Abstract
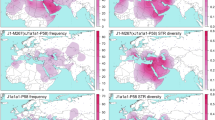
The patrilineal transmission of the Y chromosome and the fact that diversity in it accumulates along a strict genealogy imply that, by observing the current Y chromosome diversity in men, inferences can be made about the male-mediated history of humans (exactly like the female-mediated history is traced by mitochondrial DNA). By resequencing the non-recombining portion of the Y chromosome, it has been recently recognized that many branches of the Y chromosome genealogy (the so-called haplogroups) have expanded recently in bursts often tied to lifestyle changes or technological innovations. We have analysed two such bursting haplogroups: R1b-DF27 and E-M183. R1b-DF27 is prevalent in the Iberian Peninsula (40–70%), while its frequency drops to <15% north of the Pyrenees. We have estimated its age at ~4000 years ago, in line with ancient DNA discoveries and pointing to the population upheaval of the Bronze Age as a plausible agent in its origin and dispersion. Similarly, across the Strait of Gibraltar, E-M183 (equivalent to E-M81) dominates the Y chromosome landscape in NW Africa (up to 70%) and is much rarer elsewhere. However, its origins are more recent, at ~2000 years ago, and more difficult to pin down to a particular historical event. In conclusion, either in Iberia or the Maghreb, most men share a common ancestor that lived just a few hundred generations ago.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Adams SMM, Bosch E, Balaresque PLL et al (2008) The genetic legacy of religious diversity and intolerance: paternal lineages of Christians, Jews, and Muslims in the Iberian Peninsula. Am J Hum Genet 83:725–736. https://doi.org/10.1016/j.ajhg.2008.11.007
Airey J (2012) The politics of rape: sexual atrocity, propaganda wars, and the restoration stage. University of Delaware Press, Newark, NJ
Allentoft ME, Sikora M, Sjögren K-G et al (2015) Population genomics of bronze age Eurasia. Nature 522:167–172. https://doi.org/10.1038/nature14507
Alvarez-Cubero MJ, Santiago O, Martínez-Labarga C et al (2018) Methodology for Y chromosome capture: a complete genome sequence of Y chromosome using flow cytometry, laser microdissection and magnetic streptavidin-beads. Sci Rep 8:9436. https://doi.org/10.1038/s41598-018-27819-x
Arauna LR, Mendoza-Revilla J, Mas-Sandoval A et al (2017) Recent historical migrations have shaped the gene pool of Arabs and Berbers in North Africa. Mol Biol Evol 34:318–329. https://doi.org/10.1093/molbev/msw218
Arredi B, Poloni ES, Paracchini S et al (2004) A predominantly neolithic origin for Y-chromosomal DNA variation in North Africa. Am J Hum Genet 75:338–345
Auton A, Abecasis GR, Altshuler DM et al (2015) A global reference for human genetic variation. Nature 526:68–74. https://doi.org/10.1038/nature15393
Barbujani G, Magagni A, Minch E, Cavalli-Sforza LL (1997) An apportionment of human DNA diversity. Proc Natl Acad Sci U S A 94:4516–4519
Batini C, Hallast P, Zadik D et al (2015) Large-scale recent expansion of European patrilineages shown by population resequencing. Nat Commun 6:7152. https://doi.org/10.1038/ncomms8152
Bosch E, Calafell F, Comas D et al (2001) High-resolution analysis of human Y-chromosome variation shows a sharp discontinuity and limited gene flow between northwestern Africa and the Iberian Peninsula. Am J Hum Genet 68:1019–1029. https://doi.org/10.1086/319521
Bosch E, Calafell F, Pérez-Lezaun A et al (2000) Y chromosome STR haplotypes in four populations from northwest Africa. Int J Legal Med 114:36–40. https://doi.org/10.1007/s004140000136
Bosch E, Calafell F, Pérez-Lezaun A et al (1997) Population history of North Africa: evidence from classical genetic markers. Hum Biol 69:295–311
Bouckaert R, Heled J, Kühnert D et al (2014) BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput Biol 10:e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Burgarella C, Navascués M (2011) Mutation rate estimates for 110 Y-chromosome STRs combining population and father–son pair data. Eur J Hum Genet 19:70–75. https://doi.org/10.1038/ejhg.2010.154
Busby GBJ, Brisighelli F, Sanchez-Diz P et al (2012) The peopling of Europe and the cautionary tale of Y chromosome lineage R-M269. Proc R Soc B Biol Sci 279:884–892. https://doi.org/10.1098/rspb.2011.1044
Calafell F, Larmuseau MHD (2017) The Y chromosome as the most popular marker in genetic genealogy benefits interdisciplinary research. Hum Genet 136:559–573. https://doi.org/10.1007/s00439-016-1740-0
Cruciani F, Trombetta B, Antonelli C et al (2011) Strong intra- and inter-continental differentiation revealed by Y chromosome SNPs M269, U106 and U152. Forensic Sci Int Genet 5:e49–e52. https://doi.org/10.1016/j.fsigen.2010.07.006
Cruz-Dávalos DI, Nieves-Colón MA, Sockell A et al (2018) In-solution Y-chromosome capture-enrichment on ancient DNA libraries. BMC Genom 19:608. https://doi.org/10.1186/s12864-018-4945-x
de Filippo C, Bostoen K, Stoneking M, Pakendorf B (2012) Bringing together linguistic and genetic evidence to test the Bantu expansion. Proc R Soc B Biol Sci 279:3256–3263. https://doi.org/10.1098/rspb.2012.0318
Dupanloup I, Pereira L, Bertorelle G et al (2003) A recent shift from polygyny to monogamy in humans is suggested by the analysis of worldwide Y-chromosome diversity. J Mol Evol 57:85–97. https://doi.org/10.1007/s00239-003-2458-x
Fadhlaoui-Zid K, Haber M, Martínez-Cruz B et al (2013) Genome-wide and paternal diversity reveal a recent origin of human populations in North Africa. PLoS ONE 8:e80293. https://doi.org/10.1371/journal.pone.0080293
Fregel R, Gomes V, Gusmão L et al (2009) Demographic history of Canary Islands male gene-pool: replacement of native lineages by European. 14:1–14. https://doi.org/10.1186/1471-2148-9-181
Fregel R, Méndez FL, Bokbot Y et al (2017) Neolithization of North Africa involved the migration of people from both the Levant and Europe. 1–14
Fu Q, Li H, Moorjani P et al (2014) Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514:445–449. https://doi.org/10.1038/nature13810
Fu Q, Posth C, Hajdinjak M et al (2016) The genetic history of Ice Age Europe. Nature 534:200–205. https://doi.org/10.1038/nature17993
Gusmao L, Sanchez-Diz P, Calafell F et al (2005) Mutation rates at Y chromosome specific microsatellites. Hum Mutat 26:520–528. https://doi.org/10.1002/humu.20254
Haak W, Lazaridis I, Patterson N et al (2015) Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522:207–211. https://doi.org/10.1038/nature14317
Henn BM, Botigue LR, Gravel S et al (2012) Genomic ancestry of North Africans supports back-to-Africa migrations. PLoS Genet 8:e1002397
Jobling MA, Hollox EJ, Hurles ME et al (2014) Human evolutionary genetics, 2nd edn. Garland Science, New York and London
Karafet TM, Mendez FL, Meilerman MB et al (2008) New binary polymorphisms reshape and increase resolution of the human Y chromosomal haplogroup tree. Genome Res 18:830–838
Kayser M, Caglià A, Corach D et al (1997) Evaluation of Y-chromosomal STRs: a multicenter study. 110:125–133
Kohler TA, Smith ME, Bogaard A et al (2017) Greater post-Neolithic wealth disparities in Eurasia than in North America and Mesoamerica. Nature 551:619–622. https://doi.org/10.1038/nature24646
Kuderna LFK, Lizano E, Julià E et al (2019) Selective single molecule sequencing and assembly of a human Y chromosome of African origin. Nat Commun 10:4. https://doi.org/10.1038/s41467-018-07885-5
Larmuseau MHD, Calafell F, Princen SA et al (2018) The black legend on the Spanish presence in the low countries: verifying shared beliefs on genetic ancestry. Am J Phys Anthropol. https://doi.org/10.1002/ajpa.23409
Larmuseau MHD, Vanderheyden N, Van Geystelen A et al (2014) Recent radiation within Y-chromosomal haplogroup R-M269 resulted in high Y-STR haplotype resemblance. Ann Hum Genet 78:92–103. https://doi.org/10.1111/ahg.12050
McEvedy C, Jones R (1978) Atlas of world population history. Penguin, Harmondsworth (UK)
Mendizabal I, Sandoval K, Berniell-Lee G et al (2008) Genetic origin, admixture, and asymmetry in maternal and paternal human lineages in Cuba. BMC Evol Biol 8:213. https://doi.org/10.1186/1471-2148-8-213
Myres NM, Rootsi S, Lin AA et al (2011) A major Y-chromosome haplogroup R1b Holocene era founder effect in Central and Western Europe. Eur J Hum Genet 19:95–101. https://doi.org/10.1038/ejhg.2010.146
Parker G (1990a) The Dutch revolt: the revised version. Penguin Books
Parker G (1990b) Spain and the Netherlands, 1559–1659. Fontana Press, Waukegan, Il
Parker G (1973) Mutiny and discontent in the Spanish army of flanders. Past Present 58:38–52
Parker G, González de León F (1999) The grand strategy of Philip II and the revolt of the Netherlands. In: Van Nierop H, Venard M, Benedict P (eds) Reformation, revolt, and civil war in France and the Netherlands 1555–1585. Koninklijke Nederlandse Akademie van Wetenschappen, Amsterdam, pp 215–232
Poznik GD (2016) Identifying Y-chromosome haplogroups in arbitrarily large samples of sequenced or genotyped men. bioRxiv 088716. https://doi.org/10.1101/088716
Poznik GD, Xue Y, Mendez FL et al (2016) Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences. Nat Genet 48:593–599. https://doi.org/10.1038/ng.3559
Purps J, Siegert S, Willuweit S et al (2014) A global analysis of Y-chromosomal haplotype diversity for 23 STR loci. Forensic Sci Int Genet 12:12–23. https://doi.org/10.1016/j.fsigen.2014.04.008
Raghavan M, DeGiorgio M, Albrechtsen A et al (2014) The genetic prehistory of the New World Arctic. Science 345:1255832. https://doi.org/10.1126/science.1255832
Ralf A, Montiel González D, Zhong K, Kayser M (2018) Yleaf: software for human Y-chromosomal haplogroup inference from next generation sequencing data. Mol Biol Evol. https://doi.org/10.1093/molbev/msy032
Redd AJ, Agellon AB, Kearney VA et al (2002) Forensic value of 14 novel STRs on the human Y chromosome. 130:97–111
Rocca RA, Magoon G, Reynolds DF et al (2012) Discovery of Western European R1b1a2 Y chromosome variants in 1000 genomes project data: an online community approach. PLoS ONE 7:e41634
Rohde DLT, Olson S, Chang JT (2004) Modelling the recent common ancestry of all living humans. Nature 431:562–566. https://doi.org/10.1038/nature02842
Romualdi C, Balding D, Nasidze IS et al (2002) Patterns of human diversity, within and among continents, inferred from biallelic DNA polymorphisms. Genome Res 12:602–612
Seielstad MT, Minch E, Cavalli-Sforza LL (1998) Genetic evidence for a higher female migration rate in humans. 20:278–280
Soen V (2008) ¿Más allá de la leyenda negra? Léon van der Essen y la historiografía reciente en torno al castigo de las ciudades rebeldes en los Países Bajos (siglos XIV a XVI). In: Janssens G, Van der Essen L (eds) El Ejército Español en Flandes 1567–1584. Academia de Yuste, Yuste, pp 45–72
Solé-Morata N, Bertranpetit J, Comas D, Calafell F (2014a) Recent radiation of R-M269 and high Y-STR haplotype resemblance confirmed. Ann Hum Genet 78. https://doi.org/10.1111/ahg.12066
Solé-Morata N, Bertranpetit J, Comas D, Calafell F (2015) Y-chromosome diversity in Catalan surname samples: insights into surname origin and frequency. Eur J Hum Genet 23:1549–1557. https://doi.org/10.1038/ejhg.2015.14
Solé-Morata N, Bertranpetit J, Comas D, Calafell F (2014b) Recent radiation of R-M269 and high Y-STR haplotype resemblance confirmed. Ann Hum Genet 78:253–254. https://doi.org/10.1111/ahg.12066
Solé-Morata N, García-Fernández C, Urasin V et al (2017a) Whole Y-chromosome sequences reveal an extremely recent origin of the most common North African paternal lineage E-M183 (M81). Sci Rep 7:15941. https://doi.org/10.1038/s41598-017-16271-y
Solé-Morata N, Villaescusa P, García-Fernández C et al (2017b) Analysis of the R1b-DF27 haplogroup shows that a large fraction of Iberian Y-chromosome lineages originated recently in situ. Sci Rep 7. https://doi.org/10.1038/s41598-017-07710-x
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. https://doi.org/10.1093/bioinformatics/btu033
Swart K (1975) The black legend during the eighty years war. In: Bromley J, Kossmann E (eds) Britain and the Netherlands. Springer, The Hague, pp 36–57
Valverde L, Illescas MJ, Villaescusa P et al (2016) New clues to the evolutionary history of the main European paternal lineage M269: dissection of the Y-SNP S116 in Atlantic Europe and Iberia. Eur J Hum Genet 24:437–441. https://doi.org/10.1038/ejhg.2015.114
van Oven M, Van Geystelen A, Kayser M et al (2014) Seeing the wood for the trees: a minimal reference phylogeny for the human Y chromosome. Hum Mutat 35:187–191. https://doi.org/10.1002/humu.22468
Verdu P, Austerlitz F, Estoup A et al (2009) Origins and genetic diversity of pygmy hunter-gatherers from Western Central Africa. Curr Biol 19:312–318
Wei W, Ayub Q, Chen Y et al (2013) A calibrated human Y-chromosomal phylogeny based on resequencing. Genome Res 23:388–395
Wooding S, Ostler C, Prasad BV et al (2004) Directional migration in the Hindu castes: inferences from mitochondrial, autosomal and Y-chromosomal data. Hum Genet 115:221–229
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Nature Switzerland AG
About this chapter
Cite this chapter
García-Fernández, C., Calafell, F. (2019). The Parallel Lives of Human Y Chromosome Lineages Across the Strait of Gibraltar. In: Pontarotti, P. (eds) Evolution, Origin of Life, Concepts and Methods. Springer, Cham. https://doi.org/10.1007/978-3-030-30363-1_11
Download citation
DOI: https://doi.org/10.1007/978-3-030-30363-1_11
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-30362-4
Online ISBN: 978-3-030-30363-1
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)