS Genet 6:e1001074
Hangauer MJ, Vaughn IW, McManus MT (2013) Pervasive transcription of the human genome produces thousands of previously unidentified long intergenic noncoding RNAs. PLoS Genet 9:e1003569
Hardin JW, Warnasooriya C, Kondo Y et al (2015) Assembly and dynamics of the U4/U6 di-snRNP by single-molecule FRET. Nucleic Acids Res 43:10963–10974
Helwak A, Tollervey D (2016) Identification of miRNA-target RNA interactions using CLASH. Methods Mol Biol 1358:229–251
Helwak A, Kudla G, Dudnakova T et al (2013) Map** the human miRNA interactome by CLASH reveals frequent noncanonical binding. Cell 153:654–665
Hentze MW, Castello A, Schwarzl T et al (2018) A brave new world of RNA-binding proteins. Nat Rev Mol Cell Biol 19:327–341
Kertesz M, Wan Y, Mazor E et al (2010) Genome-wide measurement of RNA secondary structure in yeast. Nature 467:103–107
Kortmann J, Narberhaus F (2012) Bacterial RNA thermometers: molecular zippers and switches. Nat Rev Microbiol 10:255–265
Kretz M, Siprashvili Z, Chu C et al (2013) Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493:231–235
Kudla G, Granneman S, Hahn D et al (2011) Cross-linking, ligation, and sequencing of hybrids reveals RNA–RNA interactions in yeast. Proc Natl Acad Sci USA 108:10010
Li P, Wei Y, Mei M et al (2018) Integrative analysis of Zika virus genome RNA structure reveals critical determinants of viral infectivity. Cell Host Microbe 24:875-886 e5
Lu Z, Zhang Qiangfeng C, Lee B et al (2016) RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 165:1267–1279
Moore MJ, Proudfoot NJ (2009) Pre-mRNA processing reaches back to transcription and ahead to translation. Cell 136:688–700
Mumbach MR, Rubin AJ, Flynn RA et al (2016) HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat Methods 13:919–922
Nguyen TC, Cao X, Yu P et al (2016) Map** RNA-RNA interactome and RNA structure in vivo by MARIO. Nat Commun 7:12023
Piganeau N, Schauer UE, Schroeder R (2006) A yeast RNA-hybrid system for the detection of RNA-RNA interactions in vivo. RNA 12:177–184
Ramani V, Qiu R, Shendure J (2015) High-throughput determination of RNA structure by proximity ligation. Nat Biotechnol 33:980–984
Rinn JL, Chang HY (2012) Genome regulation by long noncoding RNAs. Annu Rev Biochem 81:145–166
Salzman J, Gawad C, Wang PL et al (2012) Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7:e30733
Sharma E, Sterne-Weiler T, O’Hanlon D et al (2016) Global map** of human RNA-RNA interactions. Mol Cell 62:618–626
Shen EZ, Chen H, Ozturk AR et al (2018) Identification of piRNA binding sites reveals the argonaute regulatory landscape of the C. elegans germline. Cell 172:937–951 e18
Siegfried NA, Busan S, Rice GM et al (2014) RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat Methods 11:959–965
Song Y, Yang W, Fu Q et al (2020) irCLASH reveals RNA substrates recognized by human ADARs. Nat Struct Mol Biol 27:351–362
Stephenson JD, Kenyon JC, Symmons MF et al (2016) Characterizing 3D RNA structure by single molecule FRET. Methods 103:57–67
Sugimoto Y, Vigilante A, Darbo E et al (2015) hiCLIP reveals the in vivo atlas of mRNA secondary structures recognized by Staufen 1. Nature 519:491–594
Sun L, Li P, Ju X et al (2021) In vivo structural characterization of the SARS-CoV-2 RNA genome identifies host proteins vulnerable to repurposed drugs. Cell 184:1865-1883 e20
Van Damme R, Li K, Zhang M et al (2022) Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells. Nat Commun 13:911
Van Treeck B, Protter DSW, Matheny T et al (2018) RNA self-assembly contributes to stress granule formation and defining the stress granule transcriptome. Proc Natl Acad Sci USA 115:2734–2739
Wang X, Ramat A, Simonelig M et al (2022) Emerging roles and functional mechanisms of PIWI-interacting RNAs. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-022-00528-0
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–361
Waters SA, McAteer SP, Kudla G et al (2017) Small RNA interactome of pathogenic E. coli revealed through crosslinking of RNase E. The EMBO J 36:374–387
Watts JM, Dang KK, Gorelick RJ et al (2009) Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460:711–716
Xu B, Zhu Y, Cao C et al (2022) Recent advances in RNA structurome. Sci China Life Sci 65:1285–1324
Xue Y (2022) Architecture of RNA-RNA interactions. Curr Opin Genet Dev 72:138–144
Xue Y, Chen R, Qu L et al (2020) Noncoding RNA: from dark matter to bright star. Sci China Life Sci 63:463–468
Ye R, Hu N, Cao C et al (2022) CRIC-seq reveals positional rule of PTBP1-mediated long-range RNA loo** in splicing regulation. bioRxiv, 2022.08.09.503273
Zhao R, Rueda D (2009) RNA folding dynamics by single-molecule fluorescence resonance energy transfer. Methods 49:112–117
Ziv O, Gabryelska MM, Lun ATL et al (2018) COMRADES determines in vivo RNA structures and interactions. Nat Methods 15:785–788
Ziv O, Price J, Shalamova L et al (2020) The short- and long-range RNA-RNA interactome of SARS-CoV-2. Mol Cell 80:1067-1077 e5
Acknowledgements
We thank Drs. Changchang Cao and Di Wang for critical reading of this manuscript. This work was supported by the National Key R&D Program (2022YFA1303300), NSFC (32025008, 32130064, 91940306, and 81921003), and the K.C. Wong Education Foundation (GJTD-2020-06) to Y.X.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this chapter
Cite this chapter
Ye, R., Cai, Z., Xue, Y. (2023). Map** In Situ RNA–RNA Interactions with RIC-seq. In: Barciszewski, J. (eds) RNA Structure and Function. RNA Technologies, vol 14. Springer, Cham. https://doi.org/10.1007/978-3-031-36390-0_3
Download citation
DOI: https://doi.org/10.1007/978-3-031-36390-0_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-36389-4
Online ISBN: 978-3-031-36390-0
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)