Abstract
The CCCH proteins play important roles in plant growth and development, hormone response, pathogen defense and abiotic stress tolerance. However, the knowledge of their roles in thermotolerance are scarce. Here, we identified a heat-inducible CCCH gene LlC3H18 from lily. LlC3H18 was localized in the cytoplasm and nucleus under normal conditions, while it translocated in the cytoplasmic foci and co-located with the markers of two messenger ribonucleoprotein (mRNP) granules, processing bodies (PBs) and stress granules (SGs) under heat stress conditions, and it also exhibited RNA-binding ability. In addition, LlC3H18 exhibited transactivation activity in both yeast and plant cells. In lily and Arabidopsis, overexpression of LlC3H18 damaged their thermotolerances, and silencing of LlC3H18 in lily also impaired its thermotolerance. Similarly, Arabidopsis atc3h18 mutant also showed decreased thermotolerance. These results indicated that the appropriate expression of C3H18 was crucial for establishing thermotolerance. Further analysis found that LlC3H18 directly bound to the promoter of LlWRKY33 and activated its expression. Besides, it was found that LlMYB305 acted as an upstream factor of LlC3H18 and activated its expression. In conclusion, we demonstrated that there may be a LlMYB305-LlC3H18-LlWRKY33 regulatory module in lily that is involved in the establishment of thermotolerance and finely regulates heat stress response.
Graphical Abstract

Similar content being viewed by others
Core
Lily heat-inducible CCCH gene LlC3H18 is directly activated by LlMYB305, and its protein is partially localized in the nucleus to act as a transcription activator of LlWRKY33, thus forming a LlMYB305-LlC3H18-LlWRKY33 regulatory module. LlC3H18 can also locate in the cytoplasm foci under high temperature conditions, and play a role of RNA-binding protein to form mRNP granules for finely regulating heat stress response.
Gene and accession numbers
Sequence data from this article can be found in the database of the National Center for Biotechnology Information (NCBI) under the accession numbers: LlC3H18 (OR094243), LlMYB305 (MW383251), LlWRKY33 (OR094247).
Introduction
With the development of industry, a large amount of fossil energy is used, the emission of carbon dioxide increases year by year, the trend of global warming is inevitable, and more and more abnormally high temperature weather occurs frequently (Grover et al. 2013; Wahid et al. 2007). As sessile organisms, plants are difficult to escape the adverse effects of environmental changes and are extremely sensitive to temperature changes, especially for some horticultural crops and food crops; high temperature often leads to reduced yield and quality (Teixeira et al. 2013). Many studies have proved that transcription factors (TFs) play an important role in the regulation of plant thermotolerance (Ohama et al. 2017). It may be feasible to improve thermotolerance in crops by screening heat-resistant regulators and genetic engineering methods.
The CCCH proteins are a class of proteins with zinc finger (ZNF) domains, many of which can function as TFs to regulate the expression of target genes (Pomeranz et al. 2011; Wang et al. 2008). The CCCH proteins contain 1–6 CCCH-type ZNF motifs which is consist of three Cys residues and one His residue (C-X5-14-C-X4-5-C-X3-H)(Wang et al. 2008). In Arabidopsis, there are 68 CCCH members and they are classified into 11 subfamilies based on the spacing between Cys and His in the ZNF motifs as well as the number of ZNF motifs (Wang et al. 2008). In addition, CCCH proteins are divided into tandem CCCH-type zinc finger (TZF) and non-TZF proteins: TZF proteins contain two tandem CCCH-type ZNF motifs whereas non-TZF proteins have fewer or greater than two CCCH-type ZNF motifs (Bogamuwa and Jang 2014; Seok et al. 2018). Except the ZNF domain, some members of CCCH family also had a LOTUS/OST-HTH (Limkain, Oskar, and TUdor-containing proteins 5 and 7) domain and an RRM (RNA-recognition motif) domain, both are putative RNA-binding domains, so they are always putative RNA-binding proteins for post-transcriptional regulation (Pomeranz et al. 2010b; Xu et al. 2015). In plants, CCCH proteins are a kind of regulators playing significant roles in plant growth, development, hormone response, defense pathogens, and resist to abiotic stresses (Bogamuwa and Jang 2014; Han et al. 2021). AtTZF2/AtOZF1, AtTZF3/AtOZF2, and cotton GhZFP1 are all associated with jasmonic acid-induced leaf senescence (Guo et al. 2009; Lee et al. 2012). AtTZF4, AtTZF5, and AtTZF6 positively regulate abscisic acid (ABA) response and play roles in seed germination and embryo formation, and AtC3H17 has pleiotropic effects on vegetative development, flowering, and seed development in Arabidopsis (Bogamuwa and Jang 2013; Seok et al. 2016). AtC3H14 and AtC3H15 are involved in the regulation of cell elongation, secondary wall thickening, male fertility, anther development, and acquisition of immunity against pathogens (Chai et al. 2015; Wang et al. 2022a, 2020). Poplar PdC3H17 and PdC3H18 positively regulate secondary wall formation in poplar (Chai et al. 2014). AtC3H11 is a subunit of polyadenylation factor and is required for Pseudomonas resistance (Bruggeman et al. 2014). Pepper CaC3H14 positively regulates the response of inoculation by Ralstonia solanacearum (Qiu et al. 2018). OsLIC promotes downstream OsWRKY30 for rice resistance to bacterial blight and leaf streak (Wang et al. 2022b). In addition, a number of CCCH proteins, such as AtTZF1, GhZFP1, GhTZF1, OsTZF1, AtSZF1/2, OsC3H47, PvC3H72, and OsDOS were found as important regulators for plant responses to salt, drought, cold, and oxidative stresses (Guo et al. 2009; Jan et al. 2013; Kong et al. 2006; Lin et al. 2011; Sun et al. 2015; Jan et al. 2013; Pomeranz et al. 2010a). Two non-TNF proteins of Arabidopsis, AtC3H18L and AtC3H18 can also locate in cytoplasmic foci after HS (Xu et al. 2015). AtWRKY33 has been reported to be involved in the regulation of plant pathogen defense, and salt, flooding, and heat tolerances, which is a central regulator of these physiological processes (Jiang and Deyholos 2008; Krishnamurthy et al. 2020; Li et al. 2011; Liu et al. 2021; Liu et al. 2015; Zheng et al. 2006). AtC3H14 shows an ability to bind DNA element and acts as a direct regulator of AtWRKY33 to activate its expression and participate in the establishment of resistance to B. cinerea (Wang et al. 2020). Similarly, we found that LlC3H18 bound to the DNA element from LlWRKY33 promoter and activated its expression (Fig. 6). Overexpression of LlC3H18 activated the expression of WRKY33 in lily and Arabidopsis, and silencing of LlC3H18 decreased the expression of LlWRKY33 (Fig. 6). These results suggested that LlC3H18 might act as a direct activator of LlWRKY33.
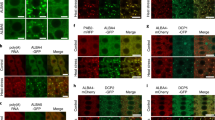
At room temperature, AtC3H18 could form mRNP granules in pollens and localize to cytoplasmic foci, but no similar phenomenon was observed in tobacco cells. However, under HS conditions, AtC3H18 can form mRNP granules in tobacco cells and localize to cytoplasmic foci, indicating the critical concentration of AtC3H18 forming mRNP granules in pollens is lower than that in tobacco cells (Xu et al. 2022). The localization of LlC3H18 also had a similar phenomenon; the localization of cytoplasmic foci of LlC3H18 was enabled by high temperature (Fig. 2A). In Chinese cabbage, the C3H18 homologous genes BcMF30a and BcMF30c play an indispensable role in pollen fertility; overexpression or mutation of them leads to abnormal pollen development, indicating that proper expression of BcMF30a and BcMF30c is extremely important for normal pollen development (Xu et al. 2020b; Xu et al. 2020c). Similarly, AtC3H18-overexpressed Arabidopsis lines also show pollen abortion, which may be caused by affecting the assembly of mRNP granules (Xu et al. 2022). Our study found that overexpression of LlC3H18 in lily and Arabidopsis would lead to the reduction of thermotolerance (Fig. 5). In LlC3H18-overexpressing plants, it was found that the expression of heat-responsive genes was activated at room temperature, indicating that LlC3H18 could act as a TF and activate HS response in the absence of high temperature; however, under HS, the induced expression of heat-responsive genes decreased, which might lead to a final decrease in thermotolerance (Fig. S4). We speculated that the excessive expression of LlC3H18 resulted in the accumulation of LlC3H18 protein, which might destroy the normal assembly process of mRNP granules under HS conditions for leading to the damage of HS response and reducing the expression of heat-protective genes; the specific mechanism need to be clarified in the future. Interesting, silencing of LlC3H18 in lily also led to the decrease of its thermotolerance (Fig. 5). Similarly, the atc3h18 Arabidopsis mutant showed decreased thermotolerance as well. We found that the heat-induced expression of heat-responsive genes was reduced in atc3h18 mutant (Fig. S6). After HS release, C3H18 was released from cytoplasmic foci and entered the nucleus to play a role as a trans-activator. The decrease or deletion of C3H18 expression may disrupt the role of trans-activator of C3H18, which is detrimental to the maintenance of HS response. These results indicated that the appropriate expression of C3H18 was crucial for establishing thermotolerance.
Meanwhile, many studies have also found that transgenic plants with overexpression of TZFs often show growth defects. For instance, the overexpression transgenic plants of AtTZF1, 4, 5, and 6 all exhibited compact and crinkled leaves, and some of homozygous overexpression plants of AtTZF1 even showed lethal phenotype (Bogamuwa and Jang 2013; Lin et al. 2011). Overexpression of AtC3H14 and AtC3H15 led to dwarfing phenotypes and male sterility in Arabidopsis, respectively (Kim et al. 2014; Shi et al. 2015). We found that overexpression of LlC3H18 also caused the growth defects of transgenic Arabidopsis plants (Fig. 4), which was very different with the growth phenotype of AtC3H18-overexpression, which did not cause any growth defects under normal conditions (Xu et al. 2022). It was speculated that LlC3H18 also showed nucleus-localization under normal conditions, which may activate some target genes to damage the growth of transgenic plants.
Arabidopsis AtC3H18 is highly expressed in anthers, and the R2R3-MYB TFs AtMYB21 and AtMYB24 function key roles in anther development and thermotolerance (Cheng et al. 2009; Huang et al. 2017, 2020; Kumar and Chattopadhyay 2018; Mandaokar and Browse 2009; Song et al. 2011; Xu et al. 2022). Lily LlMYB305 is a MYB21/24 homology, and our previous study demonstrated that LlMYB305 is induced by high temperature (Wu et al. 2021). In this study, we demonstrated that LlMYB305 directly bound to the promoter of LlC3H18 and activated its expression (Fig. 7). At the same time, we found that overexpression of LlMYB305 improved the thermotolerance of lily, silencing of LlMYB305 reduced its thermotolerance (Fig. 7); and LlMYB305 could activate the expression of LlC3H18, which did not damage thermotolerance as overexpression of LlC3H18 (Fig. 5). It was speculated that LlMYB305, as an upstream regulatory factor, could coordinate the role of LlC3H18 by simultaneously activating other factors, thereby ensuring that LlC3H18 played a role within an appropriate range. In poplar, PdMYB3 and PdMYB21 regulate the specific expression of PdC3H17 and PdC3H18 to participate in the formation of secondary cell walls (Chai et al. 2014), suggesting that MYB TFs and CCCH-type proteins may link with a conserved regulatory mechanism.
In conclusion, our study showed that LlC3H18 was a heat-inducible CCCH-type protein, and LlMYB305 could act as its upstream factor to activate its expression and participate in the establishment of thermotolerance; without HS, LlC3H18 could localize in the nucleus and acted as a trans-activator to stimulate the expression of LlWRKY33; in addition, under HS conditions, LlC3H18 could also play a role of RNA-binding protein, form mRNP granules to participate in the regulation of thermotolerance (Fig. 8). Based on these results, we speculate that there may be a LlMYB305-LlC3H18-LlWRKY33 regulatory module involved in the establishment of thermotolerance in lily.
A simple working model of the LlC3H18-mediated regulatory mechanism in lily in response to heat stress. A Under HS conditions, LlC3H18 is a heat-inducible CCCH gene, which can be directly activated by LlMYB305; LlC3H18 locates in the cytoplasm foci and acts as RNA binding protein to form mRNP granules, thus balancing the thermotolerance. B At the recovery period without HS, LlC3H18 can be transformed from cytoplasm foci to localize in the nucleus, which promotes it to act as a trans-activator, directly activating the expression of LlWRKY33, thus forming a heat-inducible LlMYB305-LlC3H18-LlWRKY33 regulatory module, and sustaining the heat stress response. The gray arrow indicates a closed state, while the yellow arrow indicates a working state. HS Heat stress, PB Processing body, SG Stress granule
Methods
Plant materials and growth conditions
The Lilium Oriental hybrid ‘Sorbonne’ and L. longiflorum ‘White heaven’ were used as the experimental materials in this study. The ‘Sorbonne’ was planted in soil and grown in a greenhouse with day/night temperatures of 22/16 °C. Sterile tissue-cultured plantlets of ‘White heaven’ were cultured in a standard culture room at 22 °C with a light–dark cycle of 16 h/8 h. Arabidopsis thaliana Col-0 and Nicotiana benthamiana (tobacco) seeds were sown in MS medium, and 10 days after germination, the seedlings were transplanted into nursery pots and grown in a standard greenhouse (22/16 °C) with 16-h/8-h light/dark photoperiod.
Isolation of LlC3H18 gene, LlC3H18 and LlWRKY33 promoters from ‘White heaven’
Two-week-old lily plants were treated with HS at 37 °C for 1 h, the leaves were collected, and total RNA was extracted using RNAprep Pure Kit (Tiangen, China), followed by M-MLV reverse transcriptase (Vazyme, China) and Oligo dT primer to synthesize cDNA. According to the transcriptome data, the specific primers were designed to amplify the open reading frame (ORF) of LlC3H18 (Table S2). The promoters of LlC3H18 and LlWRKY33 were isolated and cloned with the method of Hi-tail PCR (Liu and Chen 2007) from lily ‘White heaven’; the 1184-bp upstream fragment from ATG of LlC3H18, and the 521-bp upstream fragment from ATG of LlWRKY33 were isolated and identified, respectively.
Phylogenetic tree analysis and prediction of conserved protein domains
Phylogenetic tree analysis of LlC3H18 and its homologous proteins was performed by MEGA 7.0 software using the neighbor-joining method (n = 1000). Multiple alignment analysis of C3H18 amino acid sequences from different species was performed using ClustalW 2.0 and BioEdit 7.0 softwares.
Promoter activity analysis of LlC3H18
The LlC3H18 promoter was cloned into pGreenII0800-LUC (Hellens et al. 2005) and pCAMBIA1391-GUS (Abcam, USA). The reconstructed pGreenII0800-LUC-proLlC3H18 was introduced into Agrobacterium tumefaciens strain GV3101 (psoup). A mixed bacterial solution was infiltrated into tobacco leaves for the activity assay. After 48 h, the infiltrated leaves were treated with HS at 37 °C for 3 h, then they were removed to detect the LUC signal. The reconstructed pCAMBIA1391-GUS-proLlC3H18 was introduced into A. tumefaciens strain GV3101. The GUS-reporter gene was stably transformed into Arabidopsis and transiently transformed into lily petal discs. The transgenic Arabidopsis seedlings and the lily petal discs were treated with HS at 37 °C for 3 h, and then, they were sampled for GUS assay. All primers used for plasmid construct are listed in the Table S3.
The transcriptional activity assay of LlC3H18
The ORF of LlC3H18 was cloned into pGBKT7 (BD; Clontech, Japan) to generate BD-LlC3H18 protein. The plasmid-transformed yeast AH109 cell was used for transcriptional activity analysis; yeast containing GAL4 and BD were used as positive and negative controls, respectively. After 3 days of culture at 30 °C, the positive clones were selected and transferred to -Trp-His-deficient SD medium for identification of transcriptional activity. The ORF of LlC3H18 was cloned into the vector pEAQ (Sainsbury et al. 2009) to generate a BD fusion protein as effector. The 5 × GAL4 UAS element and the mini 35S promoter were fused and cloned into the vector pGreenII0800-LUC to construct the reporter vector. These reconstituted vectors were respectively introduced into A. tumefaciens GV3101 (psoup). The bacterial solution was resuspended, mixed according to the proportion, and then the tobacco leaves were injected. Under normal growth conditions, after culturing for 60 h, the injected leaves were cut to detect the LUC signal and the LUC intensity was also determined.
Subcellular localization analysis of LlC3H18
The ORF of LlC3H18 was cloned into pCAMBIA1300-N-GFP (Abcam, USA) vector to generate GFP-LlC3H18 fusion protein, and the reconstituted vector were introduced into A. tumefaciens GV3101, respectively. The different constructs were expressed in tobacco leaves, the mCherry-DCP2, mCherry-PABP8, and RFP-NLS were used as the PB, SG, and nucleus marker, respectively, and fluorescence signals were checked under confocal microscopy (Zeiss, Jena, Germany).
Heat treatment and gene expression analysis of lily
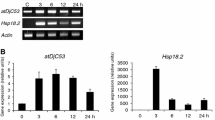
The robust tissue-cultured ‘White heaven’ seedlings with the same size were selected for gene expression analysis. For HS, lily plantlets were incubated at 37 °C for different lengths of time (0, 0.5, 1, 3, 6, 12 h). After HS finished, the leaves were collected for extracting total RNA. The RNA reverse transcribed into cDNA with a HiScript II kit (Vazyme, China), and the expression of LlC3H18 were detected by real-time quantitative PCR (RT-qPCR) with the 2−ΔΔCT method (Livak and Schmittgen 2001; Schmittgen and Livak 2008). Lily 18S rRNA was used as an internal reference gene (Table S4).
Yeast one-hybrid assay
The test and mutant fragments of the promoters of LlC3H18 and LlWRKY33 were cloned into pLacZi (Clontech, Japan) vectors. The ORFs of LlMYB305 and LlC3H18 were inserted into a pJG (Clontech, Japan) vector, respectively. The corresponding vectors were co-transformed into yeast EGY48. Successful transformants were selected by growing on -Trp-Ura deficit SD media for 3 days at 30℃. Binding was investigated using color analysis on SD media containing 80 mg L−1 x-gal.
Electrophoretic mobility shift assay (EMSA)
The ORFs of LlC3H18 and LlMYB305 were separately cloned into pGEX-4 T-1 (GE Healthcare, USA) to generate the GST fusion proteins. The fusion proteins were induced in E. coli BL21 by adding isopropyl-β-D-1-thiogalactopyranoside (200 mM, IPTG); The recombinant GST-LlC3H18 and GST-LlMYB305 proteins was purified by GST protein purification kit, and detected by SDS–polyacrylamide electrophoresis. The EMSA probe was synthesized with 5’ biotin-labeled. Binding reactions were incubated at room temperature for 20 min and then separated using electrophoresis through a 6% (40:1 acrylamide:bis-acrylamide) native gel at 4 °C. The EMSA analysis was performed using the Light Shift Chemiluminescence EMSA Kit (ThermoFisher, New York, USA).
Dual-luciferase reporter assay
The ORFs of LlC3H18 and LlMYB305 were cloned into pGreenII62-SK (SK-II) (Hellens et al. 2005) to generate effector vectors, respectively. The LlC3H18 and LlWRKY33 promoters were cloned into pGreenII0800-LUC to generate the reporter vectors. The empty vectors were used as the negative control. The vectors were introduced into A. tumefaciens strain GV3101 (psoup). A mixed bacterial solution was infiltrated into tobacco leaves for the dual-luciferase reporter assay. After 48 h, the infiltrated leaves were removed, and the LUC signal was detected and measured.
ARE-binding assay
For DNA constructs used in mRNA-binding assays, GFP-ARE was made by ligating the ARE (5’-TTATTTATTATTTATTTATTATTTATTTATTATTTATTTATTA-3’) to the end of the GFP coding region, subcloning to pCAMBIA1300-GFP vector. GFP-MutG was generated following the same procedures except the ARE region was replaced by the MutG (5’-TTGTTTGTTGTTTGTTTGTTGTTTGTTTGTTGTTTGTTTGTTA-3’). The LlC3H18 was generated by PCR and then transferring it in pGreenII62-SK (Hellens et al. 2005) vector. These reconstituted vectors were introduced into A. tumefaciens GV3101, respectively. The mixed bacterial solutions were suspended and injected in tobacco leaves. After 48 h, the leaves were removed and the intensity of GFP fluorescence was observed.
Transient overexpression in lily petals
The bacterial solutions of pGreenII62-SK and pGreenII62-SK-LlC3H18/LlWRKY33/LlMYB305 were collected by centrifugation and resuspended in the infiltration buffer (10 mM MgCl2, 200 mM acetosyringine, 10 mM MES, pH 5.6) and placed in the dark at 22℃ for 5 h before vacuum infiltration. The 10-cm length of unopened ‘Sorbonne’ flower buds was selected, and the inner petals were used to obscure the 1-cm-diameter discs with a hole-puncher (Wu et al. 2022a; Wu et al. 2023; Wu et al. 2022b). Under the vacuum condition of -0.7 MPa, the bacterial solution was infiltrated into these petal discs. Then, the discs were washed with sterile water and placed on an agar plate (0.4%), and cultivated in the dark at 22 °C for 96 h. For HS treatment, the discs were treated at 40 °C for 12 h, then harvested immediately, and their relative ion leakage determined (Wu et al. 2023).
Virus-induced gene silencing (VIGS) in lily petals
A 300-bp fragment of LlC3H18 or LlMYB305 was obtained by PCR amplification and cloned into pTRV2 to generate the pTRV2-LlC3H18 and pTRV2-LlMYB305 vector, respectively. Then, pTRV1, pTRV2, and pTRV2-LlC3H18 or pTRV2-LlMYB305 were transformed into A. tumefaciens GV3101, respectively. The bacteria of TRV1 and TRV2 were resuspended and mixed in proportion, and the petal discs were vacuumed as described above. After 5 days, the discs were treated with HS, then the discs were harvested to measure relative ion leakage.
Stable transformation of Arabidopsis
The LlC3H18 ORF was inserted into pCAMBIA1300, the LlC3H18 promoter was inserted into pCAMBIA1391, and then the reconstructed vectors were transformed into A. tumefaciens GV3101, respectively. For transformation, the floral-dip method was used with 5-week-old Arabidopsis plants. Through resistance screening and RT-PCR identification, three transgenic lines were selected for subsequent experiments.
Thermotolerance test of transgenic plants
Arabidopsis seeds were sterilized and sown on MS medium, vernalized for 3 days at 4 °C in the dark, and then transferred to a standard culture room at 22 °C (16 h light/8 h dark). The 5-day-old Arabidopsis seedlings were treated with HS in a constant temperature incubator, and recovery at 22 °C for 7 days, and the survival rate was recorded. The 5-day-old wild-type and transgenic Arabidopsis seedlings were collected for expression analysis of heat-responsive genes. Arabidopsis AtActin2 was used as internal reference gene for RT-qPCR analysis, and the relative levels of gene expression were calculated using the 2−ΔΔCT method (Livak and Schmittgen 2001; Schmittgen and Livak 2008).
Availability of data and materials
The authors confirm that all data in this study are included in this published article (and its supplementary information file).
Abbreviations
- ARE:
-
AU-rich element
- CCCH:
-
Three conserved cysteine residues and one histidine residue
- EMSA:
-
Electrophoretic mobility shift assay
- GFP:
-
Green fluorescent protein
- HS:
-
Heat stress
- LUC:
-
Luciferase
- mRNP:
-
Messenger ribonucleoprotein
- OE:
-
Overexpression
- ORF:
-
Open reading frame
- PB:
-
Processing body
- PCR:
-
Polymerase chain reaction
- SG:
-
Stress granule
- TF:
-
Transcription factor
- TRV:
-
Tobacco rattle virus
- TZF:
-
CCCH-type zinc finger
- VIGS:
-
Virus-induced gene silencing
- Y1H:
-
Yeast one-hybrid
References
Addepalli B, Hunt AG. Ribonuclease activity is a common property of Arabidopsis CCCH-containing zinc-finger proteins. FEBS Lett. 2008;582(17):2577–82.
Birkenbihl RP, Diezel C, Somssich IE. Arabidopsis WRKY33 is a key transcriptional regulator of hormonal and metabolic responses toward Botrytis cinerea infection. Plant Physiol. 2012;159(1):266–85.
Bogamuwa S, Jang JC. The Arabidopsis tandem CCCH zinc finger proteins AtTZF4, 5 and 6 are involved in light-, abscisic acid- and gibberellic acid-mediated regulation of seed germination. Plant Cell Environ. 2013;36(8):1507–19.
Bogamuwa SP, Jang JC. Tandem CCCH zinc finger proteins in plant growth, development and stress response. Plant Cell Physiol. 2014;55(8):1367–75.
Brewer BY, Malicka J, Blackshear PJ, Wilson GM. RNA sequence elements required for high affinity binding by the zinc finger domain of tristetraprolin: conformational changes coupled to the bipartite nature of Au-rich MRNA-destabilizing motifs. J Biol Chem. 2004;279(27):27870–7.
Bruggeman Q, Garmier M, de Bont L, Soubigou-Taconnat L, Mazubert C, Benhamed M, Raynaud C, Bergounioux C, Delarue M. The Polyadenylation Factor Subunit CLEAVAGE AND POLYADENYLATION SPECIFICITY FACTOR30: A Key Factor of Programmed Cell Death and a Regulator of Immunity in Arabidopsis. Plant Physiol. 2014;165(2):732–46.
Chai G, Kong Y, Zhu M, Yu L, Qi G, Tang X, Wang Z, Cao Y, Yu C, Zhou G. Arabidopsis C3H14 and C3H15 have overlap** roles in the regulation of secondary wall thickening and anther development. J Exp Bot. 2015;66(9):2595–609.
Chai G, Qi G, Cao Y, Wang Z, Yu L, Tang X, Yu Y, Wang D, Kong Y, Zhou G. Poplar PdC3H17 and PdC3H18 are direct targets of PdMYB3 and PdMYB21, and positively regulate secondary wall formation in Arabidopsis and poplar. New Phytol. 2014;203(2):520–34.
Cheng H, Song S, **ao L, Soo HM, Cheng Z, **e D, Peng J. Gibberellin acts through jasmonate to control the expression of MYB21, MYB24, and MYB57 to promote stamen filament growth in Arabidopsis. PLoS Genet. 2009;5(3): e1000440.
Decker CJ, Parker R. P-bodies and stress granules: possible roles in the control of translation and mRNA degradation. Cold Spring Harb Perspect Biol. 2012;4(9):a012286.
Deng H, Liu H, Li X, **ao J, Wang S. A CCCH-type zinc finger nucleic acid-binding protein quantitatively confers resistance against rice bacterial blight disease. Plant Physiol. 2012;158(2):876–89.
Grover A, Mittal D, Negi M, Lavania D. Generating high temperature tolerant transgenic plants: Achievements and challenges. Plant Sci. 2013;205–206:38–47.
Guo YH, Yu YP, Wang D, Wu CA, Yang GD, Huang JG, Zheng CC. GhZFP1, a novel CCCH-type zinc finger protein from cotton, enhances salt stress tolerance and fungal disease resistance in transgenic tobacco by interacting with GZIRD21A and GZIPR5. New Phytol. 2009;183(1):62–75.
Han G, Qiao Z, Li Y, Wang C, Wang B. The Roles of CCCH Zinc-Finger Proteins in Plant Abiotic Stress Tolerance. Int J Mol Sci. 2021;22(15):8327.
Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, Karunairetnam S, Gleave AP, Laing WA. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods. 2005;1(1):1–14.
Huang H, Gao H, Liu B, Qi T, Tong J, **ao L, **e D, Song S. Arabidopsis MYB24 Regulates Jasmonate-Mediated Stamen Development. Front Plant Sci. 2017;8:1525.
Huang H, Gong Y, Liu B, Wu D, Zhang M, **e D, Song S. The DELLA proteins interact with MYB21 and MYB24 to regulate filament elongation in Arabidopsis. BMC Plant Biol. 2020;20(1):64.
Hunt AG, Xu R, Addepalli B, Rao S, Forbes KP, Meeks LR, **ng D, Mo M, Zhao H, Bandyopadhyay A, et al. Arabidopsis mRNA polyadenylation machinery: comprehensive analysis of protein-protein interactions and gene expression profiling. BMC Genomics. 2008;9:220.
Jan A, Maruyama K, Todaka D, Kidokoro S, Abo M, Yoshimura E, Shinozaki K, Nakashima K, Yamaguchi-Shinozaki K. OsTZF1, a CCCH-tandem zinc finger protein, confers delayed senescence and stress tolerance in rice by regulating stress-related genes. Plant Physiol. 2013;161(3):1202–16.
Jiang Y, Deyholos MK. Functional characterization of Arabidopsis NaCl-inducible WRKY25 and WRKY33 transcription factors in abiotic stresses. Plant Mol Biol. 2008;69(1–2):91–5.
Kim WC, Kim JY, Ko JH, Kang H, Kim J, Han KH. AtC3H14, a plant-specific tandem CCCH zinc-finger protein, binds to its target mRNAs in a sequence-specific manner and affects cell elongation in Arabidopsis thaliana. Plant J. 2014;80(5):772–84.
Kong Z, Li M, Yang W, Xu W, Xue Y. A novel nuclear-localized CCCH-type zinc finger protein, OsDOS, is involved in delaying leaf senescence in rice. Plant Physiol. 2006;141(4):1376–88.
Krishnamurthy P, Vishal B, Ho WJ, Lok FCJ, Lee FSM, Kumar PP. Regulation of a Cytochrome P450 Gene CYP94B1 by WRKY33 Transcription Factor Controls Apoplastic Barrier Formation in Roots to Confer Salt Tolerance. Plant Physiol. 2020;184(4):2199–15.
Kumar D, Chattopadhyay S. Glutathione modulates the expression of heat shock proteins via the transcription factors BZIP10 and MYB21 in Arabidopsis. J Exp Bot. 2018;69(15):3729–43.
Lee SJ, Jung HJ, Kang H, Kim SY. Arabidopsis zinc finger proteins AtC3H49/AtTZF3 and AtC3H20/AtTZF2 are involved in ABA and JA responses. Plant Cell Physiol. 2012;53(4):673–86.
Li S, Fu Q, Chen L, Huang W, Yu D. Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance. Planta. 2011;233(6):1237–52.
Lin PC, Pomeranz MC, Jikumaru Y, Kang SG, Hah C, Fujioka S, Kamiya Y, Jang JC. The Arabidopsis tandem zinc finger protein AtTZF1 affects ABA- and GA-mediated growth, stress and gene expression responses. Plant J. 2011;65(2):253–68.
Liu B, Jiang Y, Tang H, Tong S, Lou S, Shao C, Zhang J, Song Y, Chen N, Bi H. The ubiquitin E3 ligase SR1 modulates the submergence response by degrading phosphorylated WRKY33 in Arabidopsis. Plant Cell. 2021;33(5):1771–89.
Liu S, Kracher B, Ziegler J, Birkenbihl RP, Somssich IE. Negative regulation of ABA signaling by WRKY33 is critical for Arabidopsis immunity towards Botrytis cinerea 2100. Elife. 2015;4:e07295.
Liu YG, Chen Y. High-efficiency thermal asymmetric interlaced PCR for amplification of unknown flanking sequences. Biotechniques. 2007;43(5):649–56.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25(4):402–8.
Mandaokar A, Browse J. MYB108 acts together with MYB24 to regulate jasmonate-mediated stamen maturation in Arabidopsis. Plant Physiol. 2009;149(2):851–62.
Ohama N, Sato H, Shinozaki K, Yamaguchi-Shinozaki K. Transcriptional Regulatory Network of Plant Heat Stress Response. Trends Plant Sci. 2017;22(1):53–65.
Pomeranz M, Finer J, Jang JC. Putative molecular mechanisms underlying tandem CCCH zinc finger protein mediated plant growth, stress, and gene expression responses. Plant Signal Behav. 2011;6(5):647–51.
Pomeranz M, Lin PC, Finer J, Jang JC. AtTZF gene family localizes to cytoplasmic foci. Plant Signal Behav. 2010a;5(2):190–2.
Pomeranz MC, Hah C, Lin PC, Kang SG, Finer JJ, Blackshear PJ, Jang JC. The Arabidopsis tandem zinc finger protein AtTZF1 traffics between the nucleus and cytoplasmic foci and binds both DNA and RNA. Plant Physiol. 2010b;152(1):151–65.
Qiu A, Lei Y, Yang S, Wu J, Li J, Bao B, Cai Y, Wang S, Lin J, Wang Y, et al. CaC3H14 encoding a tandem CCCH zinc finger protein is directly targeted by CaWRKY40 and positively regulates the response of pepper to inoculation by Ralstonia solanacearum. Mol Plant Pathol. 2018;19(10):2221–35.
Qu J, Kang SG, Wang W, Musier-Forsyth K, Jang JC. The Arabidopsis thaliana tandem zinc finger 1 (AtTZF1) protein in RNA binding and decay. Plant J. 2014;78(3):452–67.
Sainsbury F, Thuenemann EC, Lomonossoff GP. pEAQ: versatile expression vectors for easy and quick transient expression of heterologous proteins in plants. Plant Biotechnol J. 2009;7(7):682–93.
Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative CT method. Nat Protoc. 2008;3(6):1101–8.
Seok HY, Nguyen LV, Park HY, Tarte VN, Ha J, Lee SY, Moon YH. Arabidopsis non-TZF gene AtC3H17 functions as a positive regulator in salt stress response. Biochem Biophys Res Commun. 2018;498(4):954–9.
Seok HY, Woo DH, Park HY, Lee SY, Tran HT, Lee EH, Vu Nguyen L, Moon YH. AtC3H17, a Non-Tandem CCCH Zinc Finger Protein, Functions as a Nuclear Transcriptional Activator and Has Pleiotropic Effects on Vegetative Development, Flowering and Seed Development in Arabidopsis. Plant Cell Physiol. 2016;57(3):603–15.
Shi ZH, Zhang C, Xu XF, Zhu J, Zhou Q, Ma LJ, Niu J, Yang ZN. Overexpression of AtTTP affects ARF17 expression and leads to male sterility in Arabidopsis. PLoS ONE. 2015;10(3): e0117317.
Song S, Qi T, Huang H, Ren Q, Wu D, Chang C, Peng W, Liu Y, Peng J, **e D. The Jasmonate-ZIM domain proteins interact with the R2R3-MYB transcription factors MYB21 and MYB24 to affect Jasmonate-regulated stamen development in Arabidopsis. Plant Cell. 2011;23(3):1000–13.
Sun J, Jiang H, Xu Y, Li H, Wu X, **e Q, Li C. The CCCH-type zinc finger proteins AtSZF1 and AtSZF2 regulate salt stress responses in Arabidopsis. Plant Cell Physiol. 2007;48(8):1148–58.
Teixeira EI, Fischer G, van Velthuizen H, Walter C, Ewert F. Global hot-spots of heat stress on agricultural crops due to climate change. Agric for Meteorol. 2013;170:206–15.
Wahid A, Gelani S, Ashraf M, Foolad M. Heat tolerance in plants: An overview. Environ Exp Bot. 2007;61(3):199–223.
Wang D, Chai G, Xu L, Yang K, Zhuang Y, Yang A, Liu S, Kong Y, Zhou G. Phosphorylation-Mediated Inactivation of C3H14 by MPK4 Enhances Bacterial-Triggered Immunity in Arabidopsis. Plant Physiol. 2022a;190(3):1941–59.
Wang D, Guo Y, Wu C, Yang G, Li Y, Zheng C. Genome-wide analysis of CCCH zinc finger family in Arabidopsis and rice. BMC Genomics. 2008;9:44.
Wang D, Xu H, Huang J, Kong Y, AbuQamar S, Yu D, Liu S, Zhou G, Chai G. The Arabidopsis CCCH protein C3H14 contributes to basal defense against Botrytis cinerea mainly through the WRKY33-dependent pathway. Plant Cell Environ. 2020;43(7):1792–806.
Wang L, Chen J, Zhao Y, Wang S, Yuan M. OsMAPK6 phosphorylates a zinc finger protein OsLIC to promote downstream OsWRKY30 for rice resistance to bacterial blight and leaf streak. J Integr Plant Biol. 2022b;64(5):1116–30.
Wang W, Liu B, Xu M, Jamil M, Wang G. ABA-induced CCCH tandem zinc finger protein OsC3H47 decreases ABA sensitivity and promotes drought tolerance in Oryza sativa. Biochem Biophys Res Commun. 2015;464(1):33–7.
Wu Z, Li T, Cao X, Zhang D, Teng N. Lily WRKY factor LlWRKY22 promotes thermotolerance through autoactivation and activation of LlDREB2B. Hortic Res. 2022a;9:uhac186.
Wu Z, Li T, Liu X, Yuan G, Hou H, Teng N. A novel R2R3-MYB transcription factor LlMYB305 from Lilium longiflorum plays a positive role in thermotolerance via activating heat-protective genes. Environ Exp Bot. 2021;184: 104399.
Wu Z, Li T, **ang J, Teng R, Zhang D, Teng N. A lily membrane-associated NAC transcription factor LlNAC014 is involved in thermotolerance via activation of the DREB2-HSFA3 module. J Exp Bot. 2023;74(3):945–63.
Wu Z, Li T, Zhang D, Teng N. Lily HD-Zip I transcription factor LlHB16 promotes thermotolerance by activating LlHSFA2 and LlMBF1c. Plant Cell Physiol. 2022b;63(11):1729–44.
**e Z, Lin W, Yu G, Cheng Q, Xu B, Huang B. Improved cold tolerance in switchgrass by a novel CCCH-type zinc finger transcription factor gene, PvC3H72, associated with ICE1-CBF-COR regulon and ABA-responsive genes. Biotechnol Biofuels. 2019;12:224.
Xu J, Huang Z, Du H, Tang M, Fan P, Yu J, Zhou Y. SEC1-C3H39 module fine-tunes cold tolerance by mediating its target mRNA degradation in tomato. New Phytol. 2023;237(3):870–84.
Xu L, Liu T, **ong X, Liu W, Yu Y, Cao J. AtC3H18L is a stop-codon read-through gene and encodes a novel non-tandem CCCH zinc-finger protein that can form cytoplasmic foci similar to mRNP granules. Biochem Biophys Res Commun. 2020a;528(1):140–5.
Xu L, Liu T, **ong X, Liu W, Yu Y, Cao J. Overexpression of Two CCCH-type Zinc-Finger Protein Genes Leads to Pollen Abortion in Brassica campestris ssp. chinensis. Genes (Basel). 2020b;11(11):1287.
Xu L, Liu T, **ong X, Shen X, Huang L, Yu Y, Cao J. Highly Overexpressed AtC3H18 Impairs Microgametogenesis via Promoting the Continuous Assembly of mRNP Granules. Front Plant Sci. 2022;13: 932793.
Xu L, **ong X, Liu W, Liu T, Yu Y, Cao J. BcMF30a and BcMF30c, Two Novel Non-Tandem CCCH Zinc-Finger Proteins, Function in Pollen Development and Pollen Germination in Brassica campestris ssp. chinensis. Int J Mol Sci. 2020c;21(17):6428.
Zheng Z, Qamar SA, Chen Z, Mengiste T. Arabidopsis WRKY33 transcription factor is required for resistance to necrotrophic fungal pathogens. Plant J. 2006;48(4):592–605.
Zhou T, Yang X, Wang L, Xu J, Zhang X. GhTZF1 regulates drought stress responses and delays leaf senescence by inhibiting reactive oxygen species accumulation in transgenic Arabidopsis. Plant Mol Biol. 2014;85(1–2):163–77.
Acknowledgements
The authors thank many colleagues and collaborators for their contribution to our work described here.
Funding
Open access funding provided by Shanghai Jiao Tong University. This work was supported by the National Natural Science Foundation of China (32272761, 31902055), the Fundamental Research Funds for the Central Universities (KYZZ2022004), and the Project for Crop Germplasm Resources Conservation of Jiangsu (2021-SJ-011).
Author information
Authors and Affiliations
Contributions
N.T., Z.W., and J.L. conceived and designed the experiments; Z.W., J.L., and T.L. performed the experiments; D.Z. prepared the materials and provided technical help; Z.W. and J.L. analyzed the data and wrote the manuscript; all the authors revised and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Supplementary Table S1.
The promoter sequences are used for EMSA assay.Supplementary Table S2. Primers of LlC3H18 isolation.Supplementary Table S3. Primers used for vector reconstruction.Supplementary Table S4. RT-qPCR primers.Supplementary Table S5. Primers used for atc3h18 mutant identification.
Additional file 2: Supplementary Figure S1.
Heat map of the expression of CCCH-type genes in lily leaves with heat stress. Supplementary Figure S2. Phylogenetic analysis of LlC3H18 and CCCH-type proteins of Arabidopsis.Supplementary Figure S3. Sequence analysis of LlC3H18.Supplementary Figure S4. Expression analysis of heat-related genes in the wild-type and LlC3H18 transgenic plants under normal and heat stress conditions.Supplementary Figure S5. Identified T-DNA insertion atc3h18 mutant and detection its thermotolerance. Supplementary Figure S6. Expression analysis of heat-related genes in the wild-type and atc3h18 mutant plants under normal and heat stress conditions.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Wu, Z., Liang, J., Li, T. et al. A LlMYB305-LlC3H18-LlWRKY33 module regulates thermotolerance in lily. Mol Horticulture 3, 15 (2023). https://doi.org/10.1186/s43897-023-00064-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43897-023-00064-1