Abstract
Background
Hepatocellular carcinoma (HCC) is a heterogeneous disease with recognized variability in virus infection, genetic features, and clinical outcome. To date, transcriptional profilings of HCC have been used to predict recurrence or survival/prognosis. However, there remains a challenge to identify specific genomic prints associated with HCC recurrence, which could lead to novel therapies or effective treatment. Here we examine the association between biological signals and intrahepatic recurrence using global gene expression profiles and powerful analytical methods.
Materials and Methods
Gene expression profiles were generated in 24 HCC patients with hepatitis B infections (B-type HCC) and 60 HCC patients with hepatitis C infections (C-type HCC). Gene set enrichment analysis (GSEA) was applied to the entire ranked gene lists related to early intrahepatic recurrence, based on “ideal discriminator method.”
Results
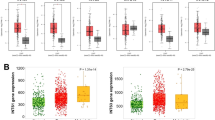
GSEA revealed Ribosomal Proteins as a common regulatory pathway in B-type (P < .001) and C-type (P = .003) HCC recurrence. In addition, Proteasome (P < .001) and Pentose Phosphate Pathway (P = .01) were identified as specific pathways in each type of HCC recurrence, respectively.
Conclusions
Understanding these biologically common and different mechanisms related to intrahepatic recurrence in B-type and C-type HCC could be useful in the development of new therapeutic strategies in our fight against HCC.



Similar content being viewed by others
References
Parkin DM. Global cancer statistics in the year 2000. Lancet Oncol. 2001;2:533–43.
Poon RT, Fan ST, Ng IO, Lo CM, Liu CL, Wong J. Different risk factors and prognosis for early and late intrahepatic recurrence after resection of hepatocellular carcinoma. Cancer. 2000;89:500–7.
Llovet JM, Burroughs A, Bruix J. Hepatocellular carcinoma. Lancet. 2003;362:1907–17.
Thiery JP. Epithelial-mesenchymal transitions in tumour progression. Nat Rev Cancer. 2002;2:442–54.
Piccart-Gebhart MJ, Procter M, Leyland-Jones B, Goldhirsch A, Untch M, Smith I, et al. Trastuzumab after adjuvant chemotherapy in HER2-positive breast cancer. N Engl J Med. 2005;353:1659–72.
Romond EH, Perez EA, Bryant J, Suman VJ, Geyer CE, Jr., Davidson NE, et al. Trastuzumab plus adjuvant chemotherapy for operable HER2-positive breast cancer. N Engl J Med. 2005;353:1673–84.
Ramaswamy S, Ross KN, Lander ES, Golub TR. A molecular signature of metastasis in primary solid tumors. Nat Genet. 2003;33:49–54.
Armstrong SA, Staunton JE, Silverman LB, Pieters R, den Boer ML, Minden MD, et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat Genet. 2002;30:41–7.
Iizuka N, Oka M, Yamada-Okabe H, Mori N, Tamesa T, Okada T, et al. Comparison of gene expression profiles between hepatitis B virus- and hepatitis C virus-infected hepatocellular carcinoma by oligonucleotide microarray data on the basis of a supervised learning method. Cancer Res. 2002;62:3939–44.
Iizuka N, Oka M, Yamada-Okabe H, Nishida M, Maeda Y, Mori N, et al. Oligonucleotide microarray for prediction of early intrahepatic recurrence of hepatocellular carcinoma after curative resection. Lancet. 2003;361:923–9.
Ye QH, Qin LX, Forgues M, He P, Kim JW, Peng AC, et al. Predicting hepatitis B virus-positive metastatic hepatocellular carcinomas using gene expression profiling and supervised machine learning. Nat Med. 2003;9:416–23.
Yoshioka S, Takemasa I, Nagano H, Kittaka N, Noda T, Wada H, et al. Molecular prediction of early recurrence after resection of hepatocellular carcinoma. Eur J Cancer. 2009;5:881–9.
Wang SM, Ooi LL, Hui KM. Identification and validation of a novel gene signature associated with the recurrence of human hepatocellular carcinoma. Clin Cancer Res. 2007;13:6275–83.
Woo HG, Park ES, Cheon JH, Kim JH, Lee JS, Park BJ, et al. Gene expression-based recurrence prediction of hepatitis B virus-related human hepatocellular carcinoma. Clin Cancer Res. 2008;14:2056–64.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Monti S, Savage KJ, Kutok JL, Feuerhake F, Kurtin P, Mihm M, et al. Molecular profiling of diffuse large B-cell lymphoma identifies robust subtypes including one characterized by host inflammatory response. Blood. 2005;105:1851–61.
Yang C, Zeisberg M, Lively JC, Nyberg P, Afdhal N, Kalluri R. Integrin alpha1beta1 and alpha2beta1 are the key regulators of hepatocarcinoma cell invasion across the fibrotic matrix microenvironment. Cancer Res. 2003;63:8312–7.
Takeno A, Takemasa I, Doki Y, Yamasaki M, Miyata H, Takiguchi S, et al. Integrative approach for differentially overexpressed genes in gastric cancer by combining large-scale gene expression profiling and network analysis. Br J Cancer. 2008;99:1307–15.
Kittaka N, Takemasa I, Takeda Y, Marubashi S, Nagano H, Umeshita K, et al. Molecular map** of human hepatocellular carcinoma provides deeper biological insight from genomic data. Eur J Cancer. 2008;44:885–97.
Troyanskaya OG, Garber ME, Brown PO, Botstein D, Altman RB. Nonparametric methods for identifying differentially expressed genes in microarray data. Bioinformatics. 2002;18:1454–61.
Veer LJ, Dai H, van de Vijver MJ, He YD, Hart AA, Mao M, et al. (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415: 530–6.
Golub TR, Slonim DK, Tamayo P, Huard C, Gaasenbeek M, Mesirov JP, et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science. 1999;286:531–7.
Dahlquist KD, Salomonis N, Vranizan K, Lawlor SC, Conklin BR. GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nat Genet. 2002;31:19–20.
Hu XT, Chen W, Wang D, Shi QL, Zhang FB, Liao YQ, et al. The proteasome subunit PSMA7 located on the 20q13 amplicon is overexpressed and associated with liver metastasis in colorectal cancer. Oncol Rep. 2008;19:441–6.
Langbein S, Zerilli M, Zur Hausen A, Staiger W, Rensch-Boschert K, Lukan N, et al. Expression of transketolase TKTL1 predicts colon and urothelial cancer patient survival: Warburg effect reinterpreted. Br J Cancer. 2006;94:578–85.
Langbein S, Frederiks WM, zur Hausen A, Popa J, Lehmann J, Weiss C, et al. Metastasis is promoted by a bioenergetic switch: new targets for progressive renal cell cancer. Int J Cancer. 2008;122:2422–8.
Acknowledgment
The authors thank Kenichi Matsubara, Ph.D. (President and Executive Director of The DNA Chip Research Inc., Yokohama Japan) for contract services on AceGene Human 30K.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kittaka, N., Takemasa, I., Seno, S. et al. Exploration of Potential Genomic Portraits Associated with Intrahepatic Recurrence in Human Hepatocellular Carcinoma. Ann Surg Oncol 17, 3145–3154 (2010). https://doi.org/10.1245/s10434-010-1150-9
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1245/s10434-010-1150-9