Abstract
Background
Chordomas are rare, slow-growing and locally aggressive bone sarcomas. At present, chordomas are difficult to manage due to their high recurrence rate, metastasis tendency and poor prognosis. The underlying mechanisms of chordoma tumorigenesis and progression urgently need to be explored to find the effective therapeutic targets. Our previous data demonstrates that EGFR plays important roles in chordoma development and CKLF-like MARVEL transmembrane domain containing (CMTM)3 suppresses gastric cancer metastasis by inhibiting the EGFR/STAT3/EMT signaling pathway. However, the roles and mechanism of CMTM3 in chordomas remain unknown.
Methods
Primary chordoma tissues and the paired adjacent non-tumor tissues were collected to examine the expression of CMTM3 by western blot. The expression of CMTM3 in chordoma cell lines was tested by Real-time PCR and western blot. CCK-8 and colony forming unit assay were performed to delineate the roles of CMTM3 in cell proliferation. Wound healing and Transwell assays were performed to assess cell migration and invasion abilities. A xenograft model in NSG mice was used to elucidate the function of CMTM3 in vivo. Signaling pathways were analyzed by western blot and IHC. RNA-seq was performed to further explore the mechanism regulated by CMTM3 in chordoma cells.
Results
CMTM3 expression was downregulated in chordoma tissues compared with paired normal tissues. CMTM3 suppressed proliferation, migration and invasion of chordoma cells in vitro and inhibited tumor growth in vivo. CMTM3 accelerated EGFR degradation, suppressed EGFR/STAT3/EMT signaling pathway, upregulated TP53 expression and enriched the TP53 signaling pathway in chordoma cells.
Conclusions
CMTM3 inhibited tumorigenesis and development of chordomas through activating the TP53 signaling pathway and suppressing the EGFR/STAT3 signaling pathway, which suppressed EMT progression. CMTM3 might be a potential therapeutic target for chordomas.
Similar content being viewed by others
Background
Chordomas, derived from embryonic remnants of the notochord, are rare and slow-growing primary malignant tumors of the spine with an incidence rate of 1–4% of all bone cancers [1, 2]. The major treatment for chordomas is surgery and postoperative radiotherapy as chordomas are refractory to cytotoxic chemotherapeutics [3]. However, it is difficult to treat chordomas due to their local destructive behavior and their preference for growing near the critical skull base, lumbosacral nerves and blood vessels [4]. In addition, high recurrence (> 50% of patients) [5] and metastasis are major causes of treatment failure in chordomas [6, 7]. Herein, it is necessary to study the mechanism of chordoma tumorigenesis and progression and find the effective therapeutic targets for chordoma treatment.
Epidermal growth factor receptor (EGFR), a member of tyrosine kinase receptors, is generally overexpressed in chordomas [8]. EGFR affects tumor growth and metastasis [9]. Our previous study shows that phosphorylated EGFR plays important roles in chordomas progress [10]. EGFR phosphorylation triggers downstream signaling pathways such as signal transducer and activator of transcription 3 (STAT3) [11], AKT [12] and ERK1/2 [13] pathways. The activation of these signaling pathways further drives epithelial–mesenchymal transition (EMT) [14], which is a critical process in cancer metastasis [15, 16]. However, only a few studies demonstrate that EMT is involved in the pathogenesis of chordomas [17]. STAT3 activation also decreases the expression of TP53 and suppresses the TP53 signaling pathway [18], which also responds to stress signals and regulates the expression of its target genes leading to various cellular responses to prevent tumorigenesis [19, 20] as a transcription factor. At present, only Ma’ group explicitly proposes that the TP53 (alias: p53) signaling pathway plays a role in chordoma development [21]. It is necessary and crucial to clarify how the EGFR and TP53 signaling pathway make contributions to chordoma tumorigenesis and progression.
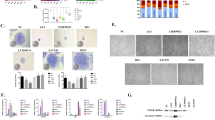
CKLF-like MARVEL transmembrane domain containing (CMTM)3 is a member of the CMTM family [22, 23]. It is located on 16q22.1, an important tumor suppressor locus with tumor suppressive properties [24]. CMTM3 is frequently reduced in multiple types of cancer, such as prostate cancer [25, 26] and renal cell carcinoma [42]. We overexpressed CMTM3 using adenovirus and investigated the whole transcriptome of JHC7 cells using Illumina RNA-Seq technology to screen the potential signaling pathway involved in CMTM3-suppressed cell proliferation. The results showed that there were 479 differently expressed genes (DEGs) between JHC7-MOCK and JHC7-CMTM3 cells with 232 upregulated genes and 247 downregulated genes (padj < 0.05, Fig. 5A, Additional files 2, 3). Among these DEGs, 68 genes were significantly upregulated and 18 genes were downregulated (Table 3, Fig. 5B, padj < 0.001).
CMTM3 induces changes in gene expression profiles. A The number of different expressed genes (padj < 0.05, MOI: 100). B A heat map summary of the gene expression values of JHC7-MOCK and JHC7-CMTM3 cells. Red indicates high and green indicates low gene expression values (padj < 0.001). C KEGG enrichment analysis. The TOP seven upregulated signaling pathways are shown of CMTM3-vs-MOCK in JHC7 cells (padj < 0.05). D The expression of TP53 was detected in CMTM3 overexpressed chordoma cell lines by Real-time PCR. E The expression of TP53 was detected in CMTM3 overexpressed JHC7 cells by western blot. F Up, the transfection efficiency of the CMTM3-siRNA was detected in JHC7 cells by western blot. 100 μg of total protein lysates were loaded; Down. the expression of TP53 in CMTM3 silenced JHC7 cells was detected by western blot. Data were representative of three independent experiments
We observed that the expression of the well-known tumor suppressor gene TP53, a target of STAT3 was obviously unregulated by overexpression of CMTM3 (ranking first besides CMTM3). In addition, The KEGG pathway enrichment analysis of JHC7 cells showed that the upregulated TP53 was involved in most of the top significant enrichment signaling pathways (padj < 0.05), particularly the TP53 signaling pathway, MAPK signaling pathway and apoptosis in CMTM3 overexpressed JHC7 cells (Fig. 5C). Wherein, TP53 signaling pathway plays important roles in regulation of tumorigenesis and progression and the expression of genes involved in the TP53 signaling pathway was significantly upregulated in CMTM3 overexpressed JHC7 cells, including TP53, SESN2, GADD45A, TP73, GADD45B and CYCS (Table 3, Fig. 5B).
Then, we confirmed these results with JHC7, U-CH1 and MUG-Chor1 cells by Real-time PCR and revealed that TP53 expression was apparently upregulated in CMTM3 overexpressed cells (Fig. 5D). Western blot was further performed in JHC7 cells and the expression of TP53 was upregulated in CMTM3 overexpressed cells (Fig. 5E). We knocked down CMTM3 in JHC7 cells (Fig. 5F, up) and found that TP53 was downregulated in CMTM3 silenced cells (Fig. 5F, down) at the protein level. Our previous study demonstrated that CMTM3 suppressed migration and invasion, but not the proliferation of gastric cancer cells [31, 32]. To clarify whether TP53 upregulation modulated CMTM3 inhibited cell proliferation, we analyzed TP53 expression in gastric cancer cell line SGC-7901 and the gastric epithelial cell line GES-1 by CMTM3 knockdown system using lentivirus transduction [31]. As shown in Additional file 4, TP53 expression was not influenced by CMTM3 knocked down in SGC-7901 and GES-1 cells. These data suggested that CMTM3 might contribute to inhibiting cell proliferation by upregulating TP53 expression. It has been reported that TP53 may respond to stress signals [43] and we found that the expression of heat shock protein family A member 6 (HSPA6), a member of the heat shock protein family was (ranking second besides CMTM3) was upregulated (Fig. 5B, Table 3). Additional file 5 further confirmed that the expression of HSPA6 was upregulated by CMTM3 at the mRNA level by Real-time PCR. These results suggested that TP53 signaling pathway involved in the CMTM3-suppressed proliferation in chordoma cells.
Taken together, we propose that CMTM3 suppresses chordomas through EGFR/STAT3 mediated EMT progression and TP53 signaling pathway.
Discussion
At present, the major treatment for chordomas is surgery and postoperative radiotherapy and they are difficult to be treated due to their destructive behavior locally and their sites [4]. Besides, it is crucial to accurately assess the dose profile outside the tumor with radiation treatment [44]. Therefore, it is necessary to explore the mechanism of chordomas tumorigenesis and development, and find its effective therapeutic targets. It has been reported that CMTM3 acts as a tumor suppressor gene to inhibit numerous cancers. Our previous data also demonstrated that CMTM3 suppressed the metastasis of gastric cancer through the EGFR/STAT3/EMT signaling pathway by interacting with Rab5 [31, 32]. In this paper, we used CMTM3 overexpressed and CMTM3 silenced systems to explore the roles and mechanism of CMTM3 in chordomas. This is the first time we found that CMTM3 inhibited chordoma with regulating EGFR/STAT3 mediated EMT progression and the TP53 signaling pathway. This study investigates the function and mechanism of CMTM3 in chordomas and provides us with a potential target for the treatment of chordomas. In the future, we may characterize the tumor tissues and cells by Raman-enhanced spectroscopy probe [45] to identify CMTM3 for potential tumor markers or diagnosis.
EGFR plays critical roles in the progression of cancers. Strong EGFR expression is significantly associated with tumor growth and metastasis [8]. Adle-Biassette demonstrated that EGFR was expressed in 244/284 (85.9%) of chordomas [46] and Yilmaz found that intensely positive EGFR was revealed in 77.5% (38/49) of chordoma patients [47]. Our previous study also found that EGFR played important roles in chordoma tumorigenesis and development [10]. In this paper, we found that CMTM3 facilitated the degradation of EGFR, and reduced EGFR expression level and p-EGFR (Fig. 4). CMTM3 colocalized with EGFR, but CMTM3 was not coimmunoprecipitated with the anti-EGFR antibody by Co-IP assay (Additional file 1B, C), which suggested that CMTM3 did not interact with EGFR. To date, the clinical relevance of EGFR in chordoma development is still controversial because of its relatively low incidence rate. Inhibitors [48, 49] or antibodies [50] against EGFR may act as potential treatments for chordomas. However, clinical trials with EGFR inhibitors and antibodies have not established a clear therapeutic benefit in all chordoma patients [51, 52]. We believe it is necessary and interesting to explore whether CMTM3 participates in facilitating the sensitivity to EGFR-targeted drugs and antibodies.
As we mentioned, CMTM3 inhibited tumorigenesis and progression mostly in epithelial-derived cancers and few studies revealed the roles of CMTM3 in mesenchymal-derived cancers. To our knowledge, this is the first study to assess the effects and mechanism of CMTM3 in mesenchymal cell-derived tumors. Our previous studies revealed that CMTM3 inhibited cell migration, but did not affect cell proliferation in epithelial-derived gastric cancer. In addition, CMTM3 reduced the phosphorylation of ERK1/2 in gastric cancer cells [32]. In this paper, CMTM3 inhibited cell proliferation and migration, but did not influence ERK1/2 phosphorylation in mesenchymal-derived chordomas. Thus, the roles of CMTM3 might vary in different cell-derived tumors.
TP53 plays crucial roles in tumor initiation and progression [53, 54]. This is the first time we revealed that CMTM3 regulates the TP53 signaling pathway in tumorigenesis and tumor progression. In this study, CMTM3 inhibited cell proliferation and suppressed p-STAT3, which significantly upregulated TP53 expression and enriched the TP53 signaling pathway in chordoma cells. However, CMTM3 has no effects on gastric cancer cell proliferation and does not alter TP53 expression after silencing CMTM3 by lentivirus (Additional file 4). These results suggest that CMTM3 suppresses cancer cell proliferation through upregulation of TP53 expression and activation of the TP53 signaling pathway.
Although our findings demonstrate the roles and mechanism of CMTM3 in the tumorigenesis and development of chordomas, this study still has limitations. First, we revealed that CMTM3 suppressed chordoma cell proliferation, migration and invasion in vitro, and tumor growth in vivo. However, tumor metastasis was not observed in the tumor-bearing mice. We speculated that this is because of the slow-growing characteristics of chordomas and that 5 months were not enough for tumor metastasis or for the observation of the chordoma-bearing mice survival. Second, we observed that CMTM3 reduced EGFR protein level and suppressed p-EGFR, and the fold change of p-EGFR is higher than that of EGFR (Fig. 4). Nevertheless, we still could not conclude that CMTM3 activated p-EGFR directly because the inhibition of EGFR phosphorylation may be a reflection of the reduced EGFR protein level.
Conclusions
In summary, our study first shows the function and mechanism of CMTM3 in chordomas. We find that CMTM3 expression is reduced in chordoma tissues and inhibits proliferation, migration and invasion of chordoma cells in vitro and suppresses tumor growth in vivo. Furthermore, CMTM3 facilitates EGFR degradation and inhibits tumorigenesis and development of chordomas via EGFR/STAT3 regulated EMT signaling pathway and TP53 signaling pathway. CMTM3 may act as a promising molecular target for chordomas, which would be clinically beneficial.
Availability of data and materials
The datasets used or analyzed during the current study are available from the corresponding author on reasonable request.
Abbreviations
- CCK8:
-
Counting Kit-8
- CFU:
-
Colony forming unit
- CMTM:
-
CKLF-like MARVEL transmembrane domain containing
- Co-IP:
-
Co-immunoprecipitation
- DEGs:
-
Differently expressed genes
- EGFR:
-
Epidermal growth factor receptor
- EMT:
-
Epithelial–mesenchymal transition
- HSPA6:
-
Heat shock protein family A member 6
- IHC:
-
Immunohistochemistry
- MOI:
-
Multiplicity of infection
- RT:
-
Reverse transcription
- Scr:
-
Scrambled siRNA
- siRNA:
-
Small interfering RNA
- STAT3:
-
Signal transducer and activator of transcription 3
References
Salisbury JR. The pathology of the human notochord. J Pathol. 1993;171(4):253–5. https://doi.org/10.1002/path.1711710404.
Williams BJ, Raper DM, Godbout E, Bourne TD, Prevedello DM, Kassam AB, Park DM. Diagnosis and treatment of chordoma. J Natl Compr Canc Netw. 2013;11(6):726–31. https://doi.org/10.6004/jnccn.2013.0089.
Ailon T, Torabi R, Fisher CG, Rhines LD, Clarke MJ, Bettegowda C, Boriani S, Yamada YJ, Kawahara N, et al. Management of locally recurrent chordoma of the mobile spine and sacrum: a systematic review. Spine (Phila Pa 1976). 2016;41:S193–8. https://doi.org/10.1097/BRS.0000000000001812.
Hai B, Ma Y, Liu X. A brief review of chordomas: pathogenesis, prognostic factors and therapeutic targets. Histol Histopathol. 2019;34(5):445–56. https://doi.org/10.14670/HH-18-080.
Yamada Y, Gounder M, Laufer I. Multidisciplinary management of recurrent chordomas. Curr Treat Options Oncol. 2013;14(3):442–53. https://doi.org/10.1007/s11864-013-0247-3.
Fatehi HM, Mansouri A, Alotaibi NM, Hazrati LN, Bernstein M. Metastatic saccrococcygeal chordoma. J Clin Neurosci. 2016;23:149–52. https://doi.org/10.1016/j.jocn.2015.05.036.
Stacchiotti S, Casali PG. Systemic therapy options for unresectable and metastatic chordomas. Curr Oncol Rep. 2011;13(4):323–30. https://doi.org/10.1007/s11912-011-0176-x.
Linder M, Glitzner E, Srivatsa S, Bakiri L, Matsuoka K, Shahrouzi P, Dumanic M, Novoszel P, Mohr T, et al. EGFR is required for FOS-dependent bone tumor development via RSK2/CREB signaling. EMBO Mol Med. 2018;10(11):e9408. https://doi.org/10.15252/emmm.201809408.
Huang F, Shi Q, Li Y, Xu L, Xu C, Chen F, Wang H, Liao H, Chang Z, et al. HER2/EGFR-AKT signaling switches TGFbeta from inhibiting cell proliferation to promoting cell migration in breast cancer. Cancer Res. 2018;78(21):6073–85. https://doi.org/10.1158/0008-5472.CAN-18-0136.
Liang C, Ma Y, Yong L, Yang C, Wang P, Liu X, Zhu B, Zhou H, Liu X, et al. Y-box binding protein-1 promotes tumorigenesis and progression via the epidermal growth factor receptor/AKT pathway in spinal chordoma. Cancer Sci. 2019;110(1):166–79. https://doi.org/10.1111/cas.13875.
Lv D, Li Y, Zhang W, Alvarez AA, Song L, Tang J, Gao WQ, Hu B, Cheng SY, et al. TRIM24 is an oncogenic transcriptional co-activator of STAT3 in glioblastoma. Nat Commun. 2017;8(1):1454. https://doi.org/10.1038/s41467-017-01731-w.
He L, Liu X, Yang J, Li W, Liu S, Liu X, Yang Z, Ren J, Wang Y, et al. Imbalance of the reciprocally inhibitory loop between the ubiquitin-specific protease USP43 and EGFR/PI3K/AKT drives breast carcinogenesis. Cell Res. 2018;28(9):934–51. https://doi.org/10.1038/s41422-018-0079-6.
Zhou Q, Huang T, Jiang Z, Ge C, Chen X, Zhang L, Zhao F, Zhu M, Chen T, et al. Upregulation of SNX5 predicts poor prognosis and promotes hepatocellular carcinoma progression by modulating the EGFR-ERK1/2 signaling pathway. Oncogene. 2020;39(10):2140–55. https://doi.org/10.1038/s41388-019-1131-9.
Jiang S, Wang X, Song D, Liu X, Gu Y, Xu Z, Wang X, Zhang X, Ye Q, et al. Cholesterol induces epithelial-to-mesenchymal transition of prostate cancer cells by suppressing degradation of EGFR through APMAP. Cancer Res. 2019;79(12):3063–75. https://doi.org/10.1158/0008-5472.CAN-18-3295.
Pastushenko I, Blanpain C. EMT transition states during tumor progression and metastasis. Trends Cell Biol. 2019;29(3):212–26. https://doi.org/10.1016/j.tcb.2018.12.001.
Caramel J, Ligier M, Puisieux A. Pleiotropic roles for ZEB1 in cancer. Cancer Res. 2018;78(1):30–5. https://doi.org/10.1158/0008-5472.CAN-17-2476.
Gulluoglu S, Tuysuz EC, Sahin M, et al. The role of TNF-alpha in chordoma progression and inflammatory pathways. Cell Oncol (Dordr). 2019;42:663–77. https://doi.org/10.1007/s13402-019-00454-y.
Niu G, Wright KL, Ma Y, Wright GM, Huang M, Irby R, Briggs J, Karras J, Cress WD, et al. Role of Stat3 in regulating p53 expression and function. Mol Cell Biol. 2005;25(17):7432–40. https://doi.org/10.1128/MCB.25.17.7432-7440.2005.
Chen J. The cell-cycle arrest and apoptotic functions of p53 in tumor initiation and progression. Cold Spring Harb Perspect Med. 2016;6(3):a26104. https://doi.org/10.1101/cshperspect.a026104.
Yu H, Yue X, Zhao Y, Li X, Wu L, Zhang C, Liu Z, Lin K, Xu-Monette ZY, et al. LIF negatively regulates tumour-suppressor p53 through Stat3/ID1/MDM2 in colorectal cancers. Nat Commun. 2014;5:5218. https://doi.org/10.1038/ncomms6218.
Ma J, Chen W, Wang K, Tian K, Li Q, Zhao, et al. Identification of the different roles and potential mechanisms of T isoforms in the tumor recurrence and cell cycle of chordomas. Onco Targets Ther. 2019;12:11777–91. https://doi.org/10.2147/OTT.S232526.
Han W, Lou Y, Tang J, Zhang Y, Chen Y, Li Y, Gu W, Huang J, Gui L, et al. Molecular cloning and characterization of chemokine-like factor 1 (CKLF1), a novel human cytokine with unique structure and potential chemotactic activity. Biochem J. 2001;357(Pt 1):127–35. https://doi.org/10.1042/0264-6021:3570127.
Han W, Ding P, Xu M, Wang L, Rui M, Shi S, Liu Y, Zheng Y, Chen Y, et al. Identification of eight genes encoding chemokine-like factor superfamily members 1–8 (CKLFSF1-8) by in silico cloning and experimental validation. Genomics. 2003;81(6):609–17. https://doi.org/10.1016/s0888-7543(03)00095-8.
Wang Y, Li J, Cui Y, Li T, Ng KM, Geng H, Li H, Shu XS, Li H, et al. CMTM3, located at the critical tumor suppressor locus 16q22.1, is silenced by CpG methylation in carcinomas and inhibits tumor cell growth through inducing apoptosis. Cancer Res. 2009;69(12):5194–201. https://doi.org/10.1158/0008-5472.CAN-08-3694.
Di Meo S, Airoldi I, Sorrentino C, Zorzoli A, Esposito S, Di Carlo E. Interleukin-30 expression in prostate cancer and its draining lymph nodes correlates with advanced grade and stage. Clin Cancer Res. 2014;20(3):585–94. https://doi.org/10.1158/1078-0432.CCR-13-2240.
Hu F, Yuan W, Wang X, Sheng Z, Yuan Y, Qin C, He C, Xu T. CMTM3 is reduced in prostate cancer and inhibits migration, invasion and growth of LNCaP cells. Clin Transl Oncol. 2015;17(8):632–9. https://doi.org/10.1007/s12094-015-1288-9.
**e J, Yuan Y, Liu Z, **ao Y, Zhang X, Qin C, Sheng Z, Xu T, Wang X. CMTM3 is frequently reduced in clear cell renal cell carcinoma and exhibits tumor suppressor activities. Clin Transl Oncol. 2014;16(4):402–9. https://doi.org/10.1007/s12094-013-1092-3.
Li Z, **e J, Wu J, Li W, Nie L, Sun X, Tang A, Li X, Liu R, et al. CMTM3 inhibits human testicular cancer cell growth through inducing cell-cycle arrest and apoptosis. PLoS ONE. 2014;9(2):e88965. https://doi.org/10.1371/journal.pone.0088965.
Zhang H, Zhang J, Nan X, Li X, Qu J, Hong Y, Sun L, Chen Y, Li T. CMTM3 inhibits cell growth and migration and predicts favorable survival in oral squamous cell carcinoma. Tumour Biol. 2015;36(10):7849–58. https://doi.org/10.1007/s13277-015-3504-1.
Lu M, Huang Y, Sun W, Li P, Li L, Li L. miR-135b-5p promotes gastric cancer progression by targeting CMTM3. Int J Oncol. 2018;52(2):589–98. https://doi.org/10.3892/ijo.2017.4222.
Yuan W, Liu B, Wang X, Li T, Xue H, Mo X, Yang S, Ding S, Han W. CMTM3 decreases EGFR expression and EGF-mediated tumorigenicity by promoting Rab5 activity in gastric cancer. Cancer Lett. 2017;386:77–86. https://doi.org/10.1016/j.canlet.2016.11.015.
Yuan W, Li T, Mo X, Wang X, Liu B, Wang W, Su Y, Xu L, Han W. Knockdown of CMTM3 promotes metastasis of gastric cancer via the STAT3/Twist1/EMT signaling pathway. Oncotarget. 2016;7(20):29507–19. https://doi.org/10.18632/oncotarget.8789.
Su Y, Lin Y, Zhang L, Liu B, Yuan W, Mo X, Wang X, Li H, **ng X, et al. CMTM3 inhibits cell migration and invasion and correlates with favorable prognosis in gastric cancer. Cancer Sci. 2014;105(1):26–34. https://doi.org/10.1111/cas.12304.
Dong W, Li J, Dong X, Shi W, Zhang Y, Liu Y. MiR-17 and miR-93 promote tumor progression by targeting p21 in patients with chordoma. Onco Targets Ther. 2021;14:3109–18. https://doi.org/10.2147/OTT.S307138.
Li L, Lv G, Wang B, Ma H. Long noncoding RNA LINC00525 promotes the aggressive phenotype of chordoma through acting as a microRNA-505-3p sponge and consequently raising HMGB1 expression. Onco Targets Ther. 2020;13:9015–27. https://doi.org/10.2147/OTT.S268678.
Chen L, Zuo Y, Pan R, et al. GSK-3beta regulates the expression of P21 to promote the progression of chordoma. Cancer Manag Res. 2021;2021(13):201–14. https://doi.org/10.2147/CMAR.S289883.
Chen H, Garbutt CC, Spentzos D, Choy E, Hornicek FJ, Duan Z. Expression and therapeutic potential of SOX9 in chordoma. Clin Cancer Res. 2017;23:5176–86. https://doi.org/10.1158/1078-0432.CCR-17-0177.
Wang J, Hu W, Du X, Sun Y, Han S, Tu G. Fingolimod inhibits proliferation and epithelial-mesenchymal transition in sacral chordoma by inactivating IL-6/STAT3 signalling. Biosci Rep. 2020. https://doi.org/10.1042/BSR20200221.
Zhao K, Li X, Chen X, et al. Inhibition of miR-140-3p or miR-155-5p by antagomir treatment sensitize chordoma cells to chemotherapy drug treatment by increasing PTEN expression. Eur J Pharmacol. 2019;854:298–306. https://doi.org/10.1016/j.ejphar.2019.03.034.
Guerriero JL, Sotayo A, Ponichtera HE, Castrillon JA, Pourzia AL, Schad S, Johnson SF, Carrasco RD, Lazo S, Bronson RT, Davis SP, Lobera M, Nolan MA, Letai A. Class IIa HDAC inhibition reduces breast tumours and metastases through anti-tumour macrophages. Nature. 2017;543:428–32. https://doi.org/10.1038/nature21409.
Faheem MM, Seligson ND, Ahmad SM, Rasool RU, Gandhi SG, Bhagat M, Goswami A. Convergence of therapy-induced senescence (TIS) and EMT in multistep carcinogenesis: current opinions and emerging perspectives. Cell Death Discov. 2020;6:51. https://doi.org/10.1038/s41420-020-0286-z.
Kosvyra A, Maramis C, Chouvarda I. Develo** an integrated genomic profile for cancer patients with the use of NGS data. Emerg Sci J. 2019;3(3):157–67. https://doi.org/10.28991/esj-2019-01178.
Choi S, Chen M, Cryns VL, Anderson RA. A nuclear phosphoinositide kinase complex regulates p53. Nat Cell Biol. 2019;21(4):462–75. https://doi.org/10.1038/s41556-019-0297-2.
Abdelaal AM, Attalla EM, Elshemey WM. Estimation of out-of-field dose variation using markus ionization chamber detector. SciMed J. 2020. https://doi.org/10.28991/SciMedJ-2020-0201-2.
Agsalda-Garcia M, Shieh T, Souza R, et al. Raman-enhanced spectroscopy (RESpect) probe for childhood non-hodgkin lymphoma. SciMed J. 2020;2:1–7. https://doi.org/10.28991/scimedj-2020-0201-1.
Tauziede-Espariat A, Bresson D, Polivka M, Bouazza S, Labrousse F, Aronica E, Pretet JL, Projetti F, Herman P, et al. Prognostic and therapeutic markers in chordomas: a study of 287 tumors. J Neuropathol Exp Neurol. 2016;75(2):111–20. https://doi.org/10.1093/jnen/nlv010.
Tosuner Z, Bozkurt SU, Kilic T, Yilmaz B. The role of EGFR, hepatocyte growth factor receptor (c-Met), c-ErbB2 (HER2-neu) and clinicopathological parameters in the pathogenesis and prognosis of chordoma. Turk Patoloji Derg. 2017;33(2):112–20. https://doi.org/10.5146/tjpath.2016.01378.
Scheipl S, Barnard M, Cottone L, Jorgensen M, Drewry DH, Zuercher WJ, Turlais F, Ye H, Leite AP, et al. EGFR inhibitors identified as a potential treatment for chordoma in a focused compound screen. J Pathol. 2016;239(3):320–34. https://doi.org/10.1002/path.4729.
Asquith C, Maffuid KA, Laitinen T, Torrice CD, Tizzard GJ, Crona DJ, Zuercher WJ. Targeting an EGFR water network with 4-anilinoquin(az)oline inhibitors for chordoma. ChemMedChem. 2019;14(19):1693–700. https://doi.org/10.1002/cmdc.201900428.
Magnaghi P, Salom B, Cozzi L, Amboldi N, Ballinari D, Tamborini E, Gasparri F, Montagnoli A, Raddrizzani L, et al. Afatinib is a new therapeutic approach in chordoma with a unique ability to target EGFR and Brachyury. Mol Cancer Ther. 2018;17(3):603–13. https://doi.org/10.1158/1535-7163.MCT-17-0324.
Eskilsson E, Rosland GV, Solecki G, Wang Q, Harter PN, Graziani G, Verhaak R, Winkler F, Bjerkvig R, et al. EGFR heterogeneity and implications for therapeutic intervention in glioblastoma. Neuro Oncol. 2018;20(6):743–52. https://doi.org/10.1093/neuonc/nox191.
Trapani D, Conforti F, De Pas T. EGFR inhibition in a pretreated sacral chordoma: a role for erlotinib? Case report and a brief review of literature. Transl Med UniSa. 2017;16:30–3.
Rupp M, Hagenbuchner J, Rass B, Fiegl H, Kiechl-Kohlendorfer U, Obexer P, Ausserlechner MJ. FOXO3-mediated chemo-protection in high-stage neuroblastoma depends on wild-type TP53 and SESN3. Oncogene. 2017;36(44):6190–203. https://doi.org/10.1038/onc.2017.288.
Donehower LA, Soussi T, Korkut A, Liu Y, Schultz A, Cardenas M, Li X, Babur O, Hsu TK, et al. Integrated analysis of TP53 gene and pathway alterations in The Cancer Genome Atlas. Cell Rep. 2019;28(5):1370–84. https://doi.org/10.1016/j.celrep.2019.07.001.
Acknowledgements
The authors would like to thank Prof. Wenling Han (Peking University Health Science Center Bei**g, China) for insightful suggestions for this study.
Funding
This work was supported by the National Key R&D Program of China (2018YFC1003702, 2018YFC1004403), the National Natural Science Foundation of China (81803095), the Key Clinical Projects of Peking University Third Hospital (BYSYZD2019041), the Bei**g Natural Science Foundation (7204327) and the Capital’s Funds for Health Improvement and Research (2020-4-40916).
Author information
Authors and Affiliations
Contributions
WY: Data curation, roles/writing-original draft and funding acquisition. FW: Data curation and methodology. HO, XR and JH: Funding acquisition. XM: Methodology and writing-review and editing. ZL: Conceptualization and writing-review and editing. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
This research was carried out after approval by the Ethics Committee of the Peking University Third Hospital Institutional Review Board (No. LM2019196). Informed consent was obtained from each patient.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflicts of interest to this work.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1.
A. The expression of EGFR in CMTM3 overexpressed JHC7 cells and U-CH1 cells and CMTM3 knocked-down MUG-Chor1 cells was detected by Real-time PCR. Representative data were shown as average number from one independent experiment. B. JHC7 cells were transfected with pEGFP-N1 empty vector or pEGFP-N1-CMTM3 plasmid for 48 h prior to fixation in 4% PFA and immunostained with antibodies against EGFR. EGFR staining is shown in red. Nuclei were visualized by DAPI (blue). The yellow color indicates the colocalization. The Pearson’s and Manders’ overlap coefficients were derived with LEICA QWin software (bar, 25 μm). C. Co-IP assay was performed with anti-EGFR antibody in CMTM3 overexpressed JHC7 cells and followed by western blot with the antibodies indicated (MOI: 100).
Additional file 2.
CMTM3 induces changes in gene expression profiles. A heat map summary reflecting gene expression values of JHC7-MOCK and JHC7-CMTM3 cells (MOI:100) (columns). Red indicates high and green indicates low gene expression values (padj < 0.05).
Additional file 3.
Upregulated and downregulated genes in JHC7-CMTM3 in comparison to JHC7-MOCK cells (padj < 0.05).
Additional file 4.
The expression of TP53 was detected by Real-time PCR in CMTM3 knocked-down SGC-7901 and GES-1 cells. Representative data were shown as average number from one independent experiment.
Additional file 5.
Overexpression of CMTM3 increased HSPA6 expression. The expression of HSPA6 in CMTM3 overexpressed JHC7 cells was determined by Real-time PCR. Representative data were shown as average number from one independent experiment.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Yuan, W., Wei, F., Ouyang, H. et al. CMTM3 suppresses chordoma progress through EGFR/STAT3 regulated EMT and TP53 signaling pathway. Cancer Cell Int 21, 510 (2021). https://doi.org/10.1186/s12935-021-02159-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12935-021-02159-5