Abstract
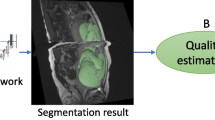
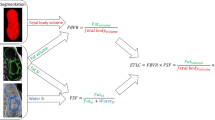
Normal fetal adipose tissue (AT) development is essential for perinatal well-being. AT, or simply fat, stores energy in the form of lipids. Malnourishment may result in excessive or depleted adiposity. Although previous studies showed a correlation between the amount of AT and perinatal outcome, prenatal assessment of AT is limited by lacking quantitative methods. Using magnetic resonance imaging (MRI), 3D fat- and water-only images of the entire fetus can be obtained from two-point Dixon images to enable AT lipid quantification. This paper is the first to present a methodology for develo** a deep learning (DL) based method for fetal fat segmentation based on Dixon MRI. It optimizes radiologists’ manual fetal fat delineation time to produce annotated training dataset. It consists of two steps: 1) model-based semi-automatic fetal fat segmentations, reviewed and corrected by a radiologist; 2) automatic fetal fat segmentation using DL networks trained on the resulting annotated dataset. Segmentation of 51 fetuses was performed with the semi-automatic method. Three DL networks were trained. We show a significant improvement in segmentation times (3:38 h \(\rightarrow \,{<}\)1 h) and observer variability (Dice of 0.738 \(\rightarrow \) 0.906) compared to manual segmentation. Automatic segmentation of 24 test cases with the 3D Residual U-Net, nn-UNet and SWIN-UNetR transformer networks yields a mean Dice score of 0.863, 0.787 and 0.856, respectively. These results are better than the manual observer variability, and comparable to automatic adult and pediatric fat segmentation. A Radiologist reviewed and corrected six new independent cases segmented using the best performing network (3D Residual U-Net), resulting in a Dice score of 0.961 and a significantly reduced correction time of 15:20 min. Using these novel segmentation methods and short MRI acquisition time, whole body subcutaneous lipids can be quantified for individual fetuses in the clinic and large-cohort research.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Banting, S.A., et al.: Estimation of neonatal body fat percentage predicts neonatal hypothermia better than birthweight centile. J. Matern.-Fetal Neonatal Med. 1–8 (2022)
Berger-Kulemann, V., et al.: Quantification of the subcutaneous fat layer with MRI in fetuses of healthy mothers with no underlying metabolic disease vs. fetuses of diabetic and obese mothers. J. Perinat. Med. (2012)
Blondiaux, E., et al.: Developmental patterns of fetal fat and corresponding signal on T1-weighted magnetic resonance imaging. Pediatr. Radiol. 48(3), 317–324 (2018)
Carberry, A.E., Raynes-Greenow, C.H., Turner, R.M., Askie, L.M., Jeffery, H.E.: Is body fat percentage a better measure of undernutrition in newborns than birth weight percentiles? Pediatr. Res. 74(6), 730–736 (2013)
Cassart, M., Garel, C.: European overview of current practice of fetal imaging by pediatric radiologists: a new task force is launched. Pediatr. Radiol. 50(12), 1794–1798 (2020)
Çiçek, Ö., Abdulkadir, A., Lienkamp, S.S., Brox, T., Ronneberger, O.: 3D U-Net: learning dense volumetric segmentation from sparse annotation. In: Ourselin, S., Joskowicz, L., Sabuncu, M.R., Unal, G., Wells, W. (eds.) MICCAI 2016. LNCS, vol. 9901, pp. 424–432. Springer, Cham (2016). https://doi.org/10.1007/978-3-319-46723-8_49
MONAI Consortium: MONAI: medical open network for AI (2022). https://doi.org/10.5281/zenodo.6639453
Dixon, W.T.: Simple proton spectroscopic imaging. Radiology 153(1), 189–194 (1984)
Dudovitch, G., Link-Sourani, D., Ben Sira, L., Miller, E., Ben Bashat, D., Joskowicz, L.: Deep learning automatic fetal structures segmentation in MRI scans with few annotated datasets. In: Martel, A.L., et al. (eds.) MICCAI 2020. LNCS, vol. 12266, pp. 365–374. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-59725-2_35
Estrada, S., et al.: FatSegNet: a fully automated deep learning pipeline for adipose tissue segmentation on abdominal Dixon MRI. Magn. Reson. Med. 83(4), 1471–1483 (2020)
Gardeil, F., Greene, R., Stuart, B., Turner, M.J.: Subcutaneous fat in the fetal abdomen as a predictor of growth restriction. Obstet. Gynecol. 94(2), 209–212 (1999)
Giza, S.A., et al.: Water-fat magnetic resonance imaging of adipose tissue compartments in the normal third trimester fetus. Pediatr. Radiol. 51(7), 1214–1222 (2021)
Hu, H.H., et al.: Linearity and bias of proton density fat fraction as a quantitative imaging biomarker: a multicenter, multiplatform, multivendor phantom study. Radiology 298(3), 640 (2021)
Isensee, F., Jaeger, P.F., Kohl, S.A., Petersen, J., Maier-Hein, K.H.: nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18(2), 203–211 (2021)
Joskowicz, L., Cohen, D., Caplan, N., Sosna, J.: Inter-observer variability of manual contour delineation of structures in CT. Eur. Radiol. 29(3), 1391–1399 (2019)
Kerfoot, E., Clough, J., Oksuz, I., Lee, J., King, A.P., Schnabel, J.A.: Left-ventricle quantification using residual U-Net. In: Pop, M., et al. (eds.) STACOM 2018. LNCS, vol. 11395, pp. 371–380. Springer, Cham (2019). https://doi.org/10.1007/978-3-030-12029-0_40
Kway, Y.M., et al.: Automated segmentation of visceral, deep subcutaneous, and superficial subcutaneous adipose tissue volumes in MRI of neonates and young children. Radiol. Artif. Intell. 3(5) (2021)
Larciprete, G., et al.: Intrauterine growth restriction and fetal body composition. Ultrasound Obstet. Gynecol. 26(3), 258–262 (2005)
Lee, W., et al.: New fetal weight estimation models using fractional limb volume. Ultrasound Obstet. Gynecol. 34(5), 556–565 (2009)
Lee, W., et al.: The fetal arm: individualized growth assessment in normal pregnancies. J. Ultrasound Med. 24(6), 817–828 (2005)
Lin, D., et al.: Automated measurement of pancreatic fat deposition on Dixon MRI using nnU-Net. J. Magn. Reson. Imaging (2022)
Mack, L.M., Kim, S.Y., Lee, S., Sangi-Haghpeykar, H., Lee, W.: A novel semiautomated fractional limb volume tool for rapid and reproducible fetal soft tissue assessment. J. Ultrasound Med. 35(7), 1573–1578 (2016)
Meshaka, R., Gaunt, T., Shelmerdine, S.C.: Artificial intelligence applied to fetal MRI: a sco** review of current research. Br. J. Radiol. 95, 20211205 (2022)
Roelants, J., et al.: Foetal fractional thigh volume: an early 3D ultrasound marker of neonatal adiposity. Pediatr. Obes. 12, 65–71 (2017)
Nobile de Santis, M., et al.: Growth of fetal lean mass and fetal fat mass in gestational diabetes. Ultrasound Obstet. Gynecol. 36(3), 328–337 (2010)
Shamshad, F., et al.: Transformers in medical imaging: a survey. ar**v preprint ar**v:2201.09873 (2022)
Shaw, M., Lutz, T., Gordon, A.: Does low body fat percentage in neonates greater than the 5th percentile birthweight increase the risk of hypoglycaemia and neonatal morbidity? J. Paediatr. Child Health 55(12), 1424–1428 (2019)
Tang, Y., et al.: Self-supervised pre-training of swin transformers for 3D medical image analysis. In: Proceedings of IEEE Computer Society Conference on Computer Vision and Pattern Recognition, pp. 20730–20740 (2022)
Torrents-Barrena, J., et al.: Segmentation and classification in MRI and us fetal imaging: recent trends and future prospects. Med. Image Anal. 51, 61–88 (2019)
Yushkevich, P.A., Gao, Y., Gerig, G.: ITK-SNAP: an interactive tool for semi-automatic segmentation of multi-modality biomedical images. In: International Conference of IEEE Engineering in Medicine and Biology Society (2016)
Acknowledgements
This research was supported by Kamin Grants [63418, 72126] from the Israel Innovation Authority.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2022 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Avisdris, N. et al. (2022). Automatic Fetal Fat Quantification from MRI. In: Licandro, R., Melbourne, A., Abaci Turk, E., Macgowan, C., Hutter, J. (eds) Perinatal, Preterm and Paediatric Image Analysis. PIPPI 2022. Lecture Notes in Computer Science, vol 13575. Springer, Cham. https://doi.org/10.1007/978-3-031-17117-8_3
Download citation
DOI: https://doi.org/10.1007/978-3-031-17117-8_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-17116-1
Online ISBN: 978-3-031-17117-8
eBook Packages: Computer ScienceComputer Science (R0)