Abstract
Background
Preeclampsia is a pregnancy-related condition that causes high blood pressure and proteinuria after 20 weeks of pregnancy. It is linked to increased maternal mortality, organ malfunction, and foetal development limitation. In this view, there is a need critical to identify biomarkers for the early detection of preeclampsia. The objective of this study is to discover critical genes and explore medications for preeclampsia treatment that may influence these genes.
Methods
Four datasets, including GSE10588, GSE25906, GSE48424 and GSE60438 were retrieved from the Gene Expression Omnibus database. The GSE10588, GSE25906, and GSE48424 datasets were then removed the batch effect using the “sva” R package and merged into a complete dataset. The differentially expressed genes (DEGs) were identified using the “limma” R package. The potential small-molecule agents for the treatment of PE was further screened using the Connective Map (CMAP) drug database based on the DEGs. Further, Weight gene Co-expression network (WGNCA) analysis was performed to identified gene module associated with preeclampsia, hub genes were then identified using the logistic regression analysis. Finally, the immune cell infiltration level of genes was evaluated through the single sample gene set enrichment analysis (ssGSEA).
Results
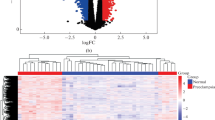
A total of 681 DEGs (376 down-regulated and 305 up-regulated genes) were identified between normal and preeclampsia samples. Then, Dexamethasone, Prednisone, Rimexolone, Piretanide, Trazodone, Buflomedil, Scoulerin, Irinotecan, and Camptothecin drugs were screened based on these DEGs through the CMAP database. Two modules including yellow and brown modules were the most associated with disease through the WGCNA analysis. KEGG analysis revealed that the chemokine signaling pathway, Th1 and Th2 cell differentiation, B cell receptor signalling pathway and oxytocin signalling pathway were significantly enriched in these modules. Moreover, two key genes, PLEK and LEP were evaluated using the univariate and multivariate logistic regression analysis from the hub modules. These two genes were further validated in the external validation cohort GSE60438 and qRT-PCR experiment. Finally, we evaluated the relationship between immune cell and two genes.
Conclusion
In conclusion, the present study investigated key genes associated with PE pathogenesis that may contribute to identifying potential biomarkers, therapeutic agents and develo** personalized treatment for PE.
Similar content being viewed by others
Introduction
Preeclampsia (PE) is a pregnancy disorder that causes high blood pressure and proteinuria [1] after 20 weeks of pregnancy [2].It affects 3%-10% of all pregnancies [3,4,5], and is associated with substantial maternal morbidity, mortality, organ dysfunction, iatrogenic premature delivery [6], and foetal development restriction [7]. Preeclampsia affects the brain development and function of the offspring, increasing the risk of intellectual disability [8], epilepsy [9], autism [10], and schizophrenia [11, 12]. Women with a history of PE are more likely to develop cardiovascular disease or hypertension,5 resulting in financial and psychological consequences on the family and society. There is no obvious therapeutic intervention for PE today; the only effective treatment is pregnancy, which might result in low birth weight and long-term detrimental health implications for infants [13, 14]. Early assessment of PE risk is critical for high-risk pregnant women because it allows for the implementation of preventive strategies to lower the incidence of PE and improve maternal and infant outcomes.
However, the etiology and pathogenesis of PE are yet unknown. Several hypotheses such as maternal-foetal (paternal) immune maladjustment [15] and inflammatory cytokine disorders [16], all suggest that PE is associated with a number of risk factors, including obesity, hypertension, diabetes, oxidative stress, foetal rejection, genetic polymorphism inheritance [17], and trophoblast insufficiency [18, 19]. The main causes are identified as immunological intolerance [20, 21] and angiogenesis imbalance, and numerous research has demonstrated the immune mechanism of PE development. The latter causes PE by generating an imbalance in immune tolerance at the mother-infant interaction [22, 23]. The pathophysiological basis of PE is shallow placental implantation. The mutual adaptation of villous trophoblast cells and the mother's immune system is required for effective placental development. The poor placental formation will result in acute-like graft rejection disease under the influence of specific immunological factors generating immune intolerance in the mother and child.
Immune system modifications have been widely recognized as the key determinants of PE [24], which is a systemic inflammatory response resulting in an imbalance between placental substances and the corresponding adaptation of the mother[25, 26]. Shah et al. [27] in their study compared the expression of CD66B, nuclear factor NF-κB, and cyclooxygenase-2 (COX-2) in the subcutaneous fat of women with PE (n = 7), normal pregnant women (n = 6), and normal non-pregnant women (n = 5). The percentages of CD66B, NF-κB, and COX-2 in PE patients were significantly higher than in normal non-pregnant or normal pregnant patients. Moreover, Xu et al. [52]. Leptin injection raised ICAM-1 and Eseltin circulation concentrations, resulting in hypertension and proteinuria in pregnant rats [53]. Our findings show that the expression level of this gene was higher in the placenta of the PE group than in the control group, which is consistent with earlier findings. PLEK, another hub gene, was found to be significantly expressed in the merged dataset. Its expression was higher in normal tissues than in PE tissues, although it was not significantly higher in the external validation set. Pleckstrin (PLEK), which is found on human chromosome 2, is a protein kinase C target [54] that is involved in signal transduction and hematopoietic cell differentiation [55]. PLEK play an important role in immune and inflammatory responses [56, 57]. Studies have been shown that PLEK is significantly over-expressed in periodontitis, cardiovascular disease, rheumatoid arthritis, and ulcerative colitis [58], and thought to be a crucial mediator in the secretion and activation pathways of pro-inflammatory cytokines TNF-α and IL-1β [59, 60]. While studies have suggested that LILRA2, EVI2A, and PLEK play a role in recurrent miscarriage [61], the underlying mechanism in placental function and regulation has yet to be fully investigated. This is the first study to examine PLEK expression in PE. An examination of its mechanisms would necessitate additional research.
PE is a complex systemic condition, and it has been shown that the immune system plays a significant role in its development [62]. As a result, we evaluated the landscape of 29 immune cell infiltration levels in PE and control samples. The results indicated that the infiltration level of neutrophils, T helper cells, Th2 cells, TIL, and Treg cells were significantly differed between PE and normal tissues. Immune cell infiltration is a new bioinformatics technique that has been used to investigate the diagnosis and prognosis of kidney cancer [63], malignant glioma [64], breast cancer [65], oral squamous cell carcinoma [66], ulcerative colitis[67], osteosarcoma [68], and variety of other diseases. Nonetheless, it has received little attention in the field of PE. The complicated connection between the maternal immune system and its semi-allogeneic foetus is critical in normal pregnancy. Moreover, establishing and maintaining the maternal and foetal immune balance is a prerequisite for normal pregnancy [69]. Abnormal maternal immunological and inflammatory responses to foetal antigens result in increased release of various toxic cytokines, which causes trophoblast cell invasion, vascular remodelling, and placental implantation disorders. Changes in the innate immune system primarily regulate this inflammatory response, with the adaptive immune system possibly playing a supporting role [70].
T cells are thought to be the most important cells in regulating immunological homeostasis [71, 72]. T lymphocytes account for 1%-3% of decidual immune cells [73]. In a normal pregnancy, the mother exhibits Th2 cell-type immunological tolerance, preventing embryo rejection [74]. However, in PE patients, the Th1/Th2 ratio increases. As a result, the Th1/Th2 balance changes towards Th1 [75]. Aside from the Th1/Th2 imbalance that contributes to the onset and progression of the disease, there is also an imbalance of Th17/ regulatory T cells. This imbalance, which was exacerbated by the Th17 immune bias, also contributed to the development of PE. Th17/ Treg cells are balanced at the maternal-foetal interface during normal pregnancy to preserve maternal immunological tolerance and inhibit the inflammatory response [76]. In our study, we identified most of T cells showed a significant difference between normal and PE samples, reveled that the T cells play an important role in PE.
Conclusion
PLEK and LEP were identified as two genes implicated in the development and progression of PE in this investigation. Although more in vivo and in vitro validations are needed, our findings help to understand the pathological process of PE and may serve as a theoretical foundation for future research. The functional annotation and pathway enrichment analysis results show that the immunological mechanism is important in the etiology of PE. Also, because of the maternal and infant complications of PE, it is critical to uncover the aetiology and molecular mechanism, develop molecular biomarkers and investigate effective drugs for the early detection, prevention, and personalized treatment of PE.
Limitations
This study not only points out several benefits of bioinformatics analysis but also highlights some limitations. The dependability of the original microarray dataset is critical to the validity of our conclusions, although the results are constrained due to the small sample size. Similarly, validation results are limited. Second, despite the identification of two hub genes as prospective biomarkers for PE immunoty**, no in vivo and in vitro studies have been conducted. More research on the functions and regulatory mechanisms of key genes in PE is still needed. As a result, this will be the focus of future efforts.
Availability of data and materials
The datasets that support the findings of this study are openly available in the Gene Expression Omnibus (GEO) database under accession ID GSE10588, GSE25906, GSE48424 and GSE60438, [http://www.ncbi.nlm.nih.gov/geo/].
Abbreviations
- PE:
-
Preeclampsia
- GEO:
-
Gene Expression Omnibus
- DEGs:
-
Differentially expressed genes
- WGCNA:
-
Weighted gene co-expression network analysis
- PPI:
-
Protein–protein interaction network
- CMAP:
-
Connectivity map
- TOM:
-
Topological overlap matrix
- GS:
-
Gene significance
- MEs:
-
Module eigengenes
- MM:
-
Module membership
- GO:
-
Gene ontology
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- ssGSEA:
-
Single-sample Gene Set Enrichment Analysis
- MOA:
-
Mechanism of actions
- ROC:
-
Receiver operating characteristic
- AUC:
-
Area under the curve
- CCR:
-
Chemokine receptor
- TIL:
-
Tumor infiltrating lymphocyte
- MCC:
-
Maximal clique centrality
References
Eiland E, Nzerue C, Faulkner M. Preeclampsia 2012. J Pregnancy. 2012;2012:586578.
Al-Jameil N, Aziz Khan F, Fareed Khan M, Tabassum H. A brief overview of preeclampsia. J Clin Med Res. 2014;6(1):1–7.
Vatten LJ, Skjaerven R. Offspring sex and pregnancy outcome by length of gestation. Early Hum Dev. 2004;76(1):47–54.
Hariharan N, Shoemaker A, Wagner S. Pathophysiology of hypertension in preeclampsia. Microvasc Res. 2017;109:34–7.
Perry H, Khalil A, Thilaganathan B. Preeclampsia and the cardiovascular system: An update. Trends Cardiovasc Med. 2018;28(8):505–13.
Mol BWJ, Roberts CT, Thangaratinam S, Magee LA, de Groot CJM, Hofmeyr GJ. Pre-eclampsia. The Lancet. 2016;387(10022):999–1011.
Genest DS, Falcao S, Gutkowska J, Lavoie JL. Impact of exercise training on preeclampsia: potential preventive mechanisms. Hypertens. 2012;60(5):1104–9.
Griffith MI, Mann JR, McDermott S. The risk of intellectual disability in children born to mothers with preeclampsia or eclampsia with partial mediation by low birth weight. Hypertens Pregnancy. 2011;30(1):108–15.
Wu CS, Sun Y, Vestergaard M, Christensen J, Ness RB, Haggerty CL, Olsen J. Preeclampsia and risk for epilepsy in offspring. Pediatr. 2008;122(5):1072–8.
Walker CK, Krakowiak P, Baker A, Hansen RL, Ozonoff S, Hertz-Picciotto I. Preeclampsia, placental insufficiency, and autism spectrum disorder or developmental delay. JAMA Pediatr. 2015;169(2):154–62.
Dalman C, Allebeck P, Cullberg J, Grunewald C, Koster M. Obstetric complications and the risk of schizophrenia: a longitudinal study of a national birth cohort. Arch Gen Psychiatry. 1999;56(3):234–40.
Ursini G, Punzi G, Chen Q, Marenco S, Robinson JF, Porcelli A, Hamilton EG, Mitjans M, Maddalena G, Begemann M, et al. Convergence of placenta biology and genetic risk for schizophrenia. Nat Med. 2018;24(6):792–801.
Coppage KH, Polzin WJ. Severe preeclampsia and delivery outcomes: is immediate cesarean delivery beneficial? Am J Obstet Gynecol. 2002;186(5):921–3.
Leavey K, Wilson SL, Bainbridge SA, Robinson WP, Cox BJ. Epigenetic regulation of placental gene expression in transcriptional subtypes of preeclampsia. Clin Epigenetics. 2018;10:28.
Saito S, Shiozaki A, Nakashima A, Sakai M, Sasaki Y. The role of the immune system in preeclampsia. Mol Aspects Med. 2007;28(2):192–209.
Ramma W, Buhimschi IA, Zhao G, Dulay AT, Nayeri UA, Buhimschi CS, Ahmed A. The elevation in circulating anti-angiogenic factors is independent of markers of neutrophil activation in preeclampsia. Angiogenesis. 2012;15(3):333–40.
Phipps EA, Thadhani R, Benzing T, Karumanchi SA. Pre-eclampsia: pathogenesis, novel diagnostics and therapies. Nat Rev Nephrol. 2019;15(5):275–89.
Agius A, Sultana R, Camenzuli C, Calleja-Agius J, Balzan R. An update on the genetics of pre-eclampsia. Minerva Ginecol. 2018;70(4):465–79.
Spinillo A, Gardella B, Adamo L, Muscettola G, Fiandrino G, Cesari S. Pathologic placental lesions in early and late fetal growth restriction. Acta Obstet Gynecol Scand. 2019;98(12):1585–94.
Lynch A, McDuffie R Jr, Murphy J, Faber K, Orleans M. Preeclampsia in multiple gestation: the role of assisted reproductive technologies. Obstet Gynecol. 2002;99(3):445–51.
Wimalasundera RC, Larbalestier N, Smith JH, de Ruiter A, Mc GTSA, Hughes AD, Poulter N, Regan L, Taylor GP. Pre-eclampsia, antiretroviral therapy, and immune reconstitution. Lancet. 2002;360(9340):1152–4.
Erlebacher A. Immunology of the maternal-fetal interface. Annu Rev Immunol. 2013;31:387–411.
Nancy P, Erlebacher A. T cell behavior at the maternal-fetal interface. Int J Dev Biol. 2014;58(2–4):189–98.
Gidlof S, Nisell H. Pre-eclampsia. Lakartidningen. 2010;107(51–52):3288–92.
LaMarca B, Cornelius D, Wallace K. Elucidating immune mechanisms causing hypertension during pregnancy. Physiology (Bethesda). 2013;28(4):225–33.
Ma Y, Ye Y, Zhang J, Ruan CC, Gao PJ. Immune imbalance is associated with the development of preeclampsia. Medicine (Baltimore). 2019;98(14):e15080.
Shah TJ, Walsh SW. Activation of NF-kappaB and expression of COX-2 in association with neutrophil infiltration in systemic vascular tissue of women with preeclampsia. Am J Obstet Gynecol. 2007;196(1):48 e41-48.
Xu H, **e Y, Sun Y, Guo R, Lv D, Li X, Li F, He M, Fan Y, Deng D. Integrated analysis of multiple microarray studies to identify potential pathogenic gene modules in preeclampsia. Exp Mol Pathol. 2021;120:104631.
Liu K, Fu Q, Liu Y, Wang C. An integrative bioinformatics analysis of microarray data for identifying hub genes as diagnostic biomarkers of preeclampsia. Biosci Rep. 2019;39(9):1-10.
Lin J, Meng Y, Song MF, Gu W. Network-Based Analysis Reveals Novel Biomarkers in Peripheral Blood of Patients With Preeclampsia. Front Mol Biosci. 2022;9:757203.
Wang Y, Li B, Zhao Y. Inflammation in Preeclampsia: Genetic Biomarkers, Mechanisms, and Therapeutic Strategies. Front Immunol. 2022;13:883404.
Sitras V, Paulssen RH, Gronaas H, Leirvik J, Hanssen TA, Vartun A, Acharya G. Differential placental gene expression in severe preeclampsia. Placenta. 2009;30(5):424–33.
Tsai S, Hardison NE, James AH, Motsinger-Reif AA, Bischoff SR, Thames BH, Piedrahita JA. Transcriptional profiling of human placentas from pregnancies complicated by preeclampsia reveals disregulation of sialic acid acetylesterase and immune signalling pathways. Placenta. 2011;32(2):175–82.
Textoris J, Ivorra D, Ben Amara A, Sabatier F, Menard JP, Heckenroth H, Bretelle F, Mege JL. Evaluation of current and new biomarkers in severe preeclampsia: a microarray approach reveals the VSIG4 gene as a potential blood biomarker. PLoS ONE. 2013;8(12):e82638.
Yong HE, Melton PE, Johnson MP, Freed KA, Kalionis B, Murthi P, Brennecke SP, Keogh RJ, Moses EK. Genome-wide transcriptome directed pathway analysis of maternal pre-eclampsia susceptibility genes. PLoS ONE. 2015;10(5):e0128230.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28(1):27–30.
Kanehisa M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 2019;28(11):1947–51.
Kanehisa M, Furumichi M, Sato Y, Ishiguro-Watanabe M, Tanabe M. KEGG: integrating viruses and cellular organisms. Nucleic Acids Res. 2021;49(D1):D545–51.
He Y, Jiang Z, Chen C, Wang X. Classification of triple-negative breast cancers based on Immunogenomic profiling. J Exp Clin Cancer Res. 2018;37(1):327.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402–8.
Chen X, Yan CC, Zhang X, You ZH. Long non-coding RNAs and complex diseases: from experimental results to computational models. Brief Bioinform. 2017;18(4):558–76.
Wang CC, Han CD, Zhao Q, Chen X. Circular RNAs and complex diseases: from experimental results computational models. Brief Bioinform. 2021;22(6):1-27.
Chen X, **e D, Zhao Q, You ZH. MicroRNAs and complex diseases: from experimental results to computational models. Brief Bioinform. 2019;20(2):515–39.
Chen X, Sun LG, Zhao Y. NCMCMDA: miRNA-disease association prediction through neighborhood constraint matrix completion. Brief Bioinform. 2021;22(1):485–96.
Chen X, Li TH, Zhao Y, Wang CC, Zhu CC. Deep-belief network for predicting potential miRNA-disease associations. Brief Bioinform. 2021;22(3):bbaa186.
Sun N, Qin S, Zhang L, Liu S. Roles of noncoding RNAs in preeclampsia. Reprod Biol Endocrinol. 2021;19(1):100.
Rokhafrooz S, Ghadiri A, Ghandil P, Ghafourian M, Hossaini SH, Daraei N, Najafian M, Rouhizadeh A. Association between HLA-G 14bp Gene Polymorphism and Serum sHLA-G Protein Concentrations in Preeclamptic Patients and Normal Pregnant Women. Immunol Invest. 2018;47(6):558–68.
Miehle K, Stepan H, Fasshauer M. Leptin, adiponectin and other adipokines in gestational diabetes mellitus and pre-eclampsia. Clin Endocrinol (Oxf). 2012;76(1):2–11.
Hauguel-de Mouzon S, Lepercq J, Catalano P. The known and unknown of leptin in pregnancy. Am J Obstet Gynecol. 2006;194(6):1537–45.
Cao R, Brakenhielm E, Wahlestedt C, Thyberg J, Cao Y. Leptin induces vascular permeability and synergistically stimulates angiogenesis with FGF-2 and VEGF. Proc Natl Acad Sci U S A. 2001;98(11):6390–5.
Muy-Rivera M, Ning Y, Frederic IO, Vadachkoria S, Luthy DA, Williams MA. Leptin, soluble leptin receptor and leptin gene polymorphism in relation to preeclampsia risk. Physiol Res. 2005;54(2):167–74.
Prior LJ, Eikelis N, Armitage JA, Davern PJ, Burke SL, Montani JP, Barzel B, Head GA. Exposure to a high-fat diet alters leptin sensitivity and elevates renal sympathetic nerve activity and arterial pressure in rabbits. Hypertension. 2010;55(4):862–8.
Ibrahim HS, Omar E, Froemming GR, Singh HJ. Leptin increases blood pressure and markers of endothelial activation during pregnancy in rats. Biomed Res Int. 2013;2013:298401.
Tyers M, Haslam RJ, Rachubinski RA, Harley CB. Molecular analysis of pleckstrin: the major protein kinase C substrate of platelets. J Cell Biochem. 1989;40(2):133–45.
Kienzle N, Young D, Silins SL, Sculley TB. Induction of pleckstrin by the Epstein-Barr virus nuclear antigen 3 family. Virology. 1996;224(1):167–74.
Liu Y, Wang L, Predina J, Han R, Beier UH, Wang LC, Kapoor V, Bhatti TR, Akimova T, Singhal S, et al. Inhibition of p300 impairs Foxp3(+) T regulatory cell function and promotes antitumor immunity. Nat Med. 2013;19(9):1173–7.
Ubel C, Sopel N, Graser A, Hildner K, Reinhardt C, Zimmermann T, Rieker RJ, Maier A, Neurath MF, Murphy KM, et al. The activating protein 1 transcription factor basic leucine zipper transcription factor, ATF-like (BATF), regulates lymphocyte- and mast cell-driven immune responses in the setting of allergic asthma. J Allergy Clin Immunol. 2014;133(1):198-206 e191-199.
Lundmark A, Davanian H, Bage T, Johannsen G, Koro C, Lundeberg J, Yucel-Lindberg T. Transcriptome analysis reveals mucin 4 to be highly associated with periodontitis and identifies pleckstrin as a link to systemic diseases. Sci Rep. 2015;5:18475.
Ding Y, Kantarci A, Badwey JA, Hasturk H, Malabanan A, Van Dyke TE. Phosphorylation of pleckstrin increases proinflammatory cytokine secretion by mononuclear phagocytes in diabetes mellitus. J Immunol. 2007;179(1):647–54.
Hasturk H, Kantarci A, Van Dyke TE. Oral inflammatory diseases and systemic inflammation: role of the macrophage. Front Immunol. 2012;3:118.
Chen H, Cheng S, Liu C, Fu J, Huang W. Bioinformatics Analysis of Differentially Expressed Genes, Methylated Genes, and miRNAs in Unexplained Recurrent Spontaneous Abortion. J Comput Biol. 2019;26(12):1418–26.
Lu HQ, Hu R. The role of immunity in the pathogenesis and development of pre-eclampsia. Scand J Immunol. 2019;90(5):e12756.
Geissler K, Fornara P, Lautenschlager C, Holzhausen HJ, Seliger B, Riemann D. Immune signature of tumor infiltrating immune cells in renal cancer. Oncoimmunology. 2015;4(1):e985082.
Matsuhisa T. Subset analysis of tumor infiltrating lymphocytes and peripheral blood lymphocytes in malignant glioma patients. No To Shinkei. 1995;47(5):466–73.
Yao S, Cheng TD, Elkhanany A, Yan L, Omilian A, Abrams SI, Evans S, Hong CC, Qi Q, Davis W, et al. Breast Tumor Microenvironment in Black Women: A Distinct Signature of CD8+ T-Cell Exhaustion. J Natl Cancer Inst. 2021;113(8):1036–43.
Parikh A, Shin J, Faquin W, Lin DT, Tirosh I, Sunwoo JB, Puram SV. Malignant cell-specific CXCL14 promotes tumor lymphocyte infiltration in oral cavity squamous cell carcinoma. J Immunother Cancer. 2020;8(2):e001048.
Xue G, Hua L, Zhou N, Li J. Characteristics of immune cell infiltration and associated diagnostic biomarkers in ulcerative colitis: results from bioinformatics analysis. Bioengineered. 2021;12(1):252–65.
Fu Y, Bao Q, Liu Z, He G, Wen J, Liu Q, Xu Y, ** Z, Zhang W. Development and Validation of a Hypoxia-Associated Prognostic Signature Related to Osteosarcoma Metastasis and Immune Infiltration. Front Cell Dev Biol. 2021;9:633607.
Perez-Sepulveda A, Torres MJ, Khoury M, Illanes SE. Innate immune system and preeclampsia. Front Immunol. 2014;5:244.
Borzychowski AM, Sargent IL, Redman CW. Inflammation and pre-eclampsia. Semin Fetal Neonatal Med. 2006;11(5):309–16.
Yang F, Zheng Q, ** L. Dynamic Function and Composition Changes of Immune Cells During Normal and Pathological Pregnancy at the Maternal-Fetal Interface. Front Immunol. 2019;10:2317.
Wang Z, Song K, Zhao W, Zhao Z. Dendritic cells in tumor microenvironment promoted the neuropathic pain via paracrine inflammatory and growth factors. Bioengineered. 2020;11(1):661–78.
Lessin DL, Hunt JS, King CR, Wood GW. Antigen expression by cells near the maternal-fetal interface. Am J Reprod Immunol Microbiol. 1988;16(1):1–7.
Perricone C, de Carolis C, Perricone R. Pregnancy and autoimmunity: a common problem. Best Pract Res Clin Rheumatol. 2012;26(1):47–60.
Vargas-Rojas MI, Solleiro-Villavicencio H, Soto-Vega E. Th1, Th2, Th17 and Treg levels in umbilical cord blood in preeclampsia. J Matern Fetal Neonatal Med. 2016;29(10):1642–5.
Eghbal-Fard S, Yousefi M, Heydarlou H, Ahmadi M, Taghavi S, Movasaghpour A, Jadidi-Niaragh F, Yousefi B, Dolati S, Hojjat-Farsangi M, et al. The imbalance of Th17/Treg axis involved in the pathogenesis of preeclampsia. J Cell Physiol. 2019;234(4):5106–16.
Acknowledgements
The researchers thank the obstetrics and gynaecology department, surgery department, and laboratory department of Qilu Hospital and The First Affiliated Hospital of USTC for their support during the data collection process.
Funding
No sources of funding exist for this research. All costs were covered by the researchers.
Author information
Authors and Affiliations
Contributions
YP wrote the proposal, participated in data collection, analyzed the data, and drafted the paper. NG,HH and AW approved the proposal with some revisions, participated in data analysis, and revised subsequent drafts of the paper. YM designed the entire study, provided administrative, technical, and material support, and gave a critical review of the intellectual content of the article. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Written informed consent was obtained from all participants. This study was approved by The First Affiliated Hospital of USTC, Division of Life Sciences and Medicine, University of Science and Technology of China (Anhui Provincial Hospital) research ethics board (NO:2021KY161) and is based on the ethical requirements of the Helsinki Declaration. All participants have the right to know.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Peng, Y., Hong, H., Gao, N. et al. Bioinformatics methods in biomarkers of preeclampsia and associated potential drug applications. BMC Genomics 23, 711 (2022). https://doi.org/10.1186/s12864-022-08937-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12864-022-08937-3