Abstract
Background
Observational studies have indicated a potential association between autoimmune diseases and the occurrence of sepsis, with an increased risk of mortality among affected patients. However, whether a causal relationship exists between the two remains unknown.
Methods
In the Mendelian randomization (MR) study, we accessed exposure Genome-wide association study (GWAS) data from both the MRC Integrative Epidemiology Unit (MRC-IEU) and the FinnGen consortium. GWAS data for sepsis and its 28-day mortality were obtained from MRC-IEU. We employed univariable, multivariable, and reverse MR analyses to explore potential associations between autoimmune disorders and sepsis and its 28-day mortality. Additionally, a two-step mediation MR analysis was performed to investigate indirect factors possibly influencing the relationship between autoimmune disorders and sepsis. Afterward, we conducted an observational analysis to further explore the relationship between autoimmune disease and occurrence as well as 28-day mortality of sepsis using a real-world database (the MIMIC-IV database). A cohort of 2537 patients diagnosed with autoimmune disease were extracted from the database for analysis. Multivariable logistic regression models were used to confirm the association between autoimmune diseases and the occurrence of sepsis, as well as the 28-day mortality associated with sepsis.
Results
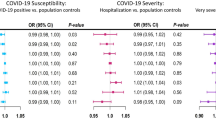
In univariable MR analysis, there appeared to be causal relationships between genetically predicted type 1 diabetes (OR = 1.036, 95% CI = 1.023–1.048, p = 9.130E-09), rheumatoid arthritis (OR = 1.077, 95% CI = 1.058–1.097, p = 1.00E-15) and sepsis, while a potential causal link was observed between celiac disease and sepsis (OR = 1.013, 95% CI = 1.002–1.024, p = 0.026). In a subsequent multivariable MR analysis, only rheumatoid arthritis was found to be independently associated with the risk of sepsis (OR = 1.138, 95% CI = 1.044–1.240, p = 3.36E-03). Furthermore, there was no causal link between autoimmune disorders and 28-day mortality from sepsis. In reverse MR analysis, sepsis was suggested to potentially trigger the onset of psoriasis (OR = 1.084, 95% CI = 1.040–1.131, p = 1.488E-04). In the real-world observational study, adjusting for multiple confounders, rheumatoid arthritis (OR = 1.34, 95% CI = 1.11–1.64, p = 0.003) and multiple sclerosis (OR = 1.31, 95% CI = 1.03–1.68, p = 0.02) were associated with a higher risk of sepsis. In addition, we did not find that autoimmune diseases were associated with 28-day mortality from sepsis.
Conclusion
Both in observational and MR analysis, only rheumatoid arthritis is highly correlated with occurrence of sepsis. However, autoimmune disease was not associated with an increased 28-day mortality in patient with sepsis. Sepsis may increase the risk of develo** psoriasis.
Similar content being viewed by others
Introduction
Autoimmune diseases are a group of diseases in which the immune system mounts an immune response against its own normal tissue components, often resulting in chronic tissue and organ damage, affecting approximately 7.6–9.4% of the global population [1]. The primary features of autoimmune diseases include the production of self-targeting antibodies and abnormalities in the function of immune cells. Often, the management of these conditions involves the use of immunomodulatory or immunosuppressive medications, which can result in compromised immune function and an elevated risk of infections [2]. Although retrospective analyses of autoimmune diseases have primarily associated patients with respiratory infections, it is important to highlight that the main drivers of ICU admissions and mortality in this group are severe infections [3, 4]. The evolving environmental changes brought about by societal industrialization have contributed to an increasing incidence of autoimmune diseases. Consequently, the associated risk of pathogenic infections is expected to rise as well [5, 6]. Therefore, prioritizing infection risks in individuals with autoimmune diseases is crucial for mitigating the emergence of life-threatening infectious conditions.
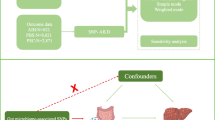
Sepsis is a complex, infection-induced systemic inflammatory response disorder characterized by an imbalance, often accompanied by acute organ dysfunction and a high mortality rate [7]. Despite a decline of 37.0% in the age-standardized incidence of sepsis and a 52.8% decrease in mortality, the burden of this severe condition persists, particularly in develo** countries [8, 9]. It is noteworthy that the increased overall burden could be attributed to severe infections resulting from autoimmune diseases and their associated treatments, such as the use of corticosteroids [10]. Some patients with autoimmune diseases were admitted to the intensive care unit at the initial diagnosis [11,12,13,14]. Among these cases, sepsis (severe infection) stands out as the primary cause of ICU mortality, followed by acute disease exacerbations [15]. The relationship between autoimmune diseases and sepsis has long been a subject of interest [2]. Therefore, we extracted information on patients with autoimmune diseases from the MIMIC-IV database to explore whether autoimmune diseases increase occurrence of sepsis and the 28-day mortality of sepsis. However, due to limitations in retrospective research, such as potential confounders and selection bias, a consistent conclusion regarding the relationship between autoimmune diseases and sepsis has not been reached [ Figure 1 shows the study design and the assumptions of MR in our study [20]. We used publicly available summary statistics from Genome-wide association study (GWAS) sources of predominantly European origin. All studies had current ethical clearance from their respective institutional review boards, including written informed consent from participants and strict quality control. As all analyses herein are based on publicly available summary data, no ethical approval from institutional review boards was required for this study. Three basic assumptions are required for the genetic variants to qualify as valid instrumental variables (IVs): (1) they should be robustly associated with the exposure; (2) they should not be associated with potential confounders of the exposure-outcome association; and (3) they should not influence the outcome by any variable other than the exposure [20]. To validate the initial MR hypothesis, we utilized independent single nucleotide polymorphisms (SNPs) that exhibited a robust association with the exposure, reaching genome-wide significance (P < 5 × 10–6). These SNPs were carefully chosen to ensure minimal linkage disequilibrium (r2 < 0.01) within a clump window larger than 5000 kb, thus ensuring their independence. If we follow the same inclusion criteria, the exposure of reverse MR analysis includes too few SNPs, so we have adopted the following criteria. The reverse MR analyses inclusion criteria for the instrumental variable SNP were as follows: P < 1 × 10–5, r2 < 0.001 within a clump window larger than 10,000 kb. To further refine the first hypothesis, we quantified the proportion of phenotypic variation explained by the entire set of SNPs and assessed the strength of our instrumental variables using the F statistic (beta2/se2). An F-statistic exceeding 10 was considered indicative of a robust instrument [21]. R2 was calculated as beta2/[beta2 + se2*(N− 2)], N being the sample size, and the genetic variability explained by each SNP was calculated [22]. Finally, after eliminating palindromic SNPs, we proceeded to utilize the remaining selected SNPs as our instrumental variables for subsequent analyses. To delve into the direct influence of distinct autoimmune diseases on sepsis, we adopted a multivariate MR approach-an extension of the conventional univariate MR. This approach duly acknowledged the inherent interplay among SNPs used in MR analyses, often manifesting shared associations across different autoimmune conditions. In our study, the SNPs utilized for multivariate MR were formulated as combinations of instrumental variables per exposure, thereby accounting for the intricate web of associations (including those associated with phenotypes of at least two autoimmune diseases). This study is reported in line with the STROBE-MR guidance, with the checklist available in the Supporting information [23]. Recognizing that utilizing diverse populations could potentially lead to biased estimates, we constrained the genetic background of the population in the MR study to individuals of European ancestry [24]. Exposures included a total of 10 distinct autoimmune diseases, and we aligned our analysis with data available from the FinnGen consortium (R9) (https://www.finngen.fi/fi.) and the MRC-IEU online database (https://gwas.mrcieu.ac.uk/). The autoimmune diseases considered for inclusion in our analysis were as follows: systemic lupus erythematosus (SLE), ankylosing spondylitis, multiple sclerosis (MS), primary biliary cholangitis, rheumatoid arthritis (RA), Crohn's disease (CD), ulcerative colitis (UC), type 1 diabetes (T1DM), celiac disease, psoriasis. We systematically reviewed and summarized the general characteristics of each autoimmune disease, and we presented these aggregated data in Additional file 3: Table S1. It is important to note that the ten autoimmune diseases from the Finngen consortium were defined using the codes of the International Classification of Diseases (ICD-9) and ICD-10. In the FinnGen consortium, individuals with undefined sex, high genotype deletion (> 5%), excess heterozygosity (± 4 standard deviations ((SDs)), and non-Finnish ancestry were excluded. All genetic association effect sizes were calculated by logistic regression, and adjusted for age, sex, and genetic principal components [25]. In the MRC-IEU database, all 10 included diseases have been previously published online. However, due to their diverse origins from different research teams, the analytical methods employed and the controlled confounding factors are not entirely uniform. For a comprehensive understanding, please refer to the cited references for detailed information [26,27,28,29,30,31,32,62]. It is well known that Neutrophil extracellular traps (NETs) are one of the major factors contributing to the severity of septic disease, and that in the early stages of sepsis, depletion of NETs does not help to prevent or contain systemic infections, and even exacerbates pathological changes [63,64,65,66]. The citrullination of nuclear histones by PAD4 leads to chromatin depolymerization, which is a key step in the formation of NETs [67]. Therefore, the abnormal structure and function of PAD4 make RA patients more susceptible to sepsis. Chitinase-3 like-protein-1 (CHI3L1, other name YKL-40) may be another important protein molecule involved in the pathological process of RA and sepsis.YKL-40 is synthesized and secreted by a wide variety of cells including macrophages, neutrophils, and chondrocytes and plays an important role in tissue injury, inflammation, tissue repair and remodeling responses [68]. It was found that YKL-40 levels were significantly increased in RA patients and induced the expression of IL-1β and TNF-α, which were involved in the inflammatory response in RA [69]. A study by Kornblit et al. found that YKL-40 levels were also significantly elevated in patients with sepsis and that YKL-40 promoted the expression of inflammatory factors [70]. Thus, RA patients may be more susceptible to sepsis due to genetic variants, and the CHI3L1 genotype (rs4950928) may be a potential locus [69, 70]. In order to further explore the potential mediation of autoimmune diseases in the occurrence of sepsis through underlying intermediary factors, we included risk factors related to autoimmune diseases associated with sepsis [36]. We found that not all autoimmune diseases lead to a decrease in blood cell counts; only SLE, celiac disease, T1DM, and reduced blood cell count showed a causal association, whereas conditions such as RA and primary biliary cholangitis had less pronounced effects. Despite an inverse causal trend between blood cell counts and sepsis, statistical significance was lacking. While some observational studies suggest a predictive relationship between changes in blood cell counts and the risk of severe infection in autoimmune diseases [71, 75,76,77,78]. Hydroxychloroquine reduces the risk of severe infections in SLE patients, while the use of glucocorticoids, especially in high doses, is closely related to severe infections [79]. Montgomery et al. found functional impairment to be a significant risk factor for severe infections in multiple sclerosis patients [17]. Therefore, patients with autoimmune diseases require closer monitoring of organ function, comorbidities, medication usage, and other factors to reduce the risk of sepsis. It is worth noting that RA is associated with an increased risk of sepsis, which can occur early in the course of the disease. This suggests that when managing patients with RA, early attention, timely treatment, and early prediction may be required to reduce the occurrence of severe infections. For example, for outpatient patients with fever or other signs of infection, infection-related markers and imaging tests should be monitored, and they should be hospitalized if necessary; for hospitalized patients, early monitoring of biomarkers and symptoms of infection, more active adjustment of antibiotics after infection, and more close monitoring for patients with high-risk factors (such as, indwelling catheters and steroid use, etc.) should be noticed. The 28-day mortality risk in sepsis is a crucial measure of disease severity, and whether autoimmune diseases increase this risk remains inconclusive. A retrospective study by Antón et al. found that autoimmune diseases often lead to a higher mortality rate in critically ill patients [4]. However, this study primarily predicted high mortality risk without correcting for concurrent confounding factors such as SOFA score, age, underlying diseases, and had a relatively small sample size. In our analysis using two-sample MR analysis, we inferred causal relationships between autoimmune diseases and sepsis at the genetic level. We did not find a causal association between autoimmune diseases and the 28-day mortality rate in sepsis. Similarly, in the retrospective analysis from MIMIC-IV, there was no observed relationship between autoimmune diseases and the 28-day mortality rate in sepsis. Therefore, we believe that autoimmune diseases do not increase the 28-day mortality rate in sepsis. This might be related to the immune dysregulation caused by autoimmune diseases, leading to imbalanced cytokines in the sepsis inflammatory cascade [80], making it difficult to form a cascading reaction. The early mortality in sepsis is closely associated with this inflammation storm. Jorge et al.'s observational study found that the risk of death in autoimmune diseases may be related to factors such as experiencing shock upon admission to the intensive care unit, having hemoglobin levels below 8 g/dL, using immunosuppressive agents before ICU admission, and having low complement C3 levels [81]. Additionally, the quality of care provided by hospitals is a key factor influencing patient mortality risk, with more experienced hospitals often having lower mortality rates [3]. Therefore, for autoimmune disease patients admitted to the ICU, it is crucial to focus on the management of complications while enhancing diagnostic and treatment capabilities specific to autoimmune diseases to reduce the risk of mortality. A key feature of sepsis is the immune dysfunction triggered by infections, leading to prolonged alterations in immune function such as changes in immune cell functionality and numbers. Similarly, the immunopathological mechanisms of autoimmune diseases are accompanied by disruptions in immune function [39, 82]. Furthermore, infections caused by pathogenic microorganisms can act as triggering factors for autoimmune diseases [83]. However, it remains uncertain whether the immune dysfunction triggered by severe infections caused by pathogenic microorganisms could lead to the development of autoimmune diseases. Through reverse MR analysis, we identified a causal relationship between sepsis and psoriasis, but no associations with other autoimmune diseases, and current research has also found that infection is an important trigger for the occurrence of psoriasis [84]. The specific mechanisms underlying this relationship require further investigation. MR provides a novel method to discover associations between different diseases at the genetic level, offering a new perspective for future observational studies. This study has several limitations in the MR analysis. First, potential horizontal pleiotropy is a concern in any MR study. In our research, we did not observe significant evidence of pleiotropic effects in all exposure analyses using the MR-Egger intercept test. Additionally, the MR-PRESSO analysis detected few outliers, and associations remained consistent after removing outlier SNPs. However, the possibility of undetected outliers still exists. Second, sample overlap might be a concern as we selected a subset of autoimmune diseases from the FinnGen consortium. Nonetheless, sample overlap is unlikely to bias our results significantly, given that our IVs were selected from large-scale GWAS. Third, due to the limited number of SNPs meeting the inclusion criteria for certain autoimmune diseases (P value < 5e− 08, R2 = 0.001 with kb = 10,000), we slightly relaxed the selection criteria, which may introduce a certain level of false positives. Fourth, genetic factors are not the sole determinants of autoimmune disease onset; environmental factors also play a role in triggering disease processes. Therefore, our MR analysis lacks associations between genetically predicted autoimmune diseases and sepsis risk, but this does not exclude the potential impact of autoimmune diseases on the pathophysiology of sepsis [39]. Fifth, the genetic associations of blood cell count, inflammatory cytokines, and are based on relatively small global genomic studies, potentially leading to issues of statistical power. Sixth, univariate MR analysis may not capture the direct impact of specific biomarkers on disease outcomes, as the effect of a biomarker might be mediated by other biomarkers within a complex network. Seventh, all our analyses are based on individuals of European ancestry; generalizability to other populations requires further investigation. In real-world retrospective studies, there are also limitations. First, our results could be influenced by diagnostic bias, where the severity of autoimmune diseases and the immunocompromised state of autoimmune disease patients might lead them to be admitted to ICUs earlier than other populations, potentially resulting in better survival rates. This selection process could lead to a higher incidence of autoimmune diseases in the ICU population. Additionally, we controlled for potential influencing factors, yet the overall confounding factors included in our analysis might not be exhaustive, such as other clinical scores that were not incorporated. We believe that further research with more diverse pre-ICU admission data from intensive care units would help fully eliminate diagnostic bias. Second, the MIMIC database is derived from a single-center research institution, which may limit the generalizability of the study outcomes. Third, due to the limitations of the database, we were unable to distinguish whether patients with autoimmune diseases were diagnosed after their first ICU admission, i.e., it was difficult to determine whether the autoimmune disease was a newly diagnosed disease or a comorbidity, although this group of patients may be rare. Fourth, because Mendelian randomization uses integrated data, whereas observational studies use individual data, it is difficult for us to achieve correction for the same confounders for the two different analytical methods. In this study, our aim was mainly to assess the association of autoimmune diseases with the development of sepsis and 28-day mortality through a MR and observational study. In our results, genetically predicted RA was independently associated with the development of sepsis. We did not find that none of the other autoimmune diseases predicted by genes were independently associated with the development of sepsis, including subsequent mediation analyses. In addition, neither observational studies nor MR analyses found autoimmune diseases to be associated with 28-day mortality from sepsis., Surprisingly, there was a causal relationship between genetically predicted sepsis and the development of psoriasis.Methods
Mendelian randomization
Study design and genetic instrument selection
Data sources for exposures, mediators and outcomes
Conclusion
Availability of data and materials
The datasets analyzed in this study are publicly available summary statistics. Data used can be obtained upon a reasonable request to the corresponding author.
References
Gutierrez-Arcelus M, Rich SS, Raychaudhuri S. Autoimmune diseases - connecting risk alleles with molecular traits of the immune system. Nat Rev Genet. 2016;17(3):160–74.
Tsonis IA, Avrameas S, Moutsopoulos HM. Autoimmunity and pathophysiology. J Autoimmun. 2007;29(4):203–5.
Tektonidou MG, Dasgupta A, Ward MM. Interhospital variation in mortality among patients with systemic lupus erythematosus and sepsis in the USA. Rheumatology (Oxford). 2019;58(10):1794–801.
Antón JM, Castro P, Espinosa G, Marcos M, Gandía M, Merchán R, et al. Mortality and long term survival prognostic factors of patients with systemic autoimmune diseases admitted to an intensive care unit: a retrospective study. Clin Exp Rheumatol. 2012;30(3):338–44.
Rose NR. Prediction and prevention of autoimmune disease in the 21st century: a review and preview. Am J Epidemiol. 2016;183(5):403–6.
Conrad N, Misra S, Verbakel JY, Verbeke G, Molenberghs G, Taylor PN, et al. Incidence, prevalence, and co-occurrence of autoimmune disorders over time and by age, sex, and socioeconomic status: a population-based cohort study of 22 million individuals in the UK. Lancet. 2023;401(10391):1878–90.
Cecconi M, Evans L, Levy M, Rhodes A. Sepsis and septic shock. Lancet. 2018;392(10141):75–87.
Rudd KE, Johnson SC, Agesa KM, Shackelford KA, Tsoi D, Kievlan DR, et al. Global, regional, and national sepsis incidence and mortality, 1990–2017: analysis for the Global burden of disease study. Lancet. 2020;395(10219):200–11.
Rudd KE, Kissoon N, Limmathurotsakul D, Bory S, Mutahunga B, Seymour CW, et al. The global burden of sepsis: barriers and potential solutions. Crit Care. 2018;22(1):232.
Tektonidou MG, Wang Z, Dasgupta A, Ward MM. Burden of serious infections in adults with systemic lupus erythematosus: a national population-based study, 1996–2011. Arthritis Care Res (Hoboken). 2015;67(8):1078–85.
Godeau B, Mortier E, Roy PM, Chevret S, Bouachour G, Schlemmer B, et al. Short and longterm outcomes for patients with systemic rheumatic diseases admitted to intensive care units: a prognostic study of 181 patients. J Rheumatol. 1997;24(7):1317–23.
Bouachour G, Roy PM, Tirot P, Guerin O, Gouello JP, Alquier P. Prognosis of systemic diseases diagnosed in intensive care units. Presse Med. 1996;25(18):837–41.
Moreels M, Mélot C, Leeman M. Prognosis of patients with systemic rheumatic diseases admitted to the intensive care unit. Intensive Care Med. 2005;31(4):591–3.
Cruz BA, Ramanoelina J, Mahr A, Cohen P, Mouthon L, Cohen Y, et al. Prognosis and outcome of 26 patients with systemic necrotizing vasculitis admitted to the intensive care unit. Rheumatology (Oxford). 2003;42(10):1183–8.
Quintero OL, Rojas-Villarraga A, Mantilla RD, Anaya JM. Autoimmune diseases in the intensive care unit. An update Autoimmun Rev. 2013;12(3):380–95.
Nelson RE, **e Y, DuVall SL, Butler J, Kamauu AW, Knippenberg K, et al. Multiple sclerosis and risk of infection-related hospitalization and death in US veterans. Int J MS Care. 2015;17(5):221–30.
Montgomery S, Hillert J, Bahmanyar S. Hospital admission due to infections in multiple sclerosis patients. Eur J Neurol. 2013;20(8):1153–60.
Sekula P, Del Greco MF, Pattaro C, Köttgen A. Mendelian randomization as an approach to assess causality using observational data. J Am Soc Nephrol. 2016;27(11):3253–65.
Verduijn M, Siegerink B, Jager KJ, Zoccali C, Dekker FW. Mendelian randomization: Use of genetics to enable causal inference in observational studies. Nephrol Dial Transplant. 2010;25(5):1394–8.
Hemani G, Zheng J, Elsworth B, Wade KH, Haberland V, Baird D, et al. The MR-Base platform supports systematic causal inference across the human phenome. Elife. 2018;7:e34408.
Burgess S, Thompson SG. Avoiding bias from weak instruments in Mendelian randomization studies. Int J Epidemiol. 2011;40(3):755–64.
Tian D, Zhang L, Zhuang Z, Huang T, Fan D. A two-sample Mendelian randomization analysis of modifiable risk factors and intracranial aneurysms. Sci Rep. 2022;12(1):7659.
Skrivankova VW, Richmond RC, Woolf BAR, Davies NM, Swanson SA, VanderWeele TJ, et al. Strengthening the reporting of observational studies in epidemiology using mendelian randomisation (STROBE-MR): explanation and elaboration. BMJ. 2021;375:n2233.
Burgess S, Labrecque JA. Mendelian randomization with a binary exposure variable: interpretation and presentation of causal estimates. Eur J Epidemiol. 2018;33(10):947–52.
Consortium TF, https://www.finngen.fi/fi. (Accessed 16 July 2023).
Ha E, Bae SC, Kim K. Large-scale meta-analysis across East Asian and European populations updated genetic architecture and variant-driven biology of rheumatoid arthritis, identifying 11 novel susceptibility loci. Ann Rheum Dis. 2021;80(5):558–65.
Bentham J, Morris DL, Graham DSC, Pinder CL, Tombleson P, Behrens TW, et al. Genetic association analyses implicate aberrant regulation of innate and adaptive immunity genes in the pathogenesis of systemic lupus erythematosus. Nat Genet. 2015;47(12):1457–64.
Liu JZ, van Sommeren S, Huang H, Ng SC, Alberts R, Takahashi A, et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat Genet. 2015;47(9):979–86.
Forgetta V, Manousaki D, Istomine R, Ross S, Tessier MC, Marchand L, et al. Rare genetic variants of large effect influence risk of type 1 diabetes. Diabetes. 2020;69(4):784–95.
Trynka G, Hunt KA, Bockett NA, Romanos J, Mistry V, Szperl A, et al. Dense genoty** identifies and localizes multiple common and rare variant association signals in celiac disease. Nat Genet. 2011;43(12):1193–201.
Tsoi LC, Spain SL, Knight J, Ellinghaus E, Stuart PE, Capon F, et al. Identification of 15 new psoriasis susceptibility loci highlights the role of innate immunity. Nat Genet. 2012;44(12):1341–8.
Cortes A, Hadler J, Pointon JP, Robinson PC, Karaderi T, Leo P, et al. Identification of multiple risk variants for ankylosing spondylitis through high-density genoty** of immune-related loci. Nat Genet. 2013;45(7):730–8.
Beecham AH, Patsopoulos NA, **fara DK, Davis MF, Kemppinen A, Cotsapas C, et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet. 2013;45(11):1353–60.
Cordell HJ, Han Y, Mells GF, Li Y, Hirschfield GM, Greene CS, et al. International genome-wide meta-analysis identifies new primary biliary cirrhosis risk loci and targetable pathogenic pathways. Nat Commun. 2015;6:8019.
Ben E, Matthew L, Tessa A, Yi L, Peter M, Jon H et al. The MRC IEU OpenGWAS data infrastructure. bioRxiv 2020:2020.2008.2010.244293.
Sheth M, Benedum CM, Celi LA, Mark RG, Markuzon N. The association between autoimmune disease and 30-day mortality among sepsis ICU patients: a cohort study. Crit Care. 2019;23(1):93.
Ye CJ, Kong LJ, Wang YY, Dou C, Zheng J, Xu M, et al. Mendelian randomization evidence for the causal effects of socio-economic inequality on human longevity among Europeans. Nat Hum Behav. 2023;7(8):1357–70.
Consortium TF, https://risteys.finregistry.fi/endpoints/AB1_SEPSIS. (Accessed 16 July 2023).
Wang L, Wang FS, Gershwin ME. Human autoimmune diseases: a comprehensive update. J Intern Med. 2015;278(4):369–95.
Dendrou CA, Petersen J, Rossjohn J, Fugger L. HLA variation and disease. Nat Rev Immunol. 2018;18(5):325–39.
Hemani G, Bowden J, Davey SG. Evaluating the potential role of pleiotropy in Mendelian randomization studies. Hum Mol Genet. 2018;27(R2):R195-r208.
Hartwig FP, Davey Smith G, Bowden J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int J Epidemiol. 2017;46(6):1985–98.
Burgess S, Thompson SG. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol. 2017;32(5):377–89.
Verbanck M, Chen CY, Neale B, Do R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet. 2018;50(5):693–8.
Rees JMB, Wood AM, Burgess S. Extending the MR-Egger method for multivariable Mendelian randomization to correct for both measured and unmeasured pleiotropy. Stat Med. 2017;36(29):4705–18.
Johnson AEW, Bulgarelli L, Shen L, Gayles A, Shammout A, Horng S, et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci Data. 2023;10(1):1.
Emerson P, McPeake J, O’Neill A, Gilmour H, Forrest E, Puxty A, et al. The utility of scoring systems in critically ill cirrhotic patients admitted to a general intensive care unit. J Crit Care. 2014;29(6):1131.e1131-1136.
Zhou J, Zhou Y, Cao S, Li S, Wang H, Niu Z, et al. Multivariate logistic regression analysis of postoperative complications and risk model establishment of gastrectomy for gastric cancer: A single-center cohort report. Scand J Gastroenterol. 2016;51(1):8–15.
Rubio I, Osuchowski MF, Shankar-Hari M, Skirecki T, Winkler MS, Lachmann G, et al. Current gaps in sepsis immunology: new opportunities for translational research. Lancet Infect Dis. 2019;19(12):e422–36.
Venet F, Demaret J, Gossez M, Monneret G. Myeloid cells in sepsis-acquired immunodeficiency. Ann N Y Acad Sci. 2021;1499(1):3–17.
Bach JF. Infections and autoimmune diseases. J Autoimmun. 2005;25(Suppl):74–80.
Oud L. Epidemiology and outcomes of sepsis among hospitalizations with systemic lupus erythematosus admitted to the ICU: a population-based cohort study. J Intensive Care. 2020;8:3.
Zandman-Goddard G, Shoenfeld Y. SLE and infections. Clin Rev Allergy Immunol. 2003;25(1):29–40.
Simard JF, Rossides M, Gunnarsson I, Svenungsson E, Arkema EV. Infection hospitalisation in systemic lupus in Sweden. Lupus Sci Med. 2021;8(1):e000510.
Feldman CH, Hiraki LT, Winkelmayer WC, Marty FM, Franklin JM, Kim SC, et al. Serious infections among adult Medicaid beneficiaries with systemic lupus erythematosus and lupus nephritis. Arthritis Rheumatol. 2015;67(6):1577–85.
Mehta B, Pedro S, Ozen G, Kalil A, Wolfe F, Mikuls T, et al. Serious infection risk in rheumatoid arthritis compared with non-inflammatory rheumatic and musculoskeletal diseases: a US national cohort study. RMD Open. 2019;5(1):e000935.
Ananthakrishnan AN, McGinley EL. Infection-related hospitalizations are associated with increased mortality in patients with inflammatory bowel diseases. J Crohns Colitis. 2013;7(2):107–12.
Lichtenstein GR, Feagan BG, Cohen RD, Salzberg BA, Diamond RH, Chen DM, et al. Serious infections and mortality in association with therapies for Crohn’s disease: TREAT registry. Clin Gastroenterol Hepatol. 2006;4(5):621–30.
Carey IM, Critchley JA, DeWilde S, Harris T, Hosking FJ, Cook DG. Risk of infection in type 1 and type 2 diabetes compared with the general population: a matched cohort study. Diabetes Care. 2018;41(3):513–21.
Takeshita J, Shin DB, Ogdie A, Gelfand JM. Risk of serious infection, opportunistic infection, and herpes zoster among patients with psoriasis in the united Kingdom. J Invest Dermatol. 2018;138(8):1726–35.
Fouque-Aubert A, Jette-Paulin L, Combescure C, Basch A, Tebib J, Gossec L. Serious infections in patients with ankylosing spondylitis with and without TNF blockers: a systematic review and meta-analysis of randomised placebo-controlled trials. Ann Rheum Dis. 2010;69(10):1756–61.
Liu X, Arfman T, Wichapong K, Reutelingsperger CPM, Voorberg J, Nicolaes GAF. PAD4 takes charge during neutrophil activation: Impact of PAD4 mediated NET formation on immune-mediated disease. J Thromb Haemost. 2021;19(7):1607–17.
Xu J, Zhang X, Pelayo R, Monestier M, Ammollo CT, Semeraro F, et al. Extracellular histones are major mediators of death in sepsis. Nat Med. 2009;15(11):1318–21.
Dwivedi DJ, Toltl LJ, Swystun LL, Pogue J, Liaw KL, Weitz JI, et al. Prognostic utility and characterization of cell-free DNA in patients with severe sepsis. Crit Care. 2012;16(4):R151.
Rhodes A, Wort SJ, Thomas H, Collinson P, Bennett ED. Plasma DNA concentration as a predictor of mortality and sepsis in critically ill patients. Crit Care. 2006;10(2):R60.
Meng W, Paunel-Görgülü A, Flohé S, Hoffmann A, Witte I, MacKenzie C, et al. Depletion of neutrophil extracellular traps in vivo results in hypersusceptibility to polymicrobial sepsis in mice. Crit Care. 2012;16(4):R137.
Clark SR, Ma AC, Tavener SA, McDonald B, Goodarzi Z, Kelly MM, et al. Platelet TLR4 activates neutrophil extracellular traps to ensnare bacteria in septic blood. Nat Med. 2007;13(4):463–9.
Faibish M, Francescone R, Bentley B, Yan W, Shao R. A YKL-40-neutralizing antibody blocks tumor angiogenesis and progression: a potential therapeutic agent in cancers. Mol Cancer Ther. 2011;10(5):742–51.
Tizaoui K, Yang JW, Lee KH, Kim JH, Kim M, Yoon S, et al. The role of YKL-40 in the pathogenesis of autoimmune diseases: a comprehensive review. Int J Biol Sci. 2022;18(9):3731–46.
Kornblit B, Hellemann D, Munthe-Fog L, Bonde J, Strøm JJ, Madsen HO, et al. Plasma YKL-40 and CHI3L1 in systemic inflammation and sepsis-experience from two prospective cohorts. Immunobiology. 2013;218(10):1227–34.
Subesinghe S, Kleymann A, Rutherford AI, Bechman K, Norton S, Benjamin GJ. The association between lymphopenia and serious infection risk in rheumatoid arthritis. Rheumatology (Oxford). 2020;59(4):762–6.
Li Z, **ao Y, Zhang L. Application of procalcitonin, white blood cell count and neutrophil-to-lymphocyte ratio in the diagnosis of systemic lupus erythematosus with a bacterial infection. Ann Palliat Med. 2020;9(6):3870–6.
Guan Q, Gao X, Wang J, Sun Y, Shekhar S. Cytokines in Autoimmune Disease. Mediators Inflamm. 2017;2017:5089815.
Chousterman BG, Swirski FK, Weber GF. Cytokine storm and sepsis disease pathogenesis. Semin Immunopathol. 2017;39(5):517–28.
Rutherford AI, Subesinghe S, Hyrich KL, Galloway JB. Serious infection across biologic-treated patients with rheumatoid arthritis: results from the British society for rheumatology biologics register for rheumatoid arthritis. Ann Rheum Dis. 2018;77(6):905–10.
Luna G, Al** P, Burman J, Fink K, Fogdell-Hahn A, Gunnarsson M, et al. Infection risks among patients with multiple sclerosis treated with fingolimod, natalizumab, rituximab, and injectable therapies. JAMA Neurol. 2020;77(2):184–91.
Tudesq JJ, Cartron G, Rivière S, Morquin D, Iordache L, Mahr A, et al. Clinical and microbiological characteristics of the infections in patients treated with rituximab for autoimmune and/or malignant hematological disorders. Autoimmun Rev. 2018;17(2):115–24.
Shoor S. Risk of serious infection associated with agents that target T-cell activation and interleukin-17 and interleukin-23 cytokines. Infect Dis Clin North Am. 2020;34(2):179–89.
Barber MRW, Clarke AE. Systemic lupus erythematosus and risk of infection. Expert Rev Clin Immunol. 2020;16(5):527–38.
Ono S, Ueno C, Aosasa S, Tsujimoto H, Seki S, Mochizuki H. Severe sepsis induces deficient interferon-gamma and interleukin-12 production, but interleukin-12 therapy improves survival in peritonitis. Am J Surg. 2001;182(5):491–7.
Carrizosa JA, Aponte J, Cartagena D, Cervera R, Ospina MT, Sanchez A. Factors associated with mortality in patients with autoimmune diseases admitted to the intensive care unit in Bogota. Colombia Front Immunol. 2017;8:337.
van der Poll T, van de Veerdonk FL, Scicluna BP, Netea MG. The immunopathology of sepsis and potential therapeutic targets. Nat Rev Immunol. 2017;17(7):407–20.
**ao ZX, Miller JS, Zheng SG. An updated advance of autoantibodies in autoimmune diseases. Autoimmun Rev. 2021;20(2): 102743.
Zhou S, Yao Z. Roles of infection in Psoriasis. Int J Mol Sci. 2022;23(13):6955.
Acknowledgements
Genetic association estimates for autoimmune disease were obtained from a genome-wide association meta-analysis of the MRC-IEU study and FinnGen consortium. The authors thank all investigators for sharing these data.
Funding
This work was supported by the National Natural Scientific Foundation of China (82172154).
Author information
Authors and Affiliations
Contributions
Author HL, XP and WL collected and processed the data, as well as wrote this article. XS, ZW and SZ provided language help and writing assistance. SH, WS, and LC proof readed the article. SZ, XZ helped review the revised manuscript. JL and DC designed the study. The author(s) read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All studies had been approved by a relevant ethical review board and participants had given informed consent. Ethical approval was not required because of the public characteristics of the data of GWAS.
Consent for publication
Not applicable.
Competing interests
The authors have stated that they have no Competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, H., Pan, X., Zhang, S. et al. Association of autoimmune diseases with the occurrence and 28-day mortality of sepsis: an observational and Mendelian randomization study. Crit Care 27, 476 (2023). https://doi.org/10.1186/s13054-023-04763-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s13054-023-04763-5