Abstract
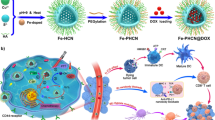
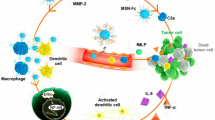
Lack of proper innate sensing inside the tumor microenvironment could reduce both innate and adaptive immunity, which remains a critical cause of immunotherapy failure in various tumor treatments. Double-stranded DNA (dsDNA) has been evidenced to be a promising immunostimulatory agent to induce type I interferons (IFN-Is) production for innate immunity activation through the cyclic GMP-AMP synthase (cGAS)-stimulator of interferon genes (STING) signaling pathway, yet the unsatisfactory delivery and susceptibility to nuclease degradation hindered its feasibility for further clinical applications. Herein, we report on the constructed tumor microenvironment-responsive DNA-based nanomedicine loaded by dendritic mesoporous organosilica nanoparticles (DMONs), which provide efficient delivery of dsDNA to induce intratumoral IFN-Is production for triggering innate sensing for enhanced anti-tumor immunotherapy. Extensive in vitro and in vivo evaluations have demonstrated the dramatic IFN-Is production induced by dsDNA@DMONs in both immune cells and tumor cells, which facilitates dendritic cells (DCs) maturation and T cells activation for eliciting the potent innate immune and adaptive immune responses. Desirable biosafety and marked therapeutic efficacy with a tumor growth inhibition (TGI) of 51.0% on the murine B16-F10 melanoma model were achieved by the single agent dsDNA@DMONs. Moreover, dsDNA@DMONs combined with anti-PD-L1 antibody further enhanced the anti-tumor efficacy and led to almost complete tumor regression. Therefore, this work highlighted the immunostimulatory DNA-based nanomedicine as a promising strategy for overcoming the resistance to immunotherapy, by promoting the IFN-Is production for innate immunity activation and remodeling the tumor microenvironment.
Graphical abstract

Similar content being viewed by others
Introduction
Cancer therapy has evolved considerably in recent decades, significantly improving the outcomes and quality of life for patients [1]. In addition to surgery, traditional cancer treatments such as chemotherapies, radiation therapies, and targeted therapies, are suffering from serious challenges such as severe systemic side effects and high treatment costs [2]. Recently, cancer immunotherapies have achieved remarkable strides in the past few years, as evidenced by the success of immune checkpoint blockade (ICB) therapies [3, 4]. However, only a small subset of cancer patients show durable clinical responses to ICB therapies and gain key outcome benefits like progression-free survival (PFS) and overall survival (OS) [5, 6]. Moreover, the magnitude of clinical benefits of ICB is highly variable both across different cancer types and between individual patients [7]. The resistance to ICB is in part due to the underlying immunosuppressive nature of the “cold” tumor, which is characterized by the low infiltration of tumor-infiltrating lymphocytes (TILs), low tumor mutational burden (TMB) and low major histocompatibility complex (MHC) class I expression in the tumor microenvironment (TME) [8, 9].
As an emerging efficient cancer immunotherapeutic modality, immunostimulatory therapies that activate innate sensing pathways are of great promise for overcoming the resistance to tumor immunotherapies by remodeling the TME [10,11,12]. As the first line of host defense against invading pathogens or dangers, the innate immune response initiation is equipped with pattern-recognition receptors (PRRs) which recognize pathogen-associated molecular patterns (PAMPs) or damage-associated molecular patterns (DMAPs) [13, 14]. The cyclic GMP-AMP synthase (cGAS)-stimulator of interferon genes (STING) signaling pathway, sensing cytoplasmic double-stranded DNA (dsDNA) of both foreign and self-origin, has been established as an essential mechanism implicated in the innate sensing [15,16,17]. Briefly, cGAS activated by the cytoplasmic dsDNA, catalyzes the synthesis of 2′-3′ cyclic GMP-AMP (cGAMP) and activates the adaptor protein STING, inducing the production of type I interferons (IFN-Is) and secretion of pro-inflammatory cytokine to trigger the innate immune response [18]. Growing evidence has indicated that IFN-Is play a key role in promoting the activation and maturation of dendritic cells (DCs), enhancing the antigen presentation for T cell priming, facilitating tumor immune infiltration, and in turn eliciting the anti-tumor immune responses [19]. Although dsDNA or other cyclic dinucleotides have been evidenced to be efficient immunostimulatory agents for cGAS-STING activation and IFN-Is production for eliciting anti-tumor immune responses, the desirable delivery and exploitation of cyclic dinucleotides remain very challenging due to their negatively charged nature and susceptibility to nuclease degradation [19, 20].
Traditional DNA delivery systems like lipid nanoparticles or cationic polymers often face severe challenges of poor stability and possible fast clearance, which is unfavorable for the effective protection and in vivo delivery of DNA [21, 22]. Dendritic mesoporous silica nanoparticles have motivated extensive research interest as versatile drug delivery systems of a broad range of drugs such as small molecules, proteins and genes, benefiting from their unique central radial pore structure with high surface area for enhanced drug loading efficiency, as well as desirable biocompatibility and feasible surface modification [23, 24]. Especially, dendritic mesoporous organosilica nanoparticles (DMONs) with structurally integrated disulfide bonds, which can be cleaved by glutathione (GSH), can achieve a TME-responsive biodegradation and drug release owing to the high expression level of GSH in cancer cells [25].
Herein, we report on the rationally designed TME-responsive immunostimulatory nanomedicine dsDNA@DMONs for efficient tumor immunotherapy, based on the DMONs with small particle size and large pore size, which facilitate the efficient intratumoral delivery of dsDNA in a TME-responsive way, for eliciting innate immunity and anti-tumor responses. The dsDNA@DMONs release dsDNA in response to GSH, which triggers innate sensing via the cGAS-STING pathway, inducing the production of IFN-Is to promote DCs maturation, antigen-priming, and T cell activation. As a result, the potent activation of adaptive anti-tumor T cell response can be achieved by the dsDNA@DMONs, and the therapeutic efficacy can be further enhanced in combination with ICB therapy (Scheme 1).
Results and discussion
Synthesis and characterization of dsDNA@DMONs nanomedicine
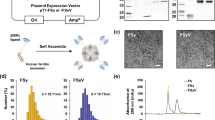
DMONs were synthesized by a triethanolamine (TEA)-catalyzed co-condensation reaction of tetraethyl orthosilicate (TEOS) and bis[3-(triethoxysilyl)propyl] tetrasulfide (BTES) based on the Stöber mechanism and sol–gel chemistry, with sodium salicylate as structural directing agents [26]. The introduction of BTES as an organosilica precursor facilitates the integration of the –S–S–S–S– functional group into the silica skeleton backbone, which in turn improves the biocompatibility and physiological stability of DMONs. Notably, benefiting from the redox reactivity of the disulfide group to GSH, the DMONs were expected to possess specific biodegradability in TME where the GSH is relatively higher than normal tissues [ The reagents used included QUANTI-Luc™ (InvivoGen), DAPI-containing anti-fade medium (P36962, Thermo Fisher Scientific), Gelstain (GS101, Transgen), Trans2K Plus DNA Marker (BM111, Transgen), Direct-zol RNA Miniprep Plus Kit (R2070, ZYMO RESEARCH), PrimeScriptTM RT reagent Kit with gDNA Eraser (RR047A, Takara), DreamTaq polymerase (EP0701, Thermo Scientific), GeneJET PCR purification kit (K0701, Thermo Scientific), 50 × Tris–acetate-EDTA (TAE, ST716, Beyotime Biotechnology), Radio Immunoprecipitation Assay buffer (RIPA, PC101, Epizyme), phosphatase (GRF102; Epizyme), protease inhibitors (GRF101, Epizyme), TRIzol Reagent (15,596,026, Invitrogen). PierceTM BCA Protein Assay Kit (23,225, Thermo Scientific), SDS-PAGE Sample Buffer (P0015F, Beyotime Biotechnology), Multicolor Prestained Protein Ladder (WJ103, Epizyme), BD Pharmingen™ Leukocyte Activation Cocktail (550,583, BD Pharmingen), Fixation/Permeabilization Solution Kit with BD GolgiPlug (555,028, BD Pharmingen), Foxp3/Transcription Factor Staining Buffer Set (00–5523-00, Thermo Scientific), Collagenase I (C10130, Sigma), Hyaluronidase (H3506, Sigma), 25 μg/mL DNase I (DN25-100, Sigma), Triethanolamine (TEA, V900257, Sigma), Hexadecyl trimethyl ammonium bromide(CTAB, H5882, Sigma), Sodium salicylate (NaSal, 54–21-7, Sigma), Tetraethyl orthosilicate (TEOS, 78–10-4, Sigma), Bis[3-(triethoxysilyl)propyl] tetrasulfide (BTES, 40,372–72-3, Sigma-Aldrich), 3-Aminopropyl triethoxysilane (APTES, 919–30-2, Sigma-Aldrich), sulfo-Cyanine5 NHS ester (43,320, Lumiprobe), DreamTaq DNA Polymerase (EP0705, Thermo Scientific), GeneJET PCR Purification Kit (K0701, Thermo Scientific), H-151 (HY-112693, MedChemExpress), Lipofectamine 2000 Transfection Reagent (11,668,019, Thermo Scientific), Simulated body fluid (SBF, R24165, Shanghai yuanye Bio-Technology). Anti-PD-L1 (ab213480, Abcam), anti-GAPDH antibody (2118, CST), anti-Phospho-TBK1/NAK antibody (5483, CST), HRP-conjugated secondary antibodies (7074, CST), anti-PD-L1(BE0101, BioXcell), anti-mouse CD16/32 antibody (553,141, BD Pharmingen), Fixable Viability Stain (565,388, BD Pharmingen), anti-CD80-BV421(562,611, BD Pharmingen), anti-CD86-PE-Cy7(560,582, BD Pharmingen), anti-mouse PD-1-PE (135,206, BioLegend), anti-mouse LAG3-PE-Cy7 (125,226, BioLegend), anti-mouse TIM3-BV421 (119,723, BioLegend), anti-mouse FOXP3-BV421 (126,419, BioLegend), anti-mouse Ki67-PE-CY7 (652,426, BioLegend), anti-mouse PD-L1-PE (124,308, BioLegend), anti-mouse CD11b-FITC (101,206, BioLegend), anti-mouse Ly6G-PE (551,461, BD Pharmingen), anti-mouse Ly6C-PE-Cy7 (128,018, BioLegend), anti-mouse F4/80-BV421 (123,132, BioLegend), anti-mouse CD206-APC (141,708, BioLegend), anti-mouse TNFα-PE (554,419, BD Pharmingen), anti-mouse CD3-BV605 (100,237, BioLegend), anti-mouse CD11C-APC (117,310, BioLegend), anti-mouse MHC-II-BV421 (107,632, BioLegend), anti-mouse NK1.1-PE-Cy7 (552,878, BD Pharmingen), anti-mouse CD80-PE (552,769, BD Pharmingen). MC38, B16-F10, 4T1, MDA-MB-231, and A375 cells were acquired from ATCC. Panc02 cells were obtained from the Cell Bank of the Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences (SIBS, CAS). RAW-Lucia ISG cells with an interferon regulatory factor (IRF)-inducible Lucia luciferase reporter construct were acquired from Invivogen. Mouse dendritic cells 2.4 (DC2.4) were kindly provided by Dr. **g Qian from Jiangsu Academy of Agricultural Sciences (JAAS). All cell lines were routinely tested using a mycoplasma contamination kit (R&D) and cultured in the indicated medium following the manufacturer’s instruction and supplemented with 10% heat-inactivated fetal bovine serum, 100 U/ml penicillin, and 100 U/ml streptomycin. All cells were kept at 37 °C with 5% CO2. Female C57BL/6 mice, six weeks old, were obtained from the Shanghai Lingchang Biotechnology Co., LTD. Mice were housed in an SPF animal facility in temperature-controlled rooms at 21 °C, with 45–65% relative humidity and 12-h light/dark cycles at the Shanghai Jiao Tong University School of Medicine. The animal experiments were performed using protocols approved by the Institutional Animal Care and Use Committee. 0.068 g of TEA was added to 25 mL of water and stirred at 300 rpm for 0.5 h at 80 °C, after which 380 mg of CTAB and 168 mg of NaSal were added and stirred for another 1 h. 2 mL of TEOS and 2 mL of BTES were added and continued to stir at 80 °C for 2 h. The products were collected by centrifugation at 20,000 rpm at the end of the reaction, after which the residual reactants were removed by washing three times using water and ethanol. The DMONs were extracted with a mixture solution of HCl and anhydrous ethanol (V HCl: V ethanol = 1: 9) at 80 °C for 12 h three times to remove the CTAB. The obtained DMONs were dried under vacuum at room temperature. 100 mg of DMONs were dispersed into 100 mL of a mixture of ethanol and water (V water: V ethanol = 1: 1), and 1 mL of APTES was added and continued to stir at 80 °C for 12 h. The reaction was carried out for 12 h, after which DMONs-NH2 was obtained by centrifugation using ethanol three times. Double-stranded DNA molecules were generated by PCR amplification with DreamTaq polymerase using pcDNA3.1 as a template as described previously [37]. The length of the PCR products is 94 base pairs. The PCR products were purified using a GeneJET PCR purification kit. Forward primer: 5’-CGATGTACGGGCCAGATATACG-3’; Reverse primer: 5’-CATATATGGGCTATGAACTAATGACC-3’. DMONs-NH2 (Si: 100 ng/mL) was dispersed into 1 mL of dsDNA aqueous solution with different mass ratios of dsDNA to Si, and stirred overnight at room temperature, after which unloaded dsDNA was removed by centrifugation. The dsDNA concentration of the supernatant was detected by Nanodrop assay to calculate the loading capacity of dsDNA. The in vitro dsDNA release profile of dsDNA@DMONs was studied in SBF (pH 7.4) with different GSH concentrations of 0, 5 and 10 mM at 37 °C with a shaking speed of 200 rpm. The concentration of dsDNA released in the supernatant at scheduled timepoints was determined using Nanodrop and followed by the replacement of buffer with fresh SBF with corresponding GSH concentrations. Transmission electron microscopy (TEM) images were acquired on a JEOL-2100F transmission electron microscope. Scanning electron microscopy (SEM) images and corresponding element map** scanning were acquired on a field-emission Magellan 400 microscope (FEI Company, USA). The quantitative analysis of the Si element was determined by inductively coupled plasma mass spectrometry (ICP-MS, NexION 2000B, PerkinElmer, US). Confocal laser scanning microscopy (CLSM) images were acquired by FV1000 (SP8, Leica, US). Nitrogen adsorption–desorption isotherms were recorded at liquid nitrogen temperature with an ASAP 2020 adsorption analyzer (Micromeritics). Murine IFN-Is activity was measured in RAW-Lucia ISG cells. Raw-Lucia ISG cells were seeded at a density of 30,000 cells per well in a 96-well flat bottom plate (Corning). After adhering to the plate, Raw-Lucia ISG cells were treated with different concentrations of dsDNA@DMONs for 24 h. Subsequently, 50 μL supernatant was transferred to a 96-well opaque white plate, and 50 μL QUANTI-Luc was added to detect luciferase activity. After mixing the sample with 6 × loading dye, the sample was loaded into the well of 1% agarose gel containing Gelstain, and the gel was marked with a DNA marker. 1 × TAE buffer was used as the running buffer, and the electrophoresis was performed at a condition of 200 mA for 30 min using a nucleic acid electrophoresis apparatus. After electrophoresis, the gel was imaged using the ChemiDoc MP gel imaging system produced by Biorad. Total RNA was extracted from cells using the TRIzol Reagent and the Direct-zol RNA Miniprep Plus Kit according to the manufacturer's instructions. The RNA quality and quantity were evaluated using a microvolume spectrophotometer. Reverse-transcribed with the PrimeScriptTM RT reagent Kit with gDNA Eraser according to the manufacturer’s instructions. RT-qPCR was performed using SYBR Green and primer pairs designed to target Ifnb, Cxcl10, Isg15, Tapbp, Tap1, Tap2, B2m, Psmb5 and β-actin (Primer sequences are listed in Supplementary Table 1). The relative mRNA expression levels were determined using the comparative CT method, with normalization to the housekee** gene β-actin, and the data was analyzed as fold induction compared to the control sample. Immunofluorescence and confocal analysis were performed as described before [38]. Sulfo-Cy5.5 NHS ester stock solution was prepared by adding 1 mL of DMSO to 1 mg of sulfo-Cy5.5 NHS ester powder. Then 100 μL of sulfo-Cy5.5 NHS ester stock solution was added to 1 mL of DMONs-NH2 and incubated overnight on a shaker protected from light. Afterward, the DMONs-Cy5.5 was obtained by centrifugation two times. B16-F10 or DC2.4 cells were cultured on glass-bottomed wells (150,680, Nunc) and treated with DMONs-NH2 for 30 min, 2 h or 6 h. After that, cells were fixed in 4% PFA in PBS for 10 min, washed twice with PBS, and then stained with DAPI for 10 min. At least eight representative images were taken for each sample using a Leica TCS SP8 confocal laser scanning microscope (CLSM). For western blotting experiments, cells were cultured with indicated treatments followed by being lysed with RIPA buffer supplemented with phosphatase and protease inhibitors and centrifuged for 15 min at 14,000 g in 4 °C. The protein concentration was determined by the PierceTM BCA Protein Assay Kit, and the protein samples were denatured using SDS-PAGE buffer by heating at 100 °C for 5 min. Subsequently, Equal protein amounts of samples were separated on 10% PAGE gels with a Multicolor Prestained Protein Ladder and then transferred to nitrocellulose membranes (Millipore) by the Trans-Blot Turbo Transfer System (Bio-Rad). The membranes were blocked in 5% BSA in TBST for 1 h at room temperature before incubation with primary antibodies at 4 °C overnight. The membranes were washed with TBST three times and incubated with HRP-conjugated secondary antibodies for 2 h at room temperature. Protein images were captured with the Tanon 5200 Series Fully Automated Chemiluminescence/Fluorescence Image Analysis System. For in vivo tumor models, 1 × 106 B16-F10 cells in 100 μL DMEM per mouse were used and injected subcutaneously into the flank of mice. The tumor volume was monitored every other day by measuring the length (a) and width (b), and the tumor volume was calculated to be 1/2a × b2. Mouse body weight was monitored during the therapeutic period. The mice in each group were intratumorally (i.t.) administrated with 50 µL/dose PBS, DMONs, dsDNA or dsDNA@DMONs for a total of four doses when the tumor volume was about 60 mm3. For the anti-PD-L1(Clone:10F.9G2) antibody, mice were administered with intraperitoneal injections of anti-PD-L1 antibody (q3d, 100 μg per mouse, a total of three doses). The study endpoint for maximum tumor volume was approximately 2000 mm3. Tumor tissues were minced and enzymatically digested with 2 mg/mL collagenase I supplemented with 1 mg/mL hyaluronidase and 25 μg/mL DNase I for 30 min at 37 °C, to acquire a single-cell suspension. The dissociated cell suspensions were passed through a 70 μm filter, counted, and resuspended in 100 μL PBS to reach a cell density of 5 × 106 cells per sample. Then the cells were blocked with anti-mouse CD16/32 antibody for 30 min. After washing with FACS buffer twice, cells were stained with Fixable Viability Stain for 15 min on ice in the dark. After washing, cells were stained with antibody mix (already titrated antibody concentrations) for 30 min on ice in the dark. After washing, cells were resuspended with 300 μL FACS buffer. For Ki67 and FOXP3 staining, cells were treated with eBioscience™ Foxp3 / Transcription Factor Staining Buffer Set by following the instructions and then stained with titrated antibodies. After washing, cells were resuspended with 300 μL FACS buffer. For intranuclear TNFα staining, cells were stimulated with Leukocyte Activation Cocktail according to the instructions before permeabilization with Fixation/Permeabilization Solution Kit, and then stained with titrated anti-TNFα antibody. Data were acquired with a FACSAriaTM III flow cytometer and analyzed by FlowJo software. Mice were euthanized at the final time point. The kidney, liver, spleen, heart and lung tissues were eviscerated, fixed in 4% formaldehyde and cut into 7-μm formalin-fixed paraffin-embedding (FFPE) slides. The FFPE slides were then stained with hematoxylin and eosin and then images were taken on an Olympus IX73 microscope.Methods
Materials and reagents
Antibodies
Cells
Mice
Synthesis of DMONs
Preparation of dsDNA
dsDNA loading
In vitro dsDNA release
Material characterization
IFN-I activity reporter assay
Agarose gel electrophoresis
RNA extraction and quantitative real-time PCR
Immunofluorescence analysis
Protein extraction and immunoblotting
In vivo tumor models
Flow cytometry analysis of the tumor microenvironment
H&E staining
Availability of data and materials
The data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Gatenby, RA, et al. Integrating evolutionary dynamics into cancer therapy. Nat Rev Clin Oncol. 2020;17:675–686.
Miller KD, et al. Cancer treatment and survivorship statistics. Cancer J Clin. 2022;72:409–36.
Kon E, Benhar I. Immune checkpoint inhibitor combinations: Current efforts and important aspects for success. Drug Resist Updates. 2019;45:13–29. https://doi.org/10.1016/j.drup.2019.07.004.
Kroemer G, Zitvogel L. Immune checkpoint inhibitors. J Exp Med. 2021;218: e20201979. https://doi.org/10.1084/jem.20201979.
Schoenfeld AJ, Hellmann MD. Acquired resistance to immune checkpoint inhibitors. Cancer Cell. 2020;37:443–55.
Jenkins RW, Barbie DA, Flaherty KT. Mechanisms of resistance to immune checkpoint inhibitors. Br J Cancer. 2018;118:9–16. https://doi.org/10.1038/bjc.2017.434.
Kraehenbuehl L, Weng C-H, Eghbali S, Wolchok JD, Merghoub T. Enhancing immunotherapy in cancer by targeting emerging immunomodulatory pathways. Nat Rev Clin Oncol. 2022;19:37–50. https://doi.org/10.1038/s41571-021-00552-7.
Zabransky DJ, Yarchoan M, Jaffee EM. Strategies for heating up cold tumors to boost immunotherapies. Ann Rev Cancer Biol. 2023;7:149–70. https://doi.org/10.1146/annurev-cancerbio-061421-040258.
Bonaventura P, et al. Cold tumors: a therapeutic challenge for immunotherapy. Front Immunol. 2019;10:168. https://doi.org/10.3389/fimmu.2019.00168.
Demaria O, et al. Harnessing innate immunity in cancer therapy. Nature. 2019;574:45–56.
Moynihan KD, Irvine DJ. Roles for innate immunity in combination immunotherapies. Can Res. 2017;77:5215–21.
Iannello A, Thompson TW, Ardolino M, Marcus A, Raulet DH. Immunosurveillance and immunotherapy of tumors by innate immune cells. Curr Opin Immunol. 2016;38:52–8. https://doi.org/10.1016/j.coi.2015.11.001.
Shekarian T, et al. Pattern recognition receptors: immune targets to enhance cancer immunotherapy. Ann Oncol. 2017;28:1756–66. https://doi.org/10.1093/annonc/mdx179.
Hernandez C, Huebener P, Schwabe RF. Damage-associated molecular patterns in cancer: a double-edged sword. Oncogene. 2016;35:5931–41. https://doi.org/10.1038/onc.2016.104.
Zhang X, Bai X-C, Chen ZJ. Structures and mechanisms in the cGAS-STING innate immunity pathway. Immunity. 2020;53:43–53.
Ding C, Song Z, Shen A, Chen T, Zhang A. Small molecules targeting the innate immune cGAS-STING-TBK1 signaling pathway. Acta Pharmaceutica Sinica B. 2020;10:2272–98. https://doi.org/10.1016/j.apsb.2020.03.001.
Samson N, Ablasser A. The cGAS–STING pathway and cancer. Nature Cancer. 2022;3:1452–63. https://doi.org/10.1038/s43018-022-00468-w.
Cattolico C, Bailey P, Barry ST. Modulation of Type I Interferon Responses to Influence Tumor-Immune Cross Talk in PDAC. Front Cell Dev Biol. 2022;10: 816517. https://doi.org/10.3389/fcell.2022.816517.
Zitvogel L, Galluzzi L, Kepp O, Smyth MJ, Kroemer G. Type I interferons in anticancer immunity. Nat Rev Immunol. 2015;15:405–14. https://doi.org/10.1038/nri3845.
Singh A, Singh N. Effect of salt concentration on the stability of heterogeneous DNA. Physica A. 2015;419:328–34. https://doi.org/10.1016/j.physa.2014.10.029.
Mendes BB, et al. Nanodelivery of nucleic acids. Nat Rev Methods Primers. 2022. https://doi.org/10.1038/s43586-022-00104-y.
Patra JK, et al. Nano based drug delivery systems: recent developments and future prospects. J Nanobiotechnol. 2018;16:71. https://doi.org/10.1186/s12951-018-0392-8.
Jafari S, et al. Mesoporous silica nanoparticles for therapeutic/diagnostic applications. Biomed Pharmacother. 2019;109:1100–11. https://doi.org/10.1016/j.biopha.2018.10.167.
Zhou Y, et al. Mesoporous silica nanoparticles for drug and gene delivery. Acta Pharmaceutica Sinica B. 2018;8:165–77. https://doi.org/10.1016/j.apsb.2018.01.007.
Du X, et al. Disulfide-bridged organosilica frameworks: designed, synthesis, redox-triggered biodegradation, and nanobiomedical applications. Adv Func Mater. 2018;28:1707325. https://doi.org/10.1002/adfm.201707325.
Gao S, et al. Nanocatalytic tumor therapy by biomimetic dual inorganic nanozyme-catalyzed cascade reaction. Adv Sci. 2019;6:1801733. https://doi.org/10.1002/advs.201801733.
**e J, et al. Emerging strategies of nanomaterial-mediated tumor radiosensitization. Adv Mater. 2019;31:1802244. https://doi.org/10.1002/adma.201802244.
Zhou Z, et al. GSH depletion liposome adjuvant for augmenting the photothermal immunotherapy of breast cancer. Sci Adv. 2020;6:4373.
Hawes MC, Wen F, Elquza E. Extracellular DNA: A Bridge to Cancer. Can Res. 2015;75:4260–4. https://doi.org/10.1158/0008-5472.CAN-15-1546.
Grieves JL, et al. Exonuclease TREX1 degrades double-stranded DNA to prevent spontaneous lupus-like inflammatory disease. Proc Natl Acad Sci U S A. 2015;112:5117–22. https://doi.org/10.1073/pnas.1423804112.
Gardner A, Ruffell B. Dendritic cells and cancer immunity. Trends Immunol. 2016;37:855–65.
Yu R, Zhu B, Chen D. Type I interferon-mediated tumor immunity and its role in immunotherapy. Cell Mol Life Sci. 2022;79:191. https://doi.org/10.1007/s00018-022-04219-z.
Schreiber RD, Old LJ, Smyth MJ. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science. 2011;331:1565–70. https://doi.org/10.1126/science.1203486.
Chen DS, Mellman I. Oncology meets immunology: the cancer-immunity cycle. Immunity. 2013;39:1–10. https://doi.org/10.1016/j.immuni.2013.07.012.
Huang KC-Y, et al. DNMT1 constrains IFNβ-mediated anti-tumor immunity and PD-L1 expression to reduce the efficacy of radiotherapy and immunotherapy. OncoImmunology. 2021;10:1989790. https://doi.org/10.1080/2162402X.2021.1989790.
Herbst RS, et al. Predictive correlates of response to the anti-PD-L1 antibody MPDL3280A in cancer patients. Nature. 2014;515:563–7. https://doi.org/10.1038/nature14011.
Luecke S, et al. cGAS is activated by DNA in a length-dependent manner. EMBO Rep. 2017;18:1707–15. https://doi.org/10.15252/embr.201744017.
Daemen S, Chan MM, Schilling JD. Comprehensive analysis of liver macrophage composition by flow cytometry and immunofluorescence in murine NASH. STAR Protocols. 2021;2: 100511. https://doi.org/10.1016/j.xpro.2021.100511.
Funding
This study was supported by Grants from the National Natural Science Foundation of China (81972820 and 82030099), the National Key R&D Program of China (2022YFD2101500), the Science and Technology Commission of Shanghai Municipality (22DZ2303000), Natural Science Foundation of Shanghai (23ZR1435900), Innovative research team of high-level local universities in Shanghai and Shanghai Jiao Tong University Key Program of Medical Engineering (YG2021ZD01).
Author information
Authors and Affiliations
Contributions
JL, XH, SG and YY performed in vivo and in vitro experiments. JL and XH analyzed the data. JL, SG, and XL wrote the manuscript. XL and HW supervised the study.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
All animal experimental protocols were approved by the Institutional Animal Care and Use Committee (IACUC, No. A-2020-001) of Shanghai Jiao Tong University School of Medicine.
Consent for publication
All authors agree to publish this manuscript.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
Supplementary figures and tables.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, J., Han, X., Gao, S. et al. Tumor microenvironment-responsive DNA-based nanomedicine triggers innate sensing for enhanced immunotherapy. J Nanobiotechnol 21, 382 (2023). https://doi.org/10.1186/s12951-023-02132-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12951-023-02132-6