Abstract
Background
Galangin, a flavonoid compound, is derived from Alpinia officinarum Hance. Previous studies have shown that galangin can inhibit the proliferation of hepatocellular carcinoma (HCC), but its mechanism is still unclear. This study aims to investigate the potential targets and molecular mechanisms of galangin on HCC through network pharmacology, bioinformatics, molecular docking, and experimental in vitro validation.
Methods
In this study, network pharmacology was used to investigate the targets and mechanisms of galangin in the treatment of HCC. AutoDockTools software was used to simulate and calculate the binding of galangin to its core targets. GO and KEGG enrichment analyses were conducted in the DAVID database to explore the main biological functions and signaling pathways impacted by galangin intervention. In addition, bioinformatics was applied to examine the correlation between the differential expressions of the anti-HCC core targets of galangin and the survival of patients with HCC. Finally, the findings obtained from network pharmacology and bioinformatics were verified in cell experiments.
Results
A total of 67 overlap** target genes of galangin and HCC were identified. Through the analysis of the protein-protein interaction (PPI) network, 10 hub genes with the highest degree of freedom were identified, including SRC, ESR1, MMP9, CDK4, CCNB1, MMP2, CDK2, CDK1, CHK1, and PLK1. These genes were found to be closely related to the degradation of the extracellular matrix, signal transduction, and the cell cycle. GO and KEGG enrichment analyses revealed that galangin exerts an anti-HCC role by affecting various signaling pathways, including the cell cycle, pathways in cancer, and the PI3K-Akt signaling pathway. The results of molecular docking indicated a significant interaction between galangin and CCNB1, CDK4, CDK1, and PLK1. Bioinformatics analysis revealed that CCNB1, CDK4, CDK1, and PLK1 were upregulated in the liver of patients with HCC at both the mRNA and protein levels. Flow cytometry analysis showed that galangin induced G0/G1 phase arrest and cell apoptosis in HepG2 and Huh7 cells. Additionally, galangin suppressed the expression of key proteins and mRNAs involved in the cell cycle pathway.
Conclusions
These results suggest that galangin inhibits the growth of HCC cells by arresting the cell cycle at the G0/G1 phase.
Similar content being viewed by others
Introduction
Hepatocellular carcinoma (HCC) is the most common type of liver cancer. Patients diagnosed with early-stage HCC, who have had the tumor completely removed, can anticipate positive treatment outcomes. Nevertheless, the early symptoms of HCC are not easily noticeable, and when it is eventually diagnosed, it is usually in the late stage. Patients suffering from advanced HCC, who have missed the optimal period for surgical treatment, are often only able to be treated with drugs. Despite its efficacy in symptom management and disease control, sorafenib, the first-line drug for advanced HCC, often triggers serious adverse reactions such as diarrhea, gastrointestinal bleeding, and hypertension during long-term use [1]. Therefore, it is the focus of current research to continuously explore therapeutic targets for HCC and develop highly efficient and specific targeted drugs for the treatment of HCC.
Evidence has shown that traditional Chinese medicine can be beneficial in alleviating symptoms and enhancing the quality of life of patients with HCC, inhibiting the recurrence of the tumor, and controlling the progression of the disease [2]. Galangin, a flavonoid derived from nature, is present in Alpinia officinarum Hance, Plantago depressa Willd., and propolis [1.
Statistical analysis
All the experimental data were expressed as mean ± SD. Statistical analyses were conducted using GraphPad Prism 9.0. Data conforming to normal distribution were analyzed by one-way ANOVA. The results of the homogeneity of variance were compared between groups using the LSD test. Dunnett’s T3 test was used for unequal variances.
Results
Network pharmacology analysis of galangin
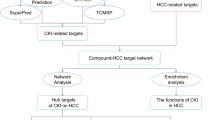
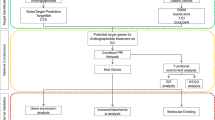
As shown in Fig. 1A, the chemical structure of galangin was obtained from the TCMSP database. Through research of the BATMAN-TCM database and the SwissTargetPrediction website, 120 potential targets of galangin were identified. HCC disease targets were collected from various databases, including 1981 in the GeneCards database, 507 in the OMIM database, 1396 in the DisGeNET database, and 284 in the PharmGKB database. After merging and deleting duplicate targets, a total of 3319 HCC targets were collected (Fig. 1B). By intersecting 120 targets of galangin with 3319 targets of HCC disease, 67 shared targets of galangin against HCC were determined (Fig. 1C).
Network pharmacology predicted the potential mechanisms of galangin for HCC. (A) The chemical structure formula of galangin. (B) The Venn diagram of the HCC-related targets in the GeneCards, OMIM, DisGeNET, and PharmGKB databases. (C) The Venn diagram of the targets shared by galangin and HCC. (D) The top ten core targets of degree of freedom values in topological analysis. (E) PPI network. (F) Network diagram of galangin-pathway-HCC-target. (G) The top 10 terms of BPs, CCs, and MFs enrichment analysis from GO enrichment analysis. (H) KEGG enrichment analysis of potential therapeutic targets for galangin against HCC
The target of galangin against HCC was inputted into the STRING database for processing, and a PPI network diagram was obtained (Fig. 1D). The relevant data was imported into Cytoscape 3.9.1 software, and network topology analysis was performed in the software to screen the top ten core targets with the highest degree of centrality (Fig. 1E). These targets include SRC, ESR1, MMP9, CDK4, CCNB1, MMP2, CDK2, CDK1, CHK1 (CHEK1), and PLK1.
67 common targets of galangin and HCC were analyzed for GO and KEGG enrichment. GO functional enrichment analysis showed that the target proteins were involved in 55 biological processes (BPs), 44 cellular components (CCs), and 41 molecular functions (MFs). The top 10 terms, according to gene counts in each category, are presented as column charts (Fig. 1G). The KEGG enrichment analysis showed that galangin can mainly affect 37 signaling pathways, including the cell cycle, pathways in cancer, the PI3K-Akt signaling pathway, microRNAs in cancer, and chemical carcinogenesis - receptor activation. The top 20 pathways enriched by KEGG are shown in Fig. 1H.
The “galangin-target-pathway-HCC” interaction network (Fig. 1F) was constructed by importing information files of galangin, common targets, signaling pathways, and HCC into Cytoscape 3.9.1. The network is composed of 89 nodes and 322 edges. The yellow circle represents the target, the blue diamond indicates the signaling pathway, the red inverted triangle denotes galangin, the green triangle represents HCC, and the lines represent the interconnections between them. As shown in Fig. 1F, the key targets of galangin therapy for HCC, including CCNB1, CDK4, CDK1, and PLK1, belong to the cell cycle signaling pathway (hsa04110).
Molecular docking analysis of galangin
When the absolute value of the binding energy between the ligand and the receptor is greater, the binding is more stable. A binding energy of less than -5 kcal/mol implies a strong affinity between the ligand and the receptor [10]. Docking experiments were conducted using galangin as a ligand with four key targets, including CCNB1, CDK4, CDK1, and PLK1, as receptors. As shown in Fig. 2, the docking results of galangin and its key targets were visualized using PyMOL software. The binding energies of galangin with CCNB1, CDK4, CDK1, and PLK1 were -5.4 kcal/mol, -5.5 kcal/mol, -7.8 kcal/mol, and -5.6 kcal/mol, respectively. The findings indicate that galangin has a good binding affinity with each of these key receptors.
Galangin suppressed the viability and proliferation of HCC cells
As shown in Fig. 3A, galangin had no inhibitory effect on the proliferation of LX-2 cells at concentrations of 40–160 µM for 12 h (p > 0.05). On the contrary, galangin showed a significant inhibitory effect on the proliferation of HepG2 cells (p < 0.001) and Huh7 cells (p < 0.05) (Fig. 3B and C). The results indicated that galangin displayed cytotoxic effects on HCC cells, including HepG2 and Huh7 cells, within the concentration range of 40–160 µM. Additionally, it impeded the proliferation of these cancer cells while showing no cytotoxicity towards the normal LX-2 cells. The half maximal inhibitory concentration (IC50) of galangin on HepG2 and Huh7 cells was 73.88 µM and 82.41 µM, respectively. Consequently, concentrations of 40, 80, and 120 µM of galangin were chosen for subsequent experiments. In addition, EDU staining experiments confirmed that cell proliferation was attenuated in HepG2 and Huh7 cells treated with galangin (Fig. 3D and E). In summary, these findings support the anti-proliferative effect of galangin on HCC cells in vitro.
Effect of galangin on viability and proliferation of three cell lines. The cytotoxicity of galangin (40, 80, 120, and 160 µM) on (A) LX-2, (B) HepG2, and (C) Huh7 cells was determined using the CCK8 method. Effect of galangin (40, 80, and 120 µM) on the proliferation of (D) HepG2 cells and (E) Huh7 cells. *p < 0.05, **p < 0.01, and ***p < 0.001 vs. control group
Galangin inhibited cell cycle progression and induced apoptosis in HCC cells
As shown in Fig. 4A and B, treatment of HepG2 cells with different concentrations of galangin (40, 80, and 120 µM) resulted in a significant increase in the rate of apoptosis. Specifically, 80 µM (p < 0.01) and 120 µM (p < 0.001) of galangin significantly increased the apoptosis rate of HepG2 cells. Meanwhile, galangin can induce apoptosis in Huh7 cells in a dose-dependent manner (p < 0.001). These results indicate that galangin can induce apoptosis in HepG2 and Huh7 cells and inhibit the proliferation of HCC cells.
Effects of galangin treatments on HCC cell apoptosis and cell cycle progression. The apoptosis of HepG2 (A) and Huh7 cells (B) was evaluated using flow cytometry. The cell cycle progression of (C) HepG2 and (D) Huh7 cells was measured using flow cytometry. *p < 0.05 and ***p < 0.001 vs. control group
The effect of galangin on the cell cycle of HepG2 cells was further detected by flow cytometry. HepG2 and Huh7 cells were exposed to different concentrations of galangin (40, 80, and 120 µM) for 12 h. As shown in Fig. 4C, compared to the control group, the percentage of HepG2 cells in the G0/G1 phase significantly increased in the galangin administration group (p < 0.001), while the percentage of HepG2 cells in the S phase significantly decreased (p < 0.001). Similarly, the percentage of Huh7 cells in the G0/G1 phase significantly increased in the galangin administration groups (p < 0.05), while the percentage of Huh7 cells in the G2/M phase significantly decreased (p < 0.05) (Fig. 4D). The results of the cell cycle experiment demonstrate that galangin could block cell cycle progression in HepG2 and Huh7 cells and inhibit the proliferation of HCC cells.
The effect of galangin on the mRNA and protein expression of core targets in HCC cells. Effect of galangin on the expression of CDC25C, CCNB1, CDK4, CDK1, and PLK1 proteins in (A) HepG2 and (B) Huh7 cells. Effect of galangin on the expression of ATR, CHK1, CDC25C, CCNB1, CDK4, CDK1, and PLK1 mRNA in (C) HepG2 and (D) Huh7 cells. *p < 0.05, **p < 0.01, and ***p < 0.001 vs. control group
Galangin downregulated the expression of the core targets
As shown in Fig. 5A and B, Western Blot was used to detect the expression of CCNB1, CDK4, CDK1, and PLK1 proteins in HepG2 and Huh7 cells treated with different concentrations of galangin (40, 80, and 120 µM) for 12 h. Compared to the control group, the expressions of CCNB1, CDK4, CDK1, and PLK1 in HepG2 cells were significantly decreased (p < 0.05, p < 0.01, or p < 0.001) in a dose-dependent manner. Similarly, galangin also inhibited the expression of CCNB1, CDK4, CDK1, and PLK1 in Huh7 cells in a dose-dependent manner (p < 0.05, p < 0.01, or p < 0.001). CDC25C, a dual-specificity phosphatase, is essential for controlling the serine/threonine kinases involved in the cell cycle. Our experiments found that galangin significantly downregulated the expression level of the CDC25C protein in HepG2 and Huh7 cells.
In order to further study the effect of galangin on cell cycle-related targets, qRT-PCR was used to detect the mRNA expression of different concentrations of galangin on the relevant targets. As shown in Fig. 5C and D, the mRNA expressions of ATR, CHK1, CDC25C, CCNB1, CDK4, CDK1, and PLK1 were significantly downregulated in HepG2 and Huh7 cells treated with different concentrations of galangin. These results further corroborated that galangin is capable of modulating the core targets of the cell cycle by suppressing the ATR/CHK1 DNA damage response pathway, thereby exhibiting anti-HCC activity.
The differential expression of core targets in HCC was examined through bioinformatics analysis
In this study, the mRNA expressions of CCNB1, CDK4, CDK1, and PLK1 in liver tissue were analyzed using the TCGA database. As shown in Fig. 6A, C, E, and G, the mRNA expression levels of CCNB1 (P = 2.66e-28), CDK4 (P = 2.494e-18), CDK1 (P = 6.273e-28), and PLK1 (P = 1.584e-27) in the HCC group were significantly higher than those in the normal group. To further clarify the expression of CCNB1, CDK4, CDK1, and PLK1 at the protein level in the liver, high-throughput proteomic data based on mass spectrometry technology was analyzed in the CPTAC database. As shown in Fig. 6B, D, F, and H, compared to the normal group, the expressions of CCNB1 (P = 3.291E-08), CDK4 (P = 5.258E-06), CDK1 (P = 3.744E-39), and PLK1 (P = 4.172E-04) proteins were upregulated in HCC patients. The results above indicate that CCNB1, CDK4, CDK1, and PLK1 are upregulated in the livers of patients with HCC at both the mRNA and the protein levels.
Differential expression of core targets in liver tissue. (A) Differential expression of CCNB1 mRNA in the liver. (B) Differential expression of CCNB1 protein in the liver. (C) Differential expression of CDK4 mRNA in the liver. (D) Differential expression of CDK4 protein in the liver. (E) Differential expression of CDK1 mRNA in the liver. (F) Differential expression of CDK1 protein in the liver. (G) Differential expression of PLK1 mRNA in the liver. (H) Differential expression of PLK1 protein in the liver
The relationship between core target expression and overall survival in HCC patients. The effect of CCNB1 (A), CDK4 (B), CDK1 (C), and PLK1 (D) mRNA expression levels on the survival of HCC patients was evaluated using the TCGA database. The effect of CCNB1 (E), CDK4 (F), CDK1 (G), and PLK1 (H) protein expression levels on the survival of HCC patients was evaluated using the CPTAC database. The effect of CCNB1 (I), CDK4 (J), CDK1 (K), and PLK1 (L) mRNA expression levels on the survival of HCC patients was further confirmed by the Kaplan-Meier plotter online database
The effects of mRNA expression of CCNB1, CDK4, CDK1, and PLK1 on the survival of HCC patients were evaluated using the TCGA database. The median mRNA expression levels of CCNB1, CDK4, CDK1, and PLK1 were used as cut-off values to divide HCC patients into high-expression and low-expression groups. Kaplan-Meier survival analysis showed that the overall survival rate of HCC patients in the high mRNA expression group of CCNB1 (P = 0.00026), CDK4 (P = 0.00016), CDK1 (P = 0.0029), and PLK1 (P = 0.00036) was significantly lower than that of those in the low mRNA expression group (Fig. 7A-D). Furthermore, the relationship between the expression levels of CCNB1, CDK4, CDK1, and PLK1 proteins in the liver and the survival of HCC patients was validated using the CPTAC database. Compared to the low-expression group of HCC patients, those with high expression levels of CCNB1 (P = 0.015), CDK4 (P = 0.023), CDK1 (P = 0.00081), and PLK1 (P = 0.00074) proteins had a lower overall survival rate (Fig. 7E-H). The Kaplan-Meier Plotter online database was utilized to ascertain the correlation between the mRNA expression levels of CCNB1, CDK4, CDK1, and PLK1 and the survival rate of HCC patients. The results showed that the mRNA expression levels of CCNB1 (P = 0.00015), CDK4 (P = 0.0021), CDK1 (P = 0.00017), and PLK1 (P = 0.00082) were negatively correlated with the overall survival rate of HCC patients (Fig. 7I-L). These findings are consistent with the analysis results obtained from the TCGA database. Therefore, the results of bioinformatics analysis demonstrate that CCNB1, CDK4, CDK1, and PLK1 are key targets for HCC therapy.
Discussion
HCC is a common primary liver malignancy. The incidence of HCC is increasing year by year, with a high mortality rate and a poor prognosis. With the advancement of research in traditional Chinese medicine, it is of great scientific and practical value to extract effective and less toxic active ingredients and reveal their anticancer mechanisms. Galangin is the main functional component extracted from Alpinia officinarum Hance [11, 12]. Many studies have shown that galangin can inhibit the growth and invasion of various malignant tumors, including lung cancer [13], breast cancer [14], liver cancer [15], bladder cancer [16], and leukemia [17]. Its anticancer spectrum is relatively wide and has a good application prospect. Although galangin has been extensively studied as an antineoplastic agent in various cancers, its mechanism of action in HCC remains unclear.
The employment of network pharmacology introduces a novel methodology to investigate the mechanisms of drug therapy for diverse diseases. In this study, the results of network pharmacology showed that the key targets of galangin against liver cancer were SRC, ESR1, MMP9, CDK4, CCNB1, MMP2, CDK2, CDK1, CHEK1, and PLK1. Among these, CCNB1, CDK1, CDK4, and PLK1 are important regulators of the cell cycle. Pathway enrichment analysis also revealed that the cell cycle pathway is a crucial pathway for galangin therapy in HCC. The cell-cycle signaling pathway is a common activation pathway in cancer development, which can regulate tumor cell apoptosis and the cell cycle [18]. As a member of the cyclin family, CCNB1 is highly expressed in a variety of human tumor tissues and is closely related to tumor cell proliferation, metastasis, and poor prognosis in patients [19]. CDK1 is an important regulatory protein in the cell cycle and is the only member of the CDK family that can independently regulate the cell cycle [20]. Studies have shown that CCNB1 and CDK1 are highly expressed in patients with HCC. The siRNA-mediated knockdown of the CCNB1 and CDK1 genes significantly induces autophagy and senescence in HCC cells [19]. The co-administration of CCNB1 and CDK1 inhibitors with immunotherapy has been shown to increase the survival time of patients with HCC [21]. CDK4 is the first cyclin-dependent protein kinase activated during the cell proliferation stage and is a rate-limiting factor of the G1/S phase transition in the cell cycle. The proliferation of human HCC cell lines is inhibited by CDK4 inhibitors through the promotion of reversible cell cycle arrest [22]. PLK1 is an important kinase that regulates the cell cycle and is abundantly expressed in HepG2 cells. Studies have shown that knocking down the PLK1 gene can significantly inhibit the migration of human liver cancer vascular endothelial cells [22, 23]. In this study, bioinformatics analysis found that HCC patients with high expression of CCNB1, CDK4, CDK1, and PLK1 proteins and mRNA had a lower overall survival rate. In vitro tests showed that the protein and mRNA expression levels of CCNB1, CDK4, CDK1, and PLK1 in HepG2 and Huh7 cells were significantly reduced after exposure to various concentrations of galangin. In addition, the molecular docking results demonstrated the robust binding affinity of galangin with these targets. Taken together, these results suggest that galangin may induce apoptosis in liver cancer cells by disrupting the cell cycle through its influence on cell cycle proteins such as CCNB1, CDK4, CDK1, and PLK1.
During evolution, cells have developed a rigorous and effective set of checkpoint mechanisms to ensure that their genetic material is accurately replicated and transmitted to subsequent generations [24]. DNA damage activates the cell cycle checkpoints, which stop the cell cycle, giving the DNA enough time to repair, or causing the cell to die if the damage is irreparable. The regulation of the cell cycle from the end of the G1 phase to the S phase and G2 phase is predominantly controlled by the ATR/CHK1/CDC25 pathway. After DNA damage, ATR accumulates at the damaged site and becomes activated. Its downstream target, CHK1, is subsequently activated, which then activates p53. This ultimately leads to cell cycle arrest. Cell division cycle 25 C (CDC25C) is crucial for regulating the activity of serine/threonine kinases involved in the cell cycle. Moreover, CDC25C can activate the CCNB1/CDK1 complex by triggering dephosphorylation of CDK1, thereby promoting the G2/M transition in mitotic cells. In the present work, the expression of ATR, CHK1, and CDC25C mRNA was detected in HepG2 and Huh7 cells treated with galangin. The results showed that galangin effectively reduced the expression of ATR, CHK1, and CDC25C mRNA in HCC cells. The findings of the cell cycle studies showed that galangin inhibited the proliferation of HCC cells in the G0/G1 phase. The cell cycle signaling pathway interfered with by galangin is shown in Fig. 8.
In this study, network pharmacology was used to predict the targets of galangin in the treatment of HCC, which were subsequently validated through in vitro experiments. Therefore, it is imperative for future research to prioritize conducting experimental validation in animal models to offer more precise evidence supporting the efficacy of galangin in treating HCC. Although molecular docking simulations can unveil the interactions between galangin and its targets, they are inadequate in fully representing its pharmacological and biological effects in the human body. Consequently, it is crucial to carry out in-depth studies on the treatment process, adverse effects, and prognosis in clinical trials. Such investigations will expedite the utilization of galangin as a therapeutic agent for liver cancer in clinical practice.
Conclusions
In summary, galangin induces apoptosis in HCC cells by blocking the cell cycle at the G0/G1 phase through the inhibition of key mRNA and protein expression in the cell cycle. The present study provides a new research direction for the treatment of HCC using galangin and offers ideas and a theoretical basis for investigating lead compounds for the treatment of HCC.
Data availability
The original contributions presented in the study are included in the article, and further inquiries can be directed to the corresponding author.
Abbreviations
- HCC:
-
Hepatocellular carcinoma
- qPCR:
-
Quantitative real-time PCR
- BP:
-
Biological processes
- CC:
-
Cellular components
- MF:
-
Molecular functions
- CDC25C:
-
Cell division cycle-25 C
- GO:
-
Gene Ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- CDK1:
-
Cyclin-dependent kinase-1
References
Tang W, Chen Z, Zhang W, Cheng Y, Zhang B, Wu F, Wang Q, Wang S, Rong D, Reiter FP, et al. The mechanisms of sorafenib resistance in hepatocellular carcinoma: theoretical basis and therapeutic aspects. Signal Transduct Target Ther. 2020;5(1):87.
Tang KY, Du SL, Wang QL, Zhang YF, Song HY. Traditional Chinese medicine targeting cancer stem cells as an alternative treatment for hepatocellular carcinoma. J Integr Med. 2020;18(3):196–202.
WANG **aoqing SY, Fangshu ZHAO, Yiming YIN, Wenting,LI NI, Baohong. HOU Lin: Research Progress on mechanism and pharmacological activities of Galangin. Pharmacol Clin Chin Materia Med. 2023;39(08):115–20.
Rampogu S, Gajula RG, Lee KW. A comprehensive review on chemotherapeutic potential of galangin. Biomed Pharmacother. 2021;141:111808.
Fang D, **ong Z, Xu J, Yin J, Luo R. Chemopreventive mechanisms of galangin against hepatocellular carcinoma: a review. Biomed Pharmacother. 2019;109:2054–61.
Liang X, Wang P, Yang C, Huang F, Wu H, Shi H, Wu X. Galangin inhibits gastric Cancer Growth through enhancing STAT3 mediated ROS Production. Front Pharmacol. 2021;12:646628.
Zhu L, Luo Q, Bi J, Ding J, Ge S, Chen F. Galangin inhibits growth of human head and neck squamous carcinoma cells in vitro and in vivo. Chem Biol Interact. 2014;224:149–56.
Murray TJ, Yang X, Sherr DH. Growth of a human mammary tumor cell line is blocked by galangin, a naturally occurring bioflavonoid, and is accompanied by down-regulation of cyclins D3, E, and a. Breast Cancer Res. 2006;8(2):R17.
Liu Zheng WH. Apoptotic effects of Galangin on the Hepatoma Carcinoma cells HepG-2. Sci Technol Food Ind. 2020;36(08):1862–5.
Santos LHS, Ferreira RS, Caffarena ER. Integrating Molecular Docking and Molecular Dynamics simulations. Methods Mol Biol. 2019;2053:13–34.
Li N, Men W, Zheng Y, Wang H, Meng X. Oroxin B induces apoptosis by Down-regulating MicroRNA-221 resulting in the inactivation of the PTEN/PI3K/AKT pathway in Liver Cancer. Molecules 2019, 24(23).
Ning LEI, Benshan LYXU, Chengmin LI, Shushan DU. Review of flavonoids in preventing and treating Hepatocellular Carcinoma. Traditional Chin Drug Res & Clin Pharmacol. 2016;27(05):726–30.
Yu S, Gong LS, Li NF, Pan YF, Zhang L. Galangin (GG) combined with cisplatin (DDP) to suppress human lung cancer by inhibition of STAT3-regulated NF-κB and Bcl-2/Bax signaling pathways. Biomed Pharmacother. 2018;97:213–24.
Qaddoori MH, Al-Shmgani HS. Galangin-Loaded Gold Nanoparticles: Molecular Mechanisms of Antiangiogenesis Properties in Breast Cancer. Int J Breast Cancer 2023, 2023:3251211.
Chien ST, Shi MD, Lee YC, Te CC, Shih YW. Galangin, a novel dietary flavonoid, attenuates metastatic feature via PKC/ERK signaling pathway in TPA-treated liver cancer HepG2 cells. Cancer Cell Int. 2015;15:15.
Long X, Chen L, Yang J, Dong T, Cheng Q, Wang W, Zou Y, Su Y, Dai W, Chen B, et al. Network-based Pharmacology and in vitro validation reveal that Galangin induces apoptosis in bladder Cancer cells by promoting the P53 signaling pathway. Anticancer Agents Med Chem. 2023;23(7):847–57.
Tolomeo M, Grimaudo S, Di Cristina A, Pipitone RM, Dusonchet L, Meli M, Crosta L, Gebbia N, Invidiata FP, Titone L, et al. Galangin increases the cytotoxic activity of imatinib mesylate in imatinib-sensitive and imatinib-resistant bcr-abl expressing leukemia cells. Cancer Lett. 2008;265(2):289–97.
Huang W, Sun H, Hu T, Zhu D, Long X, Guo H, Liu Q. Blocking the short isoform of augmenter of liver regeneration inhibits proliferation of human multiple myeloma U266 cells via the MAPK/STAT3/cell cycle signaling pathway. Oncol Lett. 2021;21(3):197.
Pinto Marques H, Gomes da Silva S, De Martin E, Agopian VG, Martins PN. Emerging biomarkers in HCC patients: current status. Int J Surg. 2020;82s:70–6.
Malumbres M, Barbacid M. Cell cycle, CDKs and cancer: a changing paradigm. Nat Rev Cancer. 2009;9(3):153–66.
Zou Y, Ruan S, ** L, Chen Z, Han H, Zhang Y, Jian Z, Lin Y, Shi N, ** H. CDK1, CCNB1, and CCNB2 are prognostic biomarkers and correlated with Immune Infiltration in Hepatocellular Carcinoma. Med Sci Monit. 2020;26:e925289.
ChengYi YDWZWHS. Expression and significance of Plk1, Chk1 /2 protein in primary hepatic carcinoma tissue and HepG2 cell. Chin J Immunol. 2015;31(06):758–60.
Huijie YU, Li WY, Li F. Influence of knocking down Plk1 on migration of human hepatocellular carcinoma tumor-derived endothelial cells. Chin J Cancer Prev Treat. 2015;22(08):584–7.
Houtgraaf JH, Versmissen J, van der Giessen WJ. A concise review of DNA damage checkpoints and repair in mammalian cells. Cardiovasc Revasc Med. 2006;7(3):165–72.
Funding
This study was supported by Joint Program on Health Science & Technology Innovation of Hainan Province (WSJK2024QN062, WSJK2024MS156), Innovative Entrepreneurship Training Program for Undergraduates of Hainan Medical University (X202311810070), The National Natural Science Foundation of China (82360836) and the Hainan Province Clinical Medical Center.
Author information
Authors and Affiliations
Contributions
Jian Xu, Junqing Zhang, and Yipeng Pan provided fund support and designed this experiment. **aoliang Li, Jiangbo Sun, Bingshu Wang, Yihang Zhao, and Gaoan Wang carried out this experiment and wrote the article. Mingyan Zhou and Weijia Chen provided assistance in conducting experiments and performing statistical analysis. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Li, X., Zhou, M., Chen, W. et al. Integrating network pharmacology, bioinformatics, and experimental validation to unveil the molecular targets and mechanisms of galangin for treating hepatocellular carcinoma. BMC Complement Med Ther 24, 208 (2024). https://doi.org/10.1186/s12906-024-04518-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12906-024-04518-x