Abstract
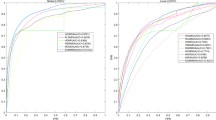
Taking into account the intrinsic high cost and time-consuming in traditional Vitro studies, a computational approach that can enable researchers to easily predict the potential miRNA-disease associations is imminently required. In this paper, we propose a computational method to predict potential associations between miRNAs and diseases via a new MeSH headings representation of diseases and eXtreme Gradient Boosting algorithm. Particularly, a novel MeSHHeading2vec method is first utilized to obtain a higher-quality MeSH heading representation of diseases, and then it is fused with miRNA functional similarity, disease semantic similarity and Gaussian interaction profile kernel similarity information to efficiently represent miRNA-disease pairs. Second, the deep auto-encoder neural network is adopted to extract the more representative feature subspace from the initial feature set. Finally, the eXtreme Gradient Boosting (XGBoost) algorithm is implemented for training and prediction. In the 5-fold cross-validation experiment, our method obtained average accuracy and AUC of 0.8668 and 0.9407, which performed better than many existing works.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Cheng, A.M., et al.: Antisense inhibition of human miRNAs and indications for an involvement of miRNA in cell growth and apoptosis. Nucleic Acids Res. 33(4), 1290–1297 (2005)
Alshalalfa, M., Alhajj, R.: Using context-specific effect of miRNAs to identify functional associations between miRNAs and gene signatures. BMC Bioinform. 14(S12), S1 (2013)
Xu, P., Guo, M., Hay, B.A.: MicroRNAs and the regulation of cell death. Trends Genet. 20(12), 617–624 (2004)
Griffiths‐Jones, S.: miRBase: microRNA sequences and annotation. Curr. Protoc. Bioinform. 29(1), 12.9.1-12.9.10 (2010)
Karp, X., Ambros, V.: Encountering microRNAs in cell fate signaling. Science 310(5752), 1288–1289 (2005)
Wang, R., et al.: MiR-185 is involved in human breast carcinogenesis by targeting Vegfa. FEBS Lett. 588(23), 4438–4447 (2014)
Ji, B.-Y., et al.: Predicting miRNA-disease association from heterogeneous information network with GraRep embedding model. Sci. Rep. 10(1), 1–12 (2020)
Guo, Z.-H., You, Z.-H., Wang, Y.-B., Huang, D.-S., Yi, H.-C., Chen, Z.-H.: Bioentity2vec: Attribute- and behavior-driven representation for predicting multi-type relationships between bioentities. GigaScience 9(6), giaa032 (2020)
Guo, Z.-H., et al.: A learning based framework for diverse biomolecule relationship prediction in molecular association network. Commun. Biol. 3(1), 1–9 (2020)
Chen, X., Zhang, D.-H., You, Z.-H.: A heterogeneous label propagation approach to explore the potential associations between miRNA and disease. J. Transl. Med. 16(1), 348 (2018)
Chen, X., et al.: BNPMDA: bipartite network projection for miRNA-disease association prediction. Bioinformatics 34(18), 3178–3186 (2018)
You, Z.-H., et al.: PBMDA: A novel and effective path-based computational model for miRNA-disease association prediction. PLoS Comput. Biol. 13(3), e1005455 (2017)
Zheng, K., et al.: MLMDA: a machine learning approach to predict and validate microRNA-disease associations by integrating of heterogenous information sources. J. Transl. Med. 17(1), 260 (2019)
Xu, J., et al.: Prioritizing candidate disease miRNAs by topological features in the miRNA target-dysregulated network: case study of prostate cancer. Mol. Cancer Ther. 10(10), 1857–1866 (2011)
Zhang, L., Chen, X., Yin, J.: Prediction of potential miRNA-disease associations through a novel unsupervised deep learning framework with variational autoencoder. Cells 8(9), 1040 (2019)
Ji, B.-Y., et al.: NEMPD: a network embedding-based method for predicting miRNA-disease associations by preserving behavior and attribute information. BMC Bioinform. 21(1), 1–17 (2020)
Ji, B.-Y., You, Z.-H., Wang, Y., Li, Z.-W., Wong, L.: DANE-MDA: predicting microRNA-disease associations via deep attributed network embedding. iScience 24(6), 102455 (2021)
Wang, L., et al.: LMTRDA: using logistic model tree to predict miRNA-disease associations by fusing multi-source information of sequences and similarities. PLOS Comput. Biol 15(3), e1006865 (2019)
Huang, Z., et al.: HMDD v3.0: a database for experimentally supported human microRNA–disease associations. Nucleic Acids Res. 47(D1), D1013–D1017 (2019)
Wang, D., et al.: Inferring the human microRNA functional similarity and functional network based on microRNA-associated diseases. Bioinformatics 26(13), 1644–1650 (2010)
van Laarhoven, T., Nabuurs, S.B., Marchiori, E.: Gaussian interaction profile kernels for predicting drug-target interaction. Bioinformatics 27(21), 3036–3043 (2011)
Guo, Z.-H., et al.: MeSHHeading2vec: a new method for representing MeSH headings as vectors based on graph embedding algorithm. Brief. Bioinform. (2020)
Wang, D., Cui, P., Zhu, W.: Structural deep network embedding. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (2016)
Tang, J., et al.: Line: large-scale information network embedding. In: Proceedings of the 24th International Conference on World Wide Web (2015)
Ou, M., et al.: Asymmetric transitivity preserving graph embedding. In: Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (2016)
Perozzi, B., Al-Rfou, R., Skiena, S.: Deepwalk: online learning of social representations. In: Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining (2014)
Belkin, M., Niyogi, P.: Laplacian eigenmaps and spectral techniques for embedding and clustering. Advances in Neural Information Processing Systems (2002)
Funding
This work is supported by the **njiang Natural Science Foundation under Grant 2017D01A78 and National Natural Science Foundation of China under Grant NO. 62002297.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Ji, BY., You, ZH., Wang, L., Wong, L., Su, XR., Zhao, BW. (2021). Predicting miRNA-Disease Associations via a New MeSH Headings Representation of Diseases and eXtreme Gradient Boosting. In: Huang, DS., Jo, KH., Li, J., Gribova, V., Premaratne, P. (eds) Intelligent Computing Theories and Application. ICIC 2021. Lecture Notes in Computer Science(), vol 12838. Springer, Cham. https://doi.org/10.1007/978-3-030-84532-2_5
Download citation
DOI: https://doi.org/10.1007/978-3-030-84532-2_5
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-84531-5
Online ISBN: 978-3-030-84532-2
eBook Packages: Computer ScienceComputer Science (R0)