Abstract
Background
The production of succinic acid (SA) from biomass has attracted worldwide interest. Saccharomyces cerevisiae is preferred for SA production due to its strong tolerance to low pH conditions, ease of genetic manipulation, and extensive application in industrial processes. However, when compared with bacterial producers, the SA titers and productivities achieved by engineered S. cerevisiae strains were relatively low. To develop efficient SA-producing strains, it’s necessary to clearly understand how S. cerevisiae cells respond to SA.
Results
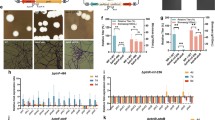
In this study, we cultivated five S. cerevisiae strains with different genetic backgrounds under different concentrations of SA. Among them, KF7 and NBRC1958 demonstrated high tolerance to SA, whereas NBRC2018 displayed the least tolerance. Therefore, these three strains were chosen to study how S. cerevisiae responds to SA. Under a concentration of 20 g/L SA, only a few differentially expressed genes were observed in three strains. At the higher concentration of 60 g/L SA, the response mechanisms of the three strains diverged notably. For KF7, genes involved in the glyoxylate cycle were significantly downregulated, whereas genes involved in gluconeogenesis, the pentose phosphate pathway, protein folding, and meiosis were significantly upregulated. For NBRC1958, genes related to the biosynthesis of vitamin B6, thiamin, and purine were significantly downregulated, whereas genes related to protein folding, toxin efflux, and cell wall remodeling were significantly upregulated. For NBRC2018, there was a significant upregulation of genes connected to the pentose phosphate pathway, gluconeogenesis, fatty acid utilization, and protein folding, except for the small heat shock protein gene HSP26. Overexpression of HSP26 and HSP42 notably enhanced the cell growth of NBRC1958 both in the presence and absence of SA.
Conclusions
The inherent activities of small heat shock proteins, the levels of acetyl-CoA and the strains’ potential capacity to consume SA all seem to affect the responses and tolerances of S. cerevisiae strains to SA. These factors should be taken into consideration when choosing host strains for SA production. This study provides a theoretical basis and identifies potential host strains for the development of robust and efficient SA-producing strains.
Similar content being viewed by others
Background
Succinic acid (SA), a C-4 building block chemical, has been widely used in medicine, agriculture, industry, and so on [1]. For example, succinate is widely used as an intermediary feedstock to produce chemicals such as 1,4-butanediol, tetrahydrofuran, γ-butyrolactone, succinate salts, and adipic acid [2]. In 2004, the U.S. Department of Energy (DOE) proposed that SA is one of the five most promising bio-based platform chemicals.
The production of SA via petrochemical processing is facing challenges posed by unsustainable fossil energy supplies and increased environmental burdens. Therefore, microbial factories have become a promising alternative. Microorganisms such as Actinobacillus succinogenes [32, 34, 63, 64]. An increased PQC activity might enhance the SA tolerance of S. cerevisiae. However, NBRC2018 showed a significant downregulation of HSP26, which could potentially undermine its ability to tolerate SA stress. Given that HSP26 and HSP42 encode the only two sHsps in S. cerevisiae, they might share similar functions in responding to SA stress. Therefore, to verify their roles in SA tolerance, we overexpressed HSP26 and HSP42 in all three strains. The constitutively strong promoter TEF1p was used to express HSP26 and HSP42. Six strains were obtained and named as KF7 + TSP26, KF7 + TSP42, N19 + TSP26, N19 + TSP42, N20 + TSP26, and N20 + TSP42, respectively. The growths of these strains under different concentrations of SA (0 g/L, 60 g/L, and 80 g/L) were compared with their original strains (Fig. 6; Table 4).
Influence of HSP26 or HSP42 overexpression on the SA tolerances of S. cerevisiae strains KF7 (a), NBRC1958 (b), and NBRC2018 (c). Cells were exposed to 0 g/L, 60 g/L, and 80 g/L SA, respectively. The initial inoculum was OD600 0.4. OD600 was measured at 10 h of cultivation. Taking the original strains KF7, NBRC1958, and NBRC2018 as controls for each group. Values and standard deviations were calculated from three repeated samples. *p < 0.05; **p < 0.001; ***p < 0.0001; ****p < 0.00001; ns, no statistically significant difference
The overexpression of HSP26 and HSP42 led to a significant improvement in the growth of NBRC1958. In detail, when HSP26 was overexpressed, the cell growth of NBRC1958 increased by 50%, 19%, and 52% at 0 g/L, 60 g/L, and 80 g/L SA, respectively. Overexpression of HSP42 had a nearly identical effect on the cell growth of NBRC1958. The results indicated that enhancing the activity of sHsps can elevate the intrinsic growth capacity of NBRC1958, thus boosting its ability to grow under SA stress. This finding aligns with prior research, which demonstrated that sHsps serve as universal effectors of longevity, and the overexpression of HSP26 extended the replicative lifespan of yeast cells [65]. Consistently, sHsps have been implicated in S. cerevisiae’s response to other weak acids, such as sorbic acid and citric acid [66, 67]. Overexpression of HSP26 has been shown to enhance the strains’ tolerance to sorbic acid [66]. However, for the other two strains, neither the overexpression of HSP26 nor HSP42 had a notable effect on cell growth. It was likely that some other key limiting factors may play a more decisive role in determining the SA tolerance of the two strains. In conclusion, overexpressing HSP26 or HSP42 in strains with inherent low sHsps activities is one of the methods to improve SA tolerance.
Genetic background has been proven to affect sugar metabolism and inhibitor tolerance of S. cerevisiae [68, 69]. In this study, we observed that the response mechanisms of different strains to SA were indeed influenced by their genetic backgrounds (Figs. 3, 4 and 5). For example, KF7 and NBRC2018 showed notable differences in the regulation of acetyl-CoA metabolism under SA stress. When KF7 was exposed to SA, genes associated with fatty acids beta-oxidation and the glyoxylate cycle were significantly downregulated. As a result, the production of acetyl-CoA from peroxisomes reduced, leading to a correspondingly reduction in endogenous succinate synthesis. Instead, exogenous SA was likely to be used for acetyl-CoA biosynthesis to maintain intracellular acetyl-CoA levels in KF7 (Fig. 3). On the contrary, when NBRC2018 encountered severe SA stress (60 g/L), genes related to fatty acid β-oxidation and the glyoxylate cycle were significantly upregulated. Despite the increased availability of acetyl-CoA, this upregulation also promoted the generation of endogenous succinate, which further exacerbating the intracellular SA stress. This differential response may be one of the main reasons why NBRC2018 was less tolerant to SA than KF7. Thus, when develo** SA-producing strains, it is crucial to carefully consider the intracellular acetyl-CoA levels and the strains’ capacity to utilize SA.
In general, weak acids typically cause toxicity in S. cerevisiae cells through several mechanisms, include intracellular acidification, membrane damage, oxidative stress, protein aggregation, carbon metabolism disruption, etc. [70]. Accordingly, yeast cells employed a variety of complex and diverse regulatory mechanisms to cope with weak acids [70, 71]. For instance, the activities of plasma membrane H+-ATPase and ABC transporters are increased in response to acetic acid [71]; the maintenance of CWI is crucial for yeast’s adaptation and tolerance to acetic and lactic acid [72]; the biosynthesis of purine and methionine is reduced in response to formic and acetic acids [73, In this study, five S. cerevisiae strains with different genetic backgrounds were compared for their SA tolerances. KF7 and NBRC1958 with excellent SA tolerances, and NBRC2018 with poor SA tolerance, were selected to investigate the response mechanisms of S. cerevisiae to SA through comparative transcriptomic analysis. Few genes were significantly regulated under 20 g/L SA in three strains. When exposed to 60 g/L SA, the three strains showed different response mechanisms. Overall, the DEGs were involved in carbon metabolism, amino acid metabolism, protein folding, meiosis, membrane proteins and cell wall structure. We conclude that the genetic background of the host strain is important for the construction of good SA producing strains. The inherent activities of sHsps, acetyl-CoA levels and the potential SA consumption capacity of the host strains must be considered. This study provides theoretical guidance and tolerant strains for the breeding of robust SA-producing strains.Conclusion
Data availability
The dataset(s) used and/or analyzed during the current study are available from the corresponding author on reasonable request. The original microarray data can be accessed in the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/) through GEO accession GSE193190.
Abbreviations
- S. cerevisiae :
-
Saccharomyces cerevisiae
- SA:
-
Succinic acid
- PPP:
-
Pentose phosphate pathway
- NADPH:
-
Nicotinamide adenine dinucleotide phosphate
- PQC:
-
Protein quality control
- sHsps:
-
Small heat-shock proteins
- E. coli :
-
Escherichia coli
- DEGs:
-
Differentially expressed genes
- FPKM:
-
Fragments per kilobase of exon per million reads mapped
- FC:
-
Fold change
- DNA:
-
Deoxyribonucleic acid
- RNA:
-
Ribonucleic Acid
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- PPI:
-
Protein-protein interaction
- acetyl-CoA:
-
Acetyl coenzyme A
- TCA cycle:
-
Tricarboxylic acid cycle
- VB6:
-
Vitamin B6
- DSB:
-
DNA double-strand breaks
- PLP:
-
Pyridoxal 5′-phosphate
- TPP:
-
Thiamine pyrophosphate
- cAMP:
-
Cyclic AMP
- PKA:
-
Protein kinase A
- CWI:
-
Cell wall integrity
- ATP:
-
Adenosine-triphosphate
- ABC:
-
ATP-binding cassette
- PDR:
-
Pleiotropic drug resistance
- GID:
-
Glucose-induced degradation
- MFS:
-
Major facilitator superfamily
- MDR:
-
Multidrug resistance
- UPR:
-
Unfolded protein response
- LB:
-
Luria-Bertani
- OD600 :
-
Absorbance at 600 nm
- SGD:
-
Saccharomyces Genome Database
- 2-PE:
-
2-phenylethanol
References
Ahn JH, Jang YS, Lee SY. Production of succinic acid by metabolically engineered microorganisms. Curr Opin Biotechnol. 2016;42:54–66.
McKinlay JB, Vieille C, Zeikus JG. Prospects for a bio-based succinate industry. Appl Microbiol Biotechnol. 2007;76(4):727–40.
Li Q, Yang M, Wang D, Li W, Wu Y, Zhang Y, **ng J, Su Z. Efficient conversion of crop stalk wastes into succinic acid production by Actinobacillus succinogenes. Bioresour Technol. 2010;101(9):3292–4.
Lee SY, Kim JM, Song H, Lee JW, Kim TY, Jang YS. From genome sequence to integrated bioprocess for succinic acid production by Mannheimia succiniciproducens. Appl Microbiol Biotechnol. 2008;79(1):11–22.
Okino S, Noburyu R, Suda M, Jojima T, Inui M, Yukawa H. An efficient succinic acid production process in a metabolically engineered Corynebacterium glutamicum strain. Appl Microbiol Biotechnol. 2008;81(3):459–64.
Cimini D, Argenzio O, D’Ambrosio S, Lama L, Finore I, Finamore R, Pepe O, Faraco V, Schiraldi C. Production of succinic acid from Basfia succiniciproducens up to the pilot scale from Arundo donax hydrolysate. Bioresour Technol. 2016;222:355–60.
Podlesny M, Jarocki P, Wyrostek J, Czernecki T, Kucharska J, Nowak A, Targonski Z. Enterobacter Sp LU1 as a novel succinic acid producer - co-utilization of glycerol and lactose. Microb Biotechnol. 2017;10(2):492–501.
Jantama K, Haupt MJ, Svoronos SA, Zhang XL, Moore JC, Shanmugam KT, Ingram LO. Combining metabolic engineering and metabolic evolution to develop nonrecombinant strains of Escherichia coli C that produce succinate and malate. Biotechnol Bioeng. 2008;99(5):1140–53.
Agren R, Otero JM, Nielsen J. Genome-scale modeling enables metabolic engineering of Saccharomyces cerevisiae for succinic acid production. J Ind Microbiol Biotechnol. 2013;40(7):735–47.
Franco-Duarte R, Bessa D, Gonçalves F, Martins R, Silva-Ferreira AC, Schuller D, Sampaio P, Pais C. Genomic and transcriptomic analysis of isolates with focus in succinic acid production. FEMS Yeast Res. 2017; 17(6).
Ito Y, Hirasawa T, Shimizu H. Metabolic engineering of Saccharomyces cerevisiae to improve succinic acid production based on metabolic profiling. Biosci Biotechnol Biochem. 2014;78(1):151–9.
Yan DJ, Wang CX, Zhou JM, Liu YL, Yang MH, **ng JM. Construction of reductive pathway in Saccharomyces cerevisiae for effective succinic acid fermentation at low pH value. Bioresour Technol. 2014;156:232–9.
**berras J, Klein M, Hulster ED, Mans R, Nevoigt E. Engineering Saccharomyces cerevisiae for succinic acid production from glycerol and carbon dioxide. Front Bioeng Biotechnol. 2020;8:566.
Malubhoy Z, Bahia FM, de Valk SC, de Hulster E, Rendulic T, Ortiz JPR, **berras J, Klein M, Mans R, Nevoigt E. Carbon dioxide fixation via production of succinic acid from glycerol in engineered Saccharomyces cerevisiae. Microb Cell Fact. 2022; 21(1).
Rendulic T, Bahia FM, Soares-Silva I, Nevoigt E, Casal M. The dicarboxylate transporters from the AceTr family and Dct-02 oppositely affect succinic acid production in S. Cerevisiae. J Fungi. 2022; 8(8).
Van De Graaf MJ, Valianpoer F, Fiey G, Delattre L, EAM S. Process for the crystallization of succinic acid. US Patent 2015, US20150057425A1.
Wang G, Tan L, Sun ZY, Gou ZX, Tang YQ, Kida K. Production of bioethanol from rice straw by simultaneous saccharification and fermentation of whole pretreated slurry using Saccharomyces cerevisiae KF-7. Environ Prog Sustain. 2015;34(2):582–8.
Kida K, Kume K, Morimura S, Sonoda Y. Repeated-batch fermentation process using a thermotolerant flocculating yeast constructed by protoplast fusion. J Ferment Bioeng. 1992;74(3):169–73.
Mans R, van Rossum HM, Wijsman M, Backx A, Kuijpers NGA, van den Broek M, Daran-Lapujade P, Pronk JT, van Maris AJA, Daran JMG. CRISPR/Cas9: a molecular Swiss army knife for simultaneous introduction of multiple genetic modifications in Saccharomyces cerevisiae. FEMS Yeast Res. 2015; 15(2).
Zeng WY, Tang YQ, Gou M, **a ZY, Kida K. Transcriptomes of a xylose-utilizing industrial flocculating Saccharomyces cerevisiae strain cultured in media containing different sugar sources. AMB Express. 2016;6(1):51.
Gibson DG. Oligonucleotide assembly in yeast to produce synthetic DNA fragments. Methods Mol Biol. 2012;852:11–21.
Sibirny AA. Yeast peroxisomes: structure, functions and biotechnological opportunities. FEMS Yeast Res. 2016; 16(4).
Strijbis K, Distel B. Intracellular acetyl unit transport in fungal carbon metabolism. Eukaryot Cell. 2010;9(12):1809–15.
Fahien LA, MacDonald MJ. The succinate mechanism of insulin release. Diabetes. 2002;51(9):2669–76.
Kunze M, Pracharoenwattana I, Smith SM, Hartig A. A central role for the peroxisomal membrane in glyoxylate cycle function. Biochim Biophys Acta. 2006;1763(12):1441–52.
Chew SY, Chee WJY, Than LTL. The glyoxylate cycle and alternative carbon metabolism as metabolic adaptation strategies of Candida Glabrata: perspectives from Candida albicans and Saccharomyces cerevisiae. J Biomed Sci. 2019; 26.
Cai L, Tu BP. Driving the cell cycle through metabolism. Annu Rev Cell Dev Biol. 2012;28:59–87.
**ao W, Wang RS, Handy DE, Loscalzo J. NAD(H) and NADP(H) redox couples and cellular energy metabolism. Antioxid Redox Signal. 2018;28(3):251–72.
Kruger A, Gruning NM, Wamelink MMC, Kerick M, Kirpy A, Parkhomchuk D, Bluemlein K, Schweiger MR, Soldatov A, Lehrach H, et al. The pentose phosphate pathway is a metabolic redox sensor and regulates transcription during the antioxidant response. Antioxid Redox Signal. 2011;15(2):311–24.
Gruning NM, Lehrach H, Ralser M. Regulatory crosstalk of the metabolic network. Trends Biochem Sci. 2010;35(4):220–7.
Andersson R, Eisele-Burger AM, Hanzen S, Vielfort K, Oling D, Eisele F, Johansson G, Gustafsson T, Kvint K, Nystrom T. Differential role of cytosolic Hsp70s in longevity assurance and protein quality control. PLoS Genet. 2021; 17(1).
Amm I, Sommer T, Wolf DH. Protein quality control and elimination of protein waste: the role of the ubiquitin-proteasome system. Biochim Biophys Acta. 2014;1843(1):182–96.
Morrissette VA, Rolfes RJ. The intersection between stress responses and inositol pyrophosphates in Saccharomyces cerevisiae. Curr Genet. 2020;66(5):901–10.
Yoshida M, Kato S, Fukuda S, Izawa S. Acquired resistance to severe ethanol stress in Saccharomyces cerevisiae protein quality control. Appl Environ Microbiol. 2021; 87(6).
Brush GS, Najor NA, Dombkowski AA, Cukovic D, Sawarynski KE. Yeast IME2 functions early in meiosis upstream of cell cycle-regulated SBF and MBF targets. PLoS ONE. 2012; 7(2).
Zhang K, Wu XC, Zheng DQ, Petes TD. Effects of temperature on the meiotic recombination landscape of the yeast Saccharomyces cerevisiae. mBio. 2017; 8(6).
Wang HT, Frackman S, Kowalisyn J, Esposito RE, Elder R. Developmental regulation of SPO13, a gene required for separation of homologous chromosomes at meiosis I. Mol Cell Biol. 1987;7(4):1425–35.
Zeng WY, Tang YQ, Gou M, Sun ZY, **a ZY, Kida K. Comparative transcriptomes reveal novel evolutionary strategies adopted by Saccharomyces cerevisiae with improved xylose utilization capability. Appl Microbiol Biotechnol. 2017;101(4):1753–67.
Perli T, Wronska AK, Ortiz-Merino RA, Pronk JT, Daran JM. Vitamin requirements and biosynthesis in Saccharomyces cerevisiae. Yeast. 2020;37(4):283–304.
Mojzita D, Hohmann S. Pdc2 coordinates expression of the THI regulon in the yeast Saccharomyces cerevisiae. Mol Genet Genomics. 2006;276(2):147–61.
Labuschagne PWJ, Divol B. Thiamine: a key nutrient for yeasts during wine alcoholic fermentation. Appl Microbiol Biotechnol. 2021;105(3):953–73.
Rodriguez-Navarro S, Llorente B, Rodriguez-Manzaneque MT, Ramne A, Uber G, Marchesan D, Dujon B, Herrero E, Sunnerhagen P, Perez-Ortin JE. Functional analysis of yeast gene families involved in metabolism of vitamins B1 and B6. Yeast. 2002;19(14):1261–76.
Walvekar AS, Laxman S. Methionine at the heart of anabolism and signaling: perspectives from budding yeast. Front Microbiol. 2019; 10.
Walvekar AS, Srinivasan R, Gupta R, Laxman S. Methionine coordinates a hierarchically organized anabolic program enabling proliferation. Mol Biol Cell. 2018;29(26):3183–200.
Zhang MM, **ong L, Tang YJ, Mehmood MA, Zhao ZK, Bai FW, Zhao XQ. Enhanced acetic acid stress tolerance and ethanol production in Saccharomyces cerevisiae by modulating expression of the de novo purine biosynthesis genes. Biotechnol Biofuels. 2019; 12.
Avrahami-Moyal L, Braun S, Engelberg D. Overexpression of PDE2 or SSD1-V in Saccharomyces cerevisiae W303-1A strain renders it ethanol-tolerant. FEMS Yeast Res. 2012;12(4):447–55.
Thevelein JM, de Winde JH. Novel sensing mechanisms and targets for the cAMP-protein kinase A pathway in the yeast Saccharomyces cerevisiae. Mol Microbiol. 1999;33(5):904–18.
Nishida N, **g DY, Kuroda K, Ueda M. Activation of signaling pathways related to cell wall integrity and multidrug resistance by organic solvent in Saccharomyces cerevisiae. Curr Genet. 2014;60(3):149–62.
Liu ZL, Wang X, Weber SA. Tolerant industrial yeast Saccharomyces cerevisiae posses a more robust cell wall integrity signaling pathway against 2-furaldehyde and 5-(hydroxymethyl)-2-furaldehyde. J Biotechnol. 2018;276:15–24.
Harris A, Wagner M, Du DJ, Raschka S, Nentwig LM, Gohlke H, Smits SHJ, Luisi B, Schmitt L. Structure and efflux mechanism of the yeast pleiotropic drug resistance transporter Pdr5. Nat Commun. 2021; 12(1).
Nygard Y, Mojzita D, Toivari M, Penttila M, Wiebe MG, Ruohonen L. The diverse role of Pdr12 in resistance to weak organic acids. Yeast. 2014;31(6):219–32.
Abe F. Induction of DAN/TIR yeast cell wall mannoprotein genes in response to high hydrostatic pressure and low temperature. FEBS Lett. 2007;581(25):4993–8.
Negoro H, Kotaka A, Matsumura K, Tsutsumi H, Hata Y. Enhancement of malate-production and increase in sensitivity to dimethyl succinate by mutation of the VID24 gene in Saccharomyces cerevisiae. J Biosci Bioeng. 2016;121(6):665–71.
Li B, Wang L, Wu YJ, **a ZY, Yang BX, Tang YQ. Improving acetic acid and furfural resistance of xylose-fermenting Saccharomyces cerevisiae strains by regulating novel transcription factors revealed via comparative transcriptomic analysis. Appl Environ Microbiol. 2021; 87(10).
Hernandez-Elvira M, Martinez-Gomez R, Dominguez-Martin E, Mendez A, Kawasaki L, Ongay-Larios L, Coria R. Tunicamycin sensitivity-suppression by high gene dosage reveals new functions of the yeast Hog1 MAP kinase. Cells. 2019; 8(7).
Piper PW, Ortiz-Calderon C, Holyoak C, Coote P, Cole M. Hsp30, the integral plasma membrane heat shock protein of Saccharomyces cerevisiae, is a stress-inducible regulator of plasma membrane H(+)-ATPase. Cell Stress Chaperones. 1997;2(1):12–24.
Meena RC, Thakur S, Chakrabarti A. Regulation of Saccharomyces cerevisiae plasma membrane H(+)-ATPase (Pma1) by dextrose and Hsp30 during exposure to thermal stress. Indian J Microbiol. 2011;51(2):153–8.
Tulha J, Lima A, Lucas C, Ferreira C. Saccharomyces cerevisiae glycerol/H + symporter Stl1p is essential for cold/near-freeze and freeze stress adaptation. A simple recipe with high biotechnological potential is given. Microb Cell Fact. 2010;9:82.
Tenreiro S, Nunes PA, Viegas CA, Neves MS, Teixeira MC, Cabral MG, Sa-Correia I. AQR1 gene (ORF YNL065w) encodes a plasma membrane transporter of the major facilitator superfamily that confers resistance to short-chain monocarboxylic acids and quinidine in Saccharomyces cerevisiae. Biochem Biophys Res Commun. 2002;292(3):741–8.
Kwon YD, Kim S, Lee SY, Kim P. Long-term continuous adaptation of Escherichia coli to high succinate stress and transcriptome analysis of the tolerant strain. J Biosci Bioeng. 2011;111(1):26–30.
Ungelenk S, Moayed F, Ho CT, Grousl T, Scharf A, Mashaghi A, Tans S, Mayer MP, Mogk A, Bukau B. Small heat shock proteins sequester misfolding proteins in near-native conformation for cellular protection and efficient refolding. Nat Commun. 2016;7:13673.
Lytras G, Zacharioudakis I, Tzamarias D. Asymmetric inheritance of the yeast chaperone Hsp26p and its functional consequences. Biochem Biophys Res Commun. 2017;491(4):1055–61.
Wang J, Pareja KA, Kaiser CA, Sevier CS. Redox signaling via the molecular chaperone BiP protects cells against endoplasmic reticulum-derived oxidative stress. Elife. 2014; 3.
Verghese J, Abrams J, Wang YY, Morano KA. Biology of the heat shock response and protein chaperones: budding yeast (Saccharomyces cerevisiae) as a model system. Microbiol Mol Biol Rev. 2012;76(2):115–58.
Campion R, Bloxam L, Burrow K, Brownridge PJ, Pentland DR, Thomas P, Gourlay CW, Eyers CE, Barclay JW, Morgan A. Proteomic analysis of dietary restriction in yeast reveals a role for Hsp26 in replicative lifespan extension. Biochem J. 2021;478(24):4153–67.
de Nobel H, Lawrie L, Brul S, Klis F, Davis M, Alloush H, Coote P. Parallel and comparative analysis of the proteome and transcriptome of sorbic acid-stressed Saccharomyces cerevisiae. Yeast. 2001;18(15):1413–28.
Lawrence CL, Botting CH, Antrobus R, Coote PJ. Evidence of a new role for the high-osmolarity glycerol mitogen-activated protein kinase pathway in yeast: regulating adaptation to citric acid stress. Mol Cell Biol. 2004;24(8):3307–23.
Feng X, Zhao H. Investigating host dependence of xylose utilization in recombinant Saccharomyces cerevisiae strains using RNA-seq analysis. Biotechnol Biofuels. 2013;6(1):96.
Sardi M, Rovinskiy N, Zhang Y, Gasch AP. Leveraging genetic-background effects in Saccharomyces cerevisiae to improve lignocellulosic hydrolysate tolerance. Appl Environ Microbiol. 2016;82(19):5838–49.
Ndukwe JK, Aliyu GO, Onwosi CO, Chukwu KO, Ezugworie FN. Mechanisms of weak acid-induced stress tolerance in yeasts: prospects for improved bioethanol production from lignocellulosic biomass. Process Biochem. 2020;90:118–30.
Mira NP, Teixeira MC, Sa-Correia I. Adaptive response and tolerance to weak acids in Saccharomyces cerevisiae: a genome-wide view. OMICS. 2010;14(5):525–40.
Ribeiro RA, Bourbon-Melo N, Sa-Correia I. The cell wall and the response and tolerance to stresses of biotechnological relevance in yeasts. Front Microbiol. 2022; 13.
Zeng L, Huang J, Feng P, Zhao X, Si Z, Long X, Cheng Q, Yi Y. Transcriptomic analysis of formic acid stress response in Saccharomyces cerevisiae. World J Microbiol Biotechnol. 2022;38(2):34.
Li B, **e C, Yang B, Gou M, **a Z, Sun Z, Tang Y. The response mechanisms of industrial Saccharomyces cerevisiae to acetic acid and formic acid during mixed glucose and xylose fermentation. Process Biochem. 2020;91:319–29.
Kawazoe N, Kimata Y, Izawa S. Acetic acid causes endoplasmic reticulum stress and induces the unfolded protein response in Saccharomyces cerevisiae. Front Microbiol. 2017;8:1192.
Berterame NM, Porro D, Ami D, Branduardi P. Protein aggregation and membrane lipid modifications under lactic acid stress in wild type and OPI1 deleted Saccharomyces cerevisiae strains. Microb Cell Fact. 2016;15:39.
Peetermans A, Foulquie-Moreno MR, Thevelein JM. Mechanisms underlying lactic acid tolerance and its influence on lactic acid production in Saccharomyces cerevisiae. Microb Cell. 2021;8(6):111–30.
Deng N, Du H, Xu Y. Cooperative response of Pichia kudriavzevii and Saccharomyces cerevisiae to lactic acid stress in Baijiu fermentation. J Agric Food Chem. 2020;68(17):4903–11.
Lin AP, Anderson SL, Minard KI, McAlister-Henn L. Effects of excess succinate and retrograde control of metabolite accumulation in yeast tricarboxylic cycle mutants. J Biol Chem. 2011;286(39):33737–46.
Terzioğlu E, Alkım C, Arslan M, Balaban BG, Holyavkin C, Kısakesen H, Topaloğlu A, Yılmaz Şahin Ü, Gündüz Işık S, Akman S, et al. Genomic, transcriptomic and physiological analyses of silver-resistant Saccharomyces cerevisiae obtained by evolutionary engineering. Yeast. 2020;37(9–10):413–26.
Holyavkin C, Turanli-Yildiz B, Yilmaz Ü, Alkim C, Arslan M, Topaloglu A, Kisakesen HI, de Billerbeck G, François JM, Çakar ZP. Genomic, transcriptomic, and metabolic characterization of 2-Phenylethanol-resistant obtained by evolutionary engineering. Front Microbiol. 2023; 14.
Sürmeli Y, Holyavkin C, Topaloglu A, Arslan M, Kisakesen HI, Çakar ZP. Evolutionary engineering and molecular characterization of a caffeine-resistant strain. World J Microb Biot. 2019; 35(12).
Kocaefe-Özsen N, Yilmaz B, Alkim C, Arslan M, Topaloglu A, Kisakesen HL, Gulsev E, Cakar ZP. Physiological and molecular characterization of an oxidative stress-resistant strain obtained by evolutionary engineering. Front Microbiol. 2022; 13.
Acknowledgements
Not applicable.
Funding
This study was financially supported by the National Key R&D Program of China (2022YFE0108500) and the National Natural Science Foundation of China (52300169).
Author information
Authors and Affiliations
Contributions
XCY and WB conducted experiments. SRR and XCY analyzed data and wrote the main manuscript. SZY provided technical assistance during data analysis. TYQ designed the study and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
**e, CY., Su, RR., Wu, B. et al. Response mechanisms of different Saccharomyces cerevisiae strains to succinic acid. BMC Microbiol 24, 158 (2024). https://doi.org/10.1186/s12866-024-03314-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12866-024-03314-4