Abstract
Background
Breast cancer patients exhibit various response patterns to neoadjuvant chemotherapy (NAC). However, it is uncertain whether diverse tumor response patterns to NAC in breast cancer patients can predict survival outcomes. We aimed to develop and validate radiomic signatures indicative of tumor shrinkage and therapeutic response for improved survival analysis.
Methods
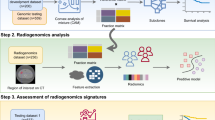
This retrospective, multicohort study included three datasets. The development dataset, consisting of preoperative and early NAC DCE-MRI data from 255 patients, was used to create an imaging signature-based multitask model for predicting tumor shrinkage patterns and pathological complete response (pCR). Patients were categorized as pCR, nonpCR with concentric shrinkage (CS), or nonpCR with non-CS, with prediction performance measured by the area under the curve (AUC). The prognostic validation dataset (n = 174) was used to assess the prognostic value of the imaging signatures for overall survival (OS) and recurrence-free survival (RFS) using a multivariate Cox model. The gene expression data (genomic validation dataset, n = 112) were analyzed to determine the biological basis of the response patterns.
Results
The multitask learning model, utilizing 17 radiomic signatures, achieved AUCs of 0.886 for predicting tumor shrinkage and 0.760 for predicting pCR. Patients who achieved pCR had the best survival outcomes, while nonpCR patients with a CS pattern had better survival than non-CS patients did, with significant differences in OS and RFS (p = 0.00012 and p = 0.00063, respectively). Gene expression analysis highlighted the involvement of the IL-17 and estrogen signaling pathways in response variability.
Conclusions
Radiomic signatures effectively predict NAC response patterns in breast cancer patients and are associated with specific survival outcomes. The CS pattern in nonpCR patients indicates better survival.
Similar content being viewed by others
Introduction
Neoadjuvant chemotherapy (NAC) is a fundamental component of treatment for early-stage breast cancer. Achieving a pathological complete response (pCR) after NAC has been consistently linked with superior long-term clinical outcomes, as evidenced by numerous studies [1,2,3]. In addition to serving as a treatment endpoint, early assessment during NAC is crucial for guiding clinical decisions and tailoring treatment strategies [4]. NAC generally involves several cycles, leading to various patterns of tumor shrinkage. Among these, concentric shrinkage (CS) is associated with a better prognosis and, in addition to pCR, may serve as an independent predictive factor for patient outcomes [5]. Nonetheless, the relationship between specific response patterns and prognosis, particularly among patients who do not achieve a pCR, is still not well understood. Elucidating the heterogeneity of these responses is essential for evaluating their prognostic value and improving treatment approaches for breast cancer patients.
Dynamic contrast-enhanced magnetic resonance imaging (DCE-MRI) is a highly specific and sensitive [30]. Our model capitalizes on these early treatment stages to provide critical prognostic information. This early prediction model holds promise for optimizing therapeutic regimens and advancing patient care by enabling clinicians to make more informed and timely decisions about treatment efficacy.
In our analysis, we delineated gene signaling pathways correlated with distinct patterns of treatment response. Genes such as BCL2 and ATF6B are involved in these pathways. Previous studies have demonstrated that downregulation of BCL2 correlates with improved disease-free survival (DFS) in breast cancer patients [31]. Additionally, the upregulation of ATF6B expression has been linked to favorable DFS outcomes [32]. Additionally, elevated PLA2G16 gene expression has been correlated with improved DFS, consistent with findings in the literature [33]. Our investigation not only confirmed a gradation of expression for these genes across patients who achieved pCR, nonpCR with CS, and nonpCR without CS but also aligned with a corresponding gradation in imaging feature changes observed in these groups. This correspondence underlines the credibility of radiomic signatures, such as sphericity, which is indicative of a tumor’s roundness and was found to be associated with a pCR and a CS pattern.
Our genomic analyses confirmed that survival differences among the distinct response pattern groups, as predicted by our model, are underpinned by biological variations. Notably, enrichment of the IL-17 signaling pathway aligns with its known contribution to the invasive progression of breast cancer [34, 35]. This observation is corroborated by evidence showing that IL-1β-induced IL-17 production by γδ T cells drives G-CSF-dependent neutrophil expansion and polarization in breast tumor models, processes critical for disease progression [36]. The complexity of ER signaling has been emphasized [37], with disruptions in ER cofactors and nongenomic mechanisms implicated in the metastasis of ER-positive breast cancer cells [28]. This discovery highlights the critical role of the estrogen pathway in the treatment of hormone-sensitive cancers and in the development of novel drug therapies [38].
Consistent with previous research findings, the extracellular matrix-receptor interaction pathway was notably prominent. This pathway included differentially expressed genes such as those in the THBS family, along with collagen and fibronectin genes, all of which are crucial in breast cancer pathogenesis [39]. These insights provide a valuable understanding of the molecular framework that dictates tumor behavior and the effectiveness of therapeutic interventions.
Our study has several limitations. First, the inclusion of diverse histopathological cancer subtypes, although representative of the clinical spectrum, adds variability to the chemotherapy protocols used. This variability may affect the pCR rate following NAC, potentially introducing selection bias. Second, the use of imaging data from multiple sources with differing imaging protocols may lead to biases in the development of the model, possibly affecting its predictive accuracy and applicability in various clinical settings. To mitigate these issues, future studies should focus on standardizing imaging protocols and taking histopathological variability into account. Such measures would enhance the validation process of the model, ensuring its dependability and usefulness across a wider range of clinical situations.
In summary, our investigation highlights the potential of noninvasive imaging as a prognostic tool for predicting responses to NAC and tumor shrinkage patterns. The study indicated that CS patterns correlate with better survival, particularly in patients who are less likely to reach a pCR. Additionally, gene expression analyses revealed distinct oncogenic pathways associated with various response patterns. These findings support the utility of imaging biomarkers in predicting therapeutic outcomes, emphasizing the role of radiomics in refining early prognosis and enabling personalized therapy.
Availability of data and codes
The gene expression data used in this study were sourced from the Gene Expression Omnibus (GEO) database and identified by the accession numbers GSE32603 and GPL14668. The ISPY-1 trial is available in TCIA on the website: (https://wiki.cancerimagingarchive.net/display/Public/ISPY1). The raw data from development dataset cannot be publicly available but can be obtained upon official request and ethical approval by contacting the corresponding author. PyRadiomics (https://pyradiomics.readthedocs.io/en/latest/) was used for radiomic feature analysis. The microenvironment cell population counter (MCP-counter) R package (http://github.com/ebecht/MCPcounter) was used for cell subpopulation estimation.
Abbreviations
- NAC:
-
Neoadjuvant chemotherapy
- pCR:
-
Pathological complete response
- AUC:
-
Area under the ROC curve
- DCE-MRI:
-
Dynamic contrast-enhanced magnetic resonance imaging
- OS:
-
Overall survival
- CS:
-
Concentric shrinkage
- RFS:
-
Recurrence-free survival
- HR:
-
Hazard ratio
References
Cortazar P, Zhang L, Untch M, Mehta K, Costantino JP, Wolmark N, Bonnefoi H, Cameron D, Gianni L, Valagussa P, et al. Pathological complete response and long-term clinical benefit in breast cancer: the CTNeoBC pooled analysis. Lancet. 2014;384:164–72.
von Minckwitz G, Untch M, Blohmer JU, Costa SD, Eidtmann H, Fasching PA, Gerber B, Eiermann W, Hilfrich J, Huober J, et al. Definition and impact of pathologic complete response on prognosis after neoadjuvant chemotherapy in various intrinsic breast cancer subtypes. J Clin Oncol. 2012;30:1796–804.
Chen D, Wang Q, Dong M, Chen F, Huang A, Chen C, Lu Y, Zhao W, Wang L. Analysis of neoadjuvant chemotherapy for breast cancer: a 20-year retrospective analysis of patients of a single institution. BMC Cancer. 2023;23:984.
Shien T, Iwata H. Adjuvant and neoadjuvant therapy for breast cancer. Jpn J Clin Oncol. 2020;50:225–9.
Fukada I, Araki K, Kobayashi K, Shibayama T, Takahashi S, Gomi N, Kokubu Y, Oikado K, Horii R, Akiyama F, et al. Pattern of tumor shrinkage during neoadjuvant chemotherapy is associated with prognosis in low-grade luminal early breast cancer. Radiology. 2018;286:49–57.
**ao J, Rahbar H, Hippe DS, Rendi MH, Parker EU, Shekar N, Hirano M, Cheung KJ, Partridge SC. Dynamic contrast-enhanced breast MRI features correlate with invasive breast cancer angiogenesis. NPJ Breast Cancer. 2021;7:42.
Fan M, Li H, Wang S, Zheng B, Zhang J, Li L. Radiomic analysis reveals DCE-MRI features for prediction of molecular subtypes of breast cancer. PLoS ONE. 2017;12: e0171683.
Reig B, Lewin AA, Du L, Heacock L, Toth HK, Heller SL, Gao Y, Moy L. Breast MRI for evaluation of response to neoadjuvant therapy. Radiographics. 2021;41:665–79.
Chamming’s F, Ueno Y, Ferre R, Kao E, Jannot AS, Chong J, Omeroglu A, Mesurolle B, Reinhold C, Gallix B. Features from computerized texture analysis of breast cancers at pretreatment MR imaging are associated with response to neoadjuvant chemotherapy. Radiology. 2018;286:412–20.
Fan M, Wu G, Cheng H, Zhang J, Shao G, Li L. Radiomic analysis of DCE-MRI for prediction of response to neoadjuvant chemotherapy in breast cancer patients. Eur J Radiol. 2017;94:140–7.
Fan M, Cui Y, You C, Liu L, Gu Y, Peng W, Bai Q, Gao X, Li L. Radiogenomic signatures of oncotype DX recurrence score enable prediction of survival in estrogen receptor-positive breast cancer: a multicohort study. Radiology. 2022;302:516–24.
Zhuang X, Chen C, Liu Z, Zhang L, Zhou X, Cheng M, Ji F, Zhu T, Lei C, Zhang J, et al. Multiparametric MRI-based radiomics analysis for the prediction of breast tumor regression patterns after neoadjuvant chemotherapy. Transl Oncol. 2020;13: 100831.
Huang YH, Zhu T, Zhang XL, Li W, Zheng XX, Cheng MY, Ji F, Zhang LL, Yang CQ, Wu ZY, et al. Longitudinal MRI-based fusion novel model predicts pathological complete response in breast cancer treated with neoadjuvant chemotherapy: a multicenter, retrospective study. Eclinicalmedicine. 2023. https://doi.org/10.1016/j.eclinm.2023.101899.
D’Angelo A, Orlandi A, Bufi E, Mercogliano S, Belli P, Manfredi R. Automated breast volume scanner (ABVS) compared to handheld ultrasound (HHUS) and contrast-enhanced magnetic resonance imaging (CE-MRI) in the early assessment of breast cancer during neoadjuvant chemotherapy: an emerging role to monitoring tumor response? Radiol Med. 2021;126:517–26.
** C, Yu H, Ke J, Ding P, Yi Y, Jiang X, Duan X, Tang J, Chang DT, Wu X, et al. Predicting treatment response from longitudinal images using multi-task deep learning. Nat Commun. 1851;2021:12.
Eun NL, Kang D, Son EJ, Park JS, Youk JH, Kim JA, Gweon HM. Texture analysis with 3.0-T MRI for association of response to neoadjuvant chemotherapy in breast cancer. Radiology. 2020;294:31–41.
Nadrljanski MM, Milosevic ZC. Tumor texture parameters of invasive ductal breast carcinoma in neoadjuvant chemotherapy: early identification of non-responders on breast MRI. Clin Imaging. 2020;65:119–23.
Hylton NM, Gatsonis CA, Rosen MA, Lehman CD, Newitt DC, Partridge SC, Bernreuter WK, Pisano ED, Morris EA, Weatherall PT, et al. Neoadjuvant chemotherapy for breast cancer: functional tumor volume by MR imaging predicts recurrence-free survival-results from the ACRIN 6657/CALGB 150007 I-SPY 1 TRIAL. Radiology. 2016;279:44–55.
Magbanua MJ, Wolf DM, Yau C, Davis SE, Crothers J, Au A, Haqq CM, Livasy C, Rugo HS, Investigators IST, et al. Serial expression analysis of breast tumors during neoadjuvant chemotherapy reveals changes in cell cycle and immune pathways associated with recurrence and response. Breast Cancer Res. 2015;17:73.
Newitt D, Hylton N, Team. obotI-SNaAT: multi-center breast DCE-MRI data and segmentations from patients in the I-SPY 1/ACRIN 6657 trials. Cancer Imag Arch. 2016. https://doi.org/10.7937/K9/TCIA.2016.HdHpgJLK.
Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 1991;19:403–10.
Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thurlimann B, Senn HJ, Panel Members. Strategies for subtypes-dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011;22:1736–47.
Hammond ME, Hayes DF, Wolff AC, Mangu PB, Temin S. American society of clinical oncology/college of american pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Oncol Pract. 2010;6:195–7.
van Griethuysen JJM, Fedorov A, Parmar C, Hosny A, Aucoin N, Narayan V, Beets-Tan RGH, Fillion-Robin JC, Pieper S, Aerts H. Computational radiomics system to decode the radiographic phenotype. Cancer Res. 2017;77:e104–7.
Fan M, Yuan W, Zhao W, Xu M, Wang S, Gao X, Li L. Joint prediction of breast cancer histological grade and Ki-67 expression level based on DCE-MRI and DWI radiomics. IEEE J Biomed Health Inform. 2020;24:1632–42.
Alinejad V, Dolati S, Motallebnezhad M, Yousefi M. The role of IL17B-IL17RB signaling pathway in breast cancer. Biomed Pharmacother. 2017;88:795–803.
Bastid J, Dejou C, Docquier A, Bonnefoy N. The emerging role of the IL-17B/IL-17RB pathway in cancer. Front Immunol. 2020;11:718.
Saha Roy S, Vadlamudi RK. Role of estrogen receptor signaling in breast cancer metastasis. Int J Breast Cancer. 2012;2012: 654698.
Yang D, Chen MB, Wang LQ, Yang L, Liu CY, Lu PH. Bcl-2 expression predicts sensitivity to chemotherapy in breast cancer: a systematic review and meta-analysis. J Exp Clin Cancer Res. 2013;32:105.
Goorts B, Dreuning KMA, Houwers JB, Kooreman LFS, Boerma EG, Mann RM, Lobbes MBI, Smidt ML. MRI-based response patterns during neoadjuvant chemotherapy can predict pathological (complete) response in patients with breast cancer. Breast Cancer Res. 2018;20:34.
Fabi A, Mottolese M, Di Benedetto A, Sperati F, Ercolani C, Buglioni S, Nistico C, Ferretti G, Vici P, Perracchio L, et al. p53 and BLC2 immunohistochemical expression across molecular subtypes in 1099 early breast cancer patients with long-term follow-up: an observational study. Clin Breast Cancer. 2020;20:e761–70.
Barron-Gallardo CA, Garcia-Chagollan M, Moran-Mendoza AJ, Delgadillo-Cristerna R, Martinez-Silva MG, Villasenor-Garcia MM, Aguilar-Lemarroy A, Jave-Suarez LF. A gene expression signature in HER2+ breast cancer patients related to neoadjuvant chemotherapy resistance, overall survival, and disease-free survival. Front Genet. 2022;13: 991706.
Yang X, Zhang Z, Jia X, Zhang Y, Mu T, Zhou B, Li L, Fu D, Hu X, **ong S. High expression of PLA2G16 is associated with a better prognosis in HER2-positive breast cancer. J Thorac Dis. 2017;9:1002–11.
Fabre JAS, Giustinniani J, Garbar C, Merrouche Y, Antonicelli F, Bensussan A. The interleukin-17 family of cytokines in breast cancer. Int J Mol Sci. 2018;19:3880.
Zhu X, Mulcahy LA, Mohammed RA, Lee AH, Franks HA, Kilpatrick L, Yilmazer A, Paish EC, Ellis IO, Patel PM, Jackson AM. IL-17 expression by breast-cancer-associated macrophages: IL-17 promotes invasiveness of breast cancer cell lines. Breast Cancer Res. 2008;10:R95.
Coffelt SB, Kersten K, Doornebal CW, Weiden J, Vrijland K, Hau CS, Verstegen NJM, Ciampricotti M, Hawinkels L, Jonkers J, de Visser KE. IL-17-producing gammadelta T cells and neutrophils conspire to promote breast cancer metastasis. Nature. 2015;522:345–8.
Khan MZI, Uzair M, Nazli A, Chen JZ. An overview on estrogen receptors signaling and its ligands in breast cancer. Eur J Med Chem. 2022;241: 114658.
Clusan L, Ferriere F, Flouriot G, Pakdel F. A basic review on estrogen receptor signaling pathways in breast cancer. Int J Mol Sci. 2023;24:6834.
Bao Y, Wang L, Shi L, Yun F, Liu X, Chen Y, Chen C, Ren Y, Jia Y. Transcriptome profiling revealed multiple genes and ECM-receptor interaction pathways that may be associated with breast cancer. Cell Mol Biol Lett. 2019;24:38.
Acknowledgements
Not applicable.
Funding
This research was funded by the National Natural Science Foundation of China under award numbers U21A20521 and 62271178 and the Natural Science Foundation of Zhejiang Province of China under award number LR23F010002.
Author information
Authors and Affiliations
Contributions
MF and LL wrote the manuscript and analyzed the data. DP, XC, ZL, SH, SX, CY and YG conducted the data collection and analysis. MF and LL had primary responsibility for the final content.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This study received ethical approval from Fudan University Cancer Hospital (2003214-7). Informed consent was obtained from all participants.
Consent for publication
All authors have read and approved the final manuscript.
Competing interests
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Fan, M., Wang, K., Pan, D. et al. Radiomic analysis reveals diverse prognostic and molecular insights into the response of breast cancer to neoadjuvant chemotherapy: a multicohort study. J Transl Med 22, 637 (2024). https://doi.org/10.1186/s12967-024-05487-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12967-024-05487-y