Abstract
Background
Primary cilia are microtubule-based organelles that protrude from the cell surface. Primary cilia play a critical role in development and disease through regulation of signaling pathways including the Hedgehog pathway. Recent mouse models have also linked ciliary dysfunction to cancer. However, little is known about the role of primary cilia in breast cancer development. Primary cilia expression was characterized in cancer cells as well as their surrounding stromal cells from 86 breast cancer patients by counting cilia and measuring cilia length. In addition, we examined cilia expression in normal epithelial and stromal cells from reduction mammoplasties as well as histologically normal adjacent tissue for comparison.
Results
We observed a statistically significant decrease in the percentage of ciliated cells on both premalignant lesions as well as in invasive cancers. This loss of cilia does not correlate with increased proliferative index (Ki67-positive cells). However, we did detect rare ciliated cancer cells present in patients with invasive breast cancer and found that these express a marker of basaloid cancers that is associated with poor prognosis (Cytokeratin 5). Interestingly, the percentage of ciliated stromal cells associated with both premalignant and invasive cancers decreased when compared to stromal cells associated with normal tissue. To understand how cilia may be lost during cancer development we analyzed the expression of genes required for ciliogenesis and/or ciliary function and compared their expression in normal versus breast cancer samples. We found that expression of ciliary genes were frequently downregulated in human breast cancers.
Conclusions
These data suggest that primary cilia are lost early in breast cancer development on both the cancer cells and their surrounding stromal cells.
Similar content being viewed by others
Background
Primary cilia are hair-like projections that extend from the plasma membrane and are found on many types of normal eukaryotic cells. Cells with primary cilia have a single cilium that is non-motile. Ciliary structure includes the basal body (also known as the mother centriole), which is anchored into the plasma membrane to nucleate microtubules of the ciliary axoneme. Ciliary assembly (or ciliogenesis) is regulated by a process known as intraflagellar transport (IFT), by which proteins are trafficked to the cilium and move along the ciliary microtubules via kinesin and dynein. IFT also regulates ciliary sensory functions, allowing the organelle to respond to physical and chemical signals to regulate critical signaling transduction pathways; for example, cilia are both negative and positive regulators of the Hedgehog pathway [1]. Cilia are known to play critical roles in developmental biology and mutations in genes required for ciliogenesis or ciliary function are associated with human genetic disorders collectively known as ciliopathies (for example, Joubert syndrome, polycystic kidney disease, Bardet-Biedl syndrome, and nephronophthisis) [2].
The role primary cilia play in cancer is not well understood [3, 4]. Mouse models demonstrate that loss of cilia can increase tumor incidence in basal cell carcinoma and medulloblastoma [5–7]. Cilia expression has been analyzed in some human cancers, demonstrating that pancreatic cancer, renal cell carcinoma, cholangiocarcinoma, melanoma, ovarian cancer, and prostate cancer all have a general loss of cilia [8–14]. These studies suggest that loss of cilia may promote cancer development in some tissues. We have also put forth the hypothesis that presence or absence of cilia may regulate targeted drug efficacy (that is, Hedgehog targeted drugs) [3]. In order to better understand the role of cilia in cancer and targeted drug efficacy, it is critical to have a comprehensive analysis of cilia expression in all human cancers.
Very little is known about the role of cilia in breast cancer development. Studies have demonstrated that human breast cancer cell lines have a low frequency of primary cilia [19]. We previously reported that basal epithelial cells in the develo** normal murine mammary gland have a higher frequency of ciliated cells compared to luminal epithelial cells [20]. For our current study, we used basal position of epithelial cells as well as co-staining with antibodies specific for basal cells including α-Smooth Muscle Actin (α-SMA) and p63 to determine the frequency of cilia on basal versus luminal epithelial cells in histologically normal human tissue (Additional file 1: Figure S2). While primary cilia were expressed on both basal and luminal epithelial cells, basal epithelial cells had a higher cilia frequency (Basal: RM, median = 23.6%; HNE, median = 22.3%) than luminal epithelial cells (Luminal: RM, median = 1.1%; HNE, median = 0.5%) (Figure 1A, 1C, and 1D). These results demonstrate that both basal and luminal epithelial cells have cilia-positive cells and that the frequency is higher for basal cells.
Invasive carcinoma (INV) of the breast is classified into three grades (INV: I, 2, and 3) of increasing atypical tissue and representing increased aggressive potential of the tumor. INV-1 and INV-2 are often grouped together as low-grade and INV-3 is considered high-grade. We quantified cilia frequency on INV cancer cells and observed a decrease in the percentage of ciliated cancer cells in INV-1, INV-2, and INV-3 (Figure 1B, 1C, 1D). The median percentage of ciliated cells in INV-1 and INV-2 when combined (median = 0.2%; P <0.001) and INV-3 (median = 0.2%; P <0.001) decreased significantly compared to normal basal epithelial cells (Figure 1C, top and Additional file 1: Table S1A). All (100%) of INV-1, INV-2, and INV-3 patients had an abnormally low (falling below the 75th percentile of normal, Q1) percentage of ciliated epithelial cells compared to basal cells (Figure 1C, bottom and Additional file 1: Table S1B). The median percentage of ciliated cells in INV-3 (median = 0.2%; P <0.01) was also significantly decreased compared to normal luminal epithelial cells (Figure 1D, top and Additional file 1: Table S1A). While INV-1, INV-2 (median = 0.2%) did not have a significant decrease in the percent ciliated cells compared to normal luminal epithelial cells (Figure 1D, top), we observed an increased number of INV-1, INV-2 (low grade), and INV-3 (high grade) patients (43% and 48%, respectively) with an abnormally low (below Q1) percentage of ciliated epithelial cells compared to luminal cells (Figure 1D, bottom and Additional file 1: Table S1C).
To determine if loss of cilia occurs early in breast cancer development we quantified cilia in carcinoma in situ (CIS) lesions, which represent a malignant, non-invasive lesion that is thought to be a precursor to invasive cancer. CIS is also classified into three grades (CIS: I, 2, and 3) of increasing cellular atypia and decreased disease-free survival. CIS-1 and CIS-2 are grouped together as low-grade and CIS-3 is considered high-grade. CIS cancer cells show no evidence of invasion and are therefore contained within a basement membrane border. We quantified cilia frequency on all CIS cells within this basement membrane and compared them to both normal basal and luminal epithelial cells. Loss of primary cilia was observed in CIS-1, CIS-2, and CIS-3. The median percentage of ciliated cells in CIS-1 and CIS-2 when combined (median = 0.8%; P <0.001) and CIS-3 (median = 0.1%; P <0.001) decreased significantly compared to normal basal epithelial cells (Figure 1C, top and Additional file 1: Table S1A). All (100%) of CIS-1, CIS-2, and CIS-3 patients had an abnormally low (below Q1) percentage of ciliated epithelial cells compared to basal cells (Figure 1C, bottom and Additional file 1: Table S1B). While the median percentage of ciliated cells in CIS-1, CIS-2 (median = 0.8%), and CIS-3 (median = 0.1%) was not significantly lower compared to normal luminal epithelial cells (Figure 1D, top), an increased number of CIS-3 patients (50%) had an abnormally low (below Q1) percentage of ciliated epithelial cells compared to luminal cells (Figure 1D, bottom and Additional file 1: Table S1C). Loss of cilia observed on cells associated with CIS lesions suggests that ciliary assembly defects occur early in breast cancer development.
Loss of cilia during breast cancer development is not due to increased proliferation or gross centrosomal defects
Ciliogenesis is a cell cycle regulated event. Primary cilia are found on cells in G0 of the cell cycle. As cells re-enter the cell cycle the cilium is resorbed into the cytoplasm [21]. Therefore, a possible cause of loss of cilia on cancer cells could be decreased percentage of cells in G0 phase of the cell cycle (increased proliferative index). Ki67 is a protein expressed in all phases of the cell cycle except G0[22]. We therefore correlated percentage of ciliated cells to Ki67 positive cells in INV to investigate if the loss of primary cilia in cancer cells is due to a high proliferative index (Figure 1E). A statistical correlation was not observed between low percent cilia and high percent Ki67 in invasive breast cancer samples (linear regression, R2 = 0.005). We found that the majority of INV samples (84% of patients) had a low proliferative index (<50% Ki67-positive cells). A small fraction of INV samples (9% of patients) did indeed have a high proliferative index (>80% Ki67-positive cells) indicating that loss of cilia may be due to increased proliferation in some patients. The percent Ki67-positive cells therefore does not account for the low percentage of cilia (median = 0.2%) observed in the majority of these breast cancer patients.
Centrosome amplification has been observed in breast cancer [23]. While we observed some breast cancer cells with centrosome amplification, we also observed patient samples with cancer cells with no evidence of centrosome amplification. We next observed that cancer cells from patients with no evidence of centrosome amplification have a significant decrease in the percentage of cilia compared to normal cells (Figure 1B). These observations suggest that, in some patients, loss of cilia is due to defects that are separate from mechanisms that cause centrosome amplification.
Cilia-positive cancer cells co-express cytokeratin 5
Cytokeratin 5 (CK5) is a marker of basal epithelial cells and progenitor cells in the normal breast [24]. CK5 was recently shown to also be a marker used in diagnosing breast cancers in the basaloid group and is associated with a poor prognosis [25]. To determine the frequency of primary cilia on CK5-positive (CK5+) versus CK5-negative (CK5-) cells in normal and cancer tissues we co-stained the RM, HNE, and INV patient samples with markers of cilia (acetylated tubulin and γ-tubulin) together with an antibody that recognizes CK5. To verify that the CK5 antibody used was specific, we compared it to another commonly used CK5 monoclonal antibody (Leica Micosystems, Inc.), and found the two CK5 antibodies co-localized in our tissues (Additional file 1: Figure S2).
CK5+ cells were present in all normal breast epithelium (Figure 2A). Primary cilia were found on both CK5- and CK5+ normal epithelial cells. The median percentage of ciliated CK5- basal epithelial cells in RM and HNE tissues (median = 26.0% and 14.9%) was not significantly different compared to CK5+ basal epithelial cells (median = 19.6% and 23.5%) (Figure 2B and Additional file 1: Table S2A).
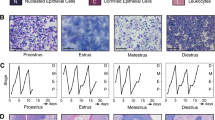
Rare cilia-positive cancer cells are positive for Cytokeratin 5. (A) Images show nuclei (blue), cytokeratin 5 (white), cilia (red), and centrosomes (green) in cells from the following tissue: reduction mammoplasties (RM) and histologically normal epithelium (HNE) adjacent to cancer and invasive cancers of four different cancer subtypes: luminal A and luminal B, Her2-positive, and triple negative. Insets show a magnified ciliated CK5-positive cell. (B) Boxplot represents median percent of cilia expressed on CK5-negative and CK5-positive cells in the following breast tissue types: basal epithelial cells of normal breast reduction mammoplasties (RM Basal, n = 12), luminal epithelial cells of normal breast reduction mammoplasties (RM Luminal, n = 12), basal cells in histologically normal epithelium adjacent to cancer (HNE Basal, n = 15), luminal cells in histologically normal epithelium adjacent to cancer (HNE Basal, n = 15), cancer cells in invasive cancers of the four breast cancer subtypes (Luminal A (LUM A, n = 37), Luminal B (LUM B, n = 10), Her2-positive (Her2+, n = 6), and Triple Negative (TN, n = 12)). Statistical significance (** = P <0.01, *** = P <0.001) was determined by performing logistic regression compared to normal RM.
The overall median percentage of ciliated cells (CK5- and CK5+ combined) in the four breast cancer subtypes was not significantly different compared to one another (median: luminal A = 0.2%, luminal B = 0.1%, Her2 = 0.5%, and Triple Negative = 0.1%; Figure 2A, Additional file 1: Table S2B). While the percentage of cancer cells with cilia is low, we found that 29.1% of these rare ciliated cancer cells were CK5+. We find CK5+ staining on these rare cilia-positive cancer cells in all four breast cancer subtypes including Luminal A, Luminal B, Her2, and Triple Negative (Figure 2B and Additional file 1: Table S2A). CK5+ staining separated out patients with a higher than the median (above 0.5%) percentage of ciliated cancer cells compared to the CK5- ciliated cancer group (Figure 2B). Further investigation is needed to better understand the significance of these rare ciliated, CK5+ cancer cells.
The frequency of ciliated stromal cells decreases as breast cancer develops
The stromal environment that surrounds a cancer can play a causal role in its development and metastasis [26]. Therefore, we investigated cilia changes on stromal cells. Stromal cells were identified based on location and elongated morphology (small round lymphocytes were excluded from this analysis). Primary cilia frequency was calculated on stromal cells surrounding normal breast (RM and HNE) and breast cancers (CIS and INV) (Figure 3A).
Fraction of ciliated stromal cells decrease as breast cancer develops. (A) Image showing stromal cells (str) next to a normal duct (left) or next to a cancer (ca) structure (right). Insets show cilia (acetylated tubulin, red) and their associated centrosomes (γ-tubulin, green) on stromal cells. (B) Boxplot represents percent of ciliated stromal cells in the following breast tissue types: normal breast reduction mammoplasties (RM, n = 12), histologically normal epithelium adjacent to cancer (HNE, n = 15), carcinoma in situ lesions grades 1 and 2 combined (CIS 1&2, n = 23), carcinoma in situ grade 3 (CIS3, n = 16), invasive cancers grades 1 and 2 combined (INV 1&2, n = 40), invasive cancer grade 3 (INV3, n = 25). The bar graph represents the percent of patients that have an abnormally low percentage of ciliated stromal cells (blue bars: Quartile 1 (Q1), less than or equal to the 75th percentile for normal tissue) or an abnormally high percentage of ciliated stromal cells (orange bars: quartile 4 (Q4), greater than or equal to the 25th percentile for normal tissue). Statistical significance (** = P <0.01, *** = P <0.001) was determined by performing logistic regression compared to normal RM.
The frequency of primary cilia on stromal cells associated with CIS-1 and CIS-2 combined (median = 4.7%; P <0.001), CIS-3 (median = 1.3%; P <0.01), INV-1 and INV-2 combined (median = 4.7%; P <0.001), INV-3 (median = 1.7%; P <0.001), decreased compared to stromal cells associated with histologically normal RM (median = 18.4%) (Figure 3B and Additional file 1: Table S3A). The types of stromal cells (that is, fibroblasts, lymphocytes, and so on) were not differentiated for this study. It is possible that the decrease in the overall percent cilia observed in the stroma is due to changes in the stromal cell type distribution during cancer development. These data also raise the possibility that primary cilia are lost on stromal cells associated with preinvasive and invasive breast cancerous cells.
Cilia expressed on breast cancer and cancer-associated stromal cells have abnormal lengths
To determine the potential functionality of cilia that are present on breast cancer cells we measured their lengths. Abnormally short and long cilia have been shown to correlate with abnormal regulation of signal transduction pathways such as Hedgehog signaling [1, 27]. This suggests that abnormal cilia length can be used as an indirect measure of the functional state of cilia.
As cilia frequency was quantified we noted the presence of rare, very long cilia expressed on breast cancer cells (Figure 4A). The median cilia length was not significantly different on cancer cells of CIS-1 & 2 (median = 0.79 μm), CIS-3 (median = 1.27 μm), or INV-1 & 2 (median = 1.10 μm) compared to normal tissue. However, the median cilia length was significantly longer on cancer cells of INV-3 (median = 1.20 μm; P <0.01) compared to cilia length on cells associated with normal tissue (RM: median = 0.76 μm; HNE: median = 0.9 μm) (Figure 4B and 4C, top and Additional file 1: Table S4A). Nevertheless, we did observe an increased number of CIS-1&2, CIS-3, INV-1&2, and INV-3 patients (38%, 75%, 57%, and 77% respectively) with cilia that were abnormally long (above Q4) compared to normal RM and HNE (Figure 4B and 4C, bottom and Additional file 1: Table S4B).
Cilia length abnormalities associated with breast cancer and their stromal cells. (A) Images of cilia representing typical lengths on basal epithelial cell in normal breast reduction mammoplasties, on invasive cancer cells, on stromal cells associated with normal breast reduction mammoplasties, and stromal cells associated with invasive cancers. (B, C) Boxplot represents median length of cilia expressed on cells in the following breast tissue types: basal epithelial cells of normal breast reduction mammoplasties (RM Basal, n = 12), luminal epithelial cells of normal breast reduction mammoplasties (RM Luminal, n = 12), basal cells in histologically normal epithelium adjacent to cancer (HNE Basal, n = 15), luminal cells in histologically normal epithelium adjacent to cancer (HNE Basal, n = 15), cancer cells in carcinoma in situ lesions grades 1 and 2 combined (CIS 1&2, n = 23), cancer cells in carcinoma in situ grade 3 (CIS3, n = 16), cancer cells in invasive cancers grades 1 and 2 combined (INV 1&2, n = 40), cancer cells in invasive cancer grade 3 (INV3, n = 25). (D) Boxplot represents the median length of cilia on stromal cells in the following breast tissue types: normal breast reduction mammoplasties (RM, n = 12), histologically normal epithelium adjacent to cancer (HNE, n = 15), carcinoma in situ lesions grades 1 and 2 combined (CIS 1&2, n = 23), carcinoma in situ grade 3 (CIS3, n = 16), invasive cancers grades 1 and 2 combined (INV 1&2, n = 40), invasive cancer grade 3 (INV3, n = 25). (B-D) The bar graph represents the percent of patients that have an abnormally low percentage of ciliated cells (blue bars: quartile 1 (Q1), less than or equal to the 75th percentile for normal tissue) or an abnormally high percentage of ciliated cells (orange bars: quartile 4 (Q4), greater than or equal to the 25th percentile for normal tissue). (** = P <0.01) was determined by performing logistic regression compared to normal RM.
No statistically significant difference was detected in the median cilia length on stromal cells associated with CIS-1 & 2 combined (median = 0.85 μm), CIS-3 (median = 0.78 μm; P = 0.15), INV-1 & 2 combined (median = 0.95 μm), or INV-3 (median = 0.93 μm) when we compared cilia length on stromal cells associated with normal tissue (RM: median = 1.00 μm; HNE: median = 0.68 μm) (Figure 4D, top and Additional file 1: Table S4D). While median cilia lengths were not different, we observed an increased number of HNE, CIS-1&2, CIS-3, INV-1&2, and INV-3 patients (73%, 74%, 69%, 50%, and 46% respectively) that had cilia with abnormally short (below Q1) cilia compared to normal RM (Figure 4D, bottom and Additional file 1: Table S4E). Taken together these findings may indicate that rare ciliated cancer cells and ciliated cancer-associated stromal cells have dysfunctional cilia.
Loss of primary cilia on breast cancers correlates with decreased expression of genes required for ciliogenesis
To determine if alterations in the expression of ciliary-associated genes plays a role in the loss of cilia we examined expression of genes required for ciliary assembly (ciliogenesis) or ciliary function in human breast cancers. We assembled a list of genes reported in the literature to be associated with ciliogenesis and/or cilia-related genetic disorders (ciliopathies) [2, 28]. This list included 69 genes from the following cilia-associated categories: Intraflagellar Transport (IFT) complex A, IFT complex B, ciliary motor subunits, BBSome complex proteins, and other ciliopathy-associated genes. The human subject microarray used for this analysis was carried out using a previously published dataset [29]. We determined the average expression of each cilia-associated gene and performed ANOVA (Analysis of Variance) between normal mammary tissue (n = 7) and breast cancer samples (n = 18) (Figure 5A). Gene expression was considered significantly different between normal and cancer samples with adjusted P value less than 0.05 and B value greater than 3.0. Among the 69 ciliary-associated genes analyzed, seven genes showed a statistically significant decrease in expression in breast cancers (log fold change: DYNC2H1 = -0.66; IFT46 = -1.07; PKD2 = -1.67; NPHP3 = -1.10; BBS2 = -1.51; BBS4 = -0.78; TTC8 = -1.46) (Figure 5B and Additional file 1: Table S5). These data demonstrate that a subset of cilia-related genes is commonly downregulated in breast cancers.
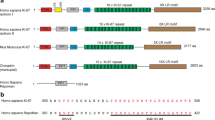
Characterization of human breast cancer samples for expression of ciliogenesis genes. (A) Heat map displays the relative gene expression change seen in basal-like cancer samples compared to normal breast tissue. Each column represents a breast cancer tissue sample (sample name listed on top) and each row represents the relative expression results of a different ciliary gene (gene names are to the right). Overexpression is depicted in red, and underexpression is depicted in green. (B) The boxplots represent distribution of expression values for individual genes comparing breast cancer tissue samples to the normal tissues samples. The ciliary-associated genes shown in boxplots are those that were found to be statistically significant by ANOVA (Analysis of Variance) between normal and basal-like cancer samples with adjusted *P value <0.05. The log odds that the gene is differentially expressed are provided by B-statistics and B >3.0 was used as a cutoff for significantly changing genes. (C) Images show staining of normal breast reduction mammoplaties or breast cancer tissue with secondary antibody alone or with an antibody recognizing DYNC2H1 (red). Inset in normal breast tissue shows that tissue was also co-stained for antibodies to identify cilia (acetylated tubulin, green; γ-tubulin, white) and analyzed for co-localization in the same cell. The arrow points to a cilium with a pool of DYNC2H1 at the base.
Next, we examined if this subset of cilia-related genes is downregulated in a larger cohort of invasive ductal breast carcinomas. We analyzed the full set of cilia-related genes in the publically available TCGA breast cancer dataset using Oncomine [18]. The gene expression data were log transformed, median centered per array, and the standard deviation was normalized to one per array for each of the cilia-related genes. We compared the median difference in gene expression between normal mammary tissue (n = 61) and invasive ductal breast carcinoma (n = 389). Of the seven identified in the Richardson dataset above, six were also significantly downregulated in the TCGA dataset (log fold change: DYNC2H1 = -2.35; IFT46 = -1.17; PKD2 = -2.62; NPHP3 = -2.40; BBS2 = -1.91; TTC8 = -1.31) (Additional file 1: Table S5). Gene expression was considered significantly different between normal and cancer samples if the P value was less than 0.05. BBS4 was downregulated (log fold change -1.02) in the TCGA dataset but this was not statistically significant (P = 0.28). Nine additional cilia-related genes were significantly downregulated when analyzed in the TCGA dataset; however, these were not significantly downregulated in the Richardson dataset (B value was less than 3). Also, we did not identify genes in either dataset that showed a statistically significant increase in expression in breast cancers compared to normal tissues.
DYNC2H1 is a cytoplasmic dynein involved in intraflagellar transport in the cilium and therefore is required for ciliogenesis. To determine if protein expression of the cilia-related gene DYNC2H1 is decreased in breast cancers we performed immunofluorescent staining of three specimens taken from normal breast reduction mammoplasties (RM) as well as five patients with invasive breast carcinoma. Note that no other antibodies were identified to the other six significantly downregulated cilia genes that worked for staining of paraffin embedded tissue. Staining of normal breast tissue demonstrated that the basal and luminal cells both have highly positive staining for DYNC2H1 as compared to secondary only control. Cancers cells have decreased positive signal in all five of the breast cancers analyzed. All five of these breast cancers were also stained for primary cilia and showed very low percentages of primary cilia. These data indicate that DYNC2H1 protein expression is decreased in breast cancers. Loss of expression of DYNC2H1 has been shown to have abnormal, shortened cilia due to the lack of transport of proteins from the tip of the cilium back to the cell body (retrograde IFT) [30, 31]. Decreased expression of DYNC2H1 as well as the other ciliary-related genes may turn off ciliogenesis and explain the loss of cilia seen in breast cancers.
Discussion
We hypothesize that primary cilia loss is an early event that promotes breast cancer development. We have knocked out genes required for ciliogenesis (IFT88 and Ki67) and found no evidence of tumor promotion in the mammary glands of these mice [20]. Therefore, loss of cilia does not appear to be sufficient to initiate tumorigenesis in the mammary gland. We hypothesize the loss of cilia cooperates/synergizes with other oncogenic events to promote tumorigenesis. Indeed, loss of cilia has been shown to cooperate with constitutive activation of the Hh pathway (overexpression of constitutively active Gli transcription factor) to promote tumorigenesis in mouse models of basal cell carcinoma and medulloblastoma [5–7]. Overexpression of mutant forms of Gli transcription factors has not been observed in breast cancer patients. However, it is possible that loss of cilia predisposes cancer cells to non-canonical activation of the Hh pathway. Specifically, cilia are required for processing of Gli3 into its repressor form. In normal cells, the Gli3-repressor sits on Hh-target genes and blocks transcriptional activation. Loss of cilia is therefore equivalent to loss of the Gli3-repressor. We hypothesize that when cilia are lost in cancer cells, the Hh-target genes are vulnerable to activation by oncogenic transcription factors. Myc has recently been shown to upregulate transcription of Hh-target genes [32]. Our working model is that loss of cilia, in the context of overexpression or increased activation of oncogenic transcription factors such as Myc, will result in increased Hh-target gene expression and thereby promote tumorigenesis. Loss of cilia has also been reported to abrogate other oncogenic signaling pathways including increasing Wnt signaling [33, 34]. Cross-reactivity with the Wnt pathway may be another mechanism by which cilia promote tumorigenesis.
A decrease in primary cilia expression was found in pre-invasive breast carcinoma in situ (CIS). Not all patients with CIS will progress to invasive breast cancer. However, there is molecular evidence that CIS is a precursor to invasive disease and diagnosis with CIS represents an increased risk of develo** invasive breast cancer [35]. Understanding of the molecular mechanisms that underlie these early stages of tumor development in breast cancer is critical so that we can design targeted therapies for early intervention. Identification of ciliary dysfunction in these precursor lesions may indicate that cilia are involved in early stages of breast cancer promotion and therefore provide a new target for therapy.
We also investigated the cause of loss of primary cilia in breast cancers. We demonstrated that increased proliferation and centrosome amplification did not account for the loss of cilia seen in breast cancer. We also observe that luminal epithelial cells have low expression of cilia in normal breast tissue. If luminal cells are the tumor-initiating cell for some cancer types, they may therefore have this first ‘hit’ already in place, predisposing them to oncogenic events that cooperate with absence of cilia. We next examined expression of genes known to be involved in the formation (ciliogenesis) and function of the primary cilium. Ciliogenesis is a highly regulated process [36]. Hundreds of proteins that localize to the primary cilium have been identified by proteomic analysis [37–45]. Many of these proteins are involved in ciliogenesis. Other proteins localized to the cilium are involved in the sensory or signaling functions of the primary cilium. Transport machinery that is critical for both ciliogenesis as well as ciliary sensory function is mediated by a process known as intraflagellar transport (IFT) [46, 47]. The Kinesin-2 motor complex transports the IFT complex as well as other protein ‘cargo’ for anterograde movement of proteins to the tip of the cilium (plus end of microtubules). The cytoplasmic Dynein 2 motor complex transports the IFT complex plus ‘cargo’ for retrograde movement from the tip of the cilium towards the cell body (minus end of microtubules). The IFT complex is made up of several proteins and mutations in IFT genes cause loss of ciliary assembly and consequently result in loss of sensory functions [48]. Another complex of proteins (BBSome complex) is also required for ciliogenesis and for targeted delivery of proteins into the cilium [49]. Mutations in IFT, BBSome, and other ciliary genes that are required for ciliogenesis and ciliary function are now known to be causal for a large number of genetic disorders classified as ciliopathies. Ciliopathies include Joubert syndrome (JBTS), polycystic kidney disease (PKD), Bardet-Biedl syndrome (BBS), and nephronophthisis (NPHP) [50]. We assembled a list of 69 ciliary-associated genes from the following categories: Intraflagellar Transport (IFT) complex A, IFT complex B, ciliary motor subunits, BBSome complex proteins, and other ciliopathy-associated genes. We observed decreased expression of genes from each of these categories (downregulation of DYNC2H1, IFT46, PKD2, NPHP3, BBS2, BBS4, and TTC8) in two separate microarray datasets. We further validated downregulation of DYNC2H1 protein expression in breast cancers. Decreased expression of these cilia-related genes is known to abrogate ciliogenesis and/or ciliary function and therefore may explain the loss of cilia seen in breast cancers. DYNC2H1 is a cytoplasmic dynein required for intraflagellar transport. Indeed, loss of DYNC2H1 expression alone is known to be sufficient to abrogate ciliogenesis [30, 31]. Further work is needed to understand if the decreased expression of these cilia-related genes is due to an altered transcriptional program or genomic instability.
While the majority of breast cancer cells were devoid of primary cilia, we observed rare cilia-expressing cancer cells. We measured the length of these ciliary axonemes and found that many of the remaining ciliated cells had ciliary axonemes that were abnormally long (longer than ciliary axonemes found on normal epithelial cells). Increases in ciliary length are known to be triggered by environmental or molecular/genetic events. For example, cilia found on renal epithelial cells have increased length in response to renal injury [51]. Also, changes in the actin cytoskeleton result in increased ciliary length [52]. Finally, mutations in MKS3 that are associated with the human ciliopathy Meckel-Gruber syndrome have many of the phenotypic hallmarks of ciliopathies (renal cysts, central nervous system defects, and polydactyly) and the associated cilia are longer than normal [53]. This increased ciliary length associated with a ciliopathy suggests that increases in the length of the ciliary axoneme can abrogate ciliary function.
The rare ciliated cancer cells found in invasive breast cancer samples frequently co-expressed cytokeratin 5 (CK5). CK5 is expressed in progenitor cells that can give rise to luminal and myoepithelial lineages [54]. In basal-like breast tumors, CK5 is a marker of poor prognosis [55]. In addition, CK5+ breast cancer cells have been shown to be more resistant to chemotherapy than CK5- cancer cells [24, 56, 57]. Detection of cilia on the CK5+ breast cancer cells may provide a new molecular target for therapies to eradicate these chemo-resistant cells in patients.
We observed a decrease in the fraction of ciliated stromal cells associated with pre-invasive and invasive breast cancer. The remaining cilia found on stromal cells were shorter than cilia found on normal stromal cells. Absence of an elongated ciliary axoneme has been associated with ciliary dysfunction in a number of animal models and human diseases [2]. Together these finding suggest that there is ciliary dysfunction on stromal cells during breast cancer development. There is growing evidence that the stromal environment surrounding cancer cells is an important regulator of breast cancer development. Boyd et al. have demonstrated that mammographically dense breasts have an increased risk in develo** breast cancer [58]. This increased breast density is attributed to increased cellular and extracellular matrix components in the stroma. The cellular changes in cancer-associated stroma include changes in the number and activation of carcinoma-associated fibroblasts [59, 60]. The activation of cancer-associated fibroblasts is thought to alter the environment by changing the composition of growth factors and ECM proteins and thereby plays a role in breast cancer development. Our observation of ciliary dysfunction in breast cancer-associated stroma prompts the need for future studies to examine the role of cilia on cancer-associated fibroblasts to determine if loss of stromal cilia contributes to breast cancer development.
Our findings have shown that breast cancer cells have a very low frequency of primary cilia. An important question for therapeutics is whether this loss of primary cilia is a driver of breast cancer. Loss of cilia has been associated with increased Hedgehog signaling in mouse models of cancer [5, 6]. Hedgehog-targeted drugs are being investigated both in clinical trials and in laboratories. Most of these drugs inhibit Smoothened (Smo), a positive regulator of the pathway that requires primary cilia to be functional [3]. Since primary cilia are lost in breast tumors, we hypothesize that breast cancer patients would not be good candidates for Smo-inhibitor drugs. However, treatment of patients with a cilia-independent Hedgehog pathway inhibitor may be efficacious [3].
Conclusions
In the present study, we determined that ciliogenesis is decreased in low- and high-grade invasive breast cancers. Our studies also revealed for the first time that primary cilia expression was lost on pre-invasive breast cancer lesions demonstrating that this is an early event in cancer development. Furthermore, cilia expression and function were decreased on cancer-associated stromal cells. Finally, we correlated loss of cilia with downregulation of genes required for ciliogenesis and/or ciliary function in human breast cancers. Future studies are needed to test this hypothesis and determine if ciliary dysfunction plays a causal role in breast cancer development.
Abbreviations
- α-SMA:
-
α-smooth muscle actin
- γ-tub:
-
γ-tubulin
- Ac-Tub:
-
Acetylated tubulin
- ANOVA:
-
Analysis of variance
- BBS:
-
Bardet-Biedl syndrome
- ca:
-
Cancer
- CIS:
-
Carcinoma in situ
- CK5:
-
Cytokeratin 5
- H&E:
-
Hematoxylin and Eosin
- HNE:
-
Histologically normal epithelium adjacent to cancer
- IFT:
-
Intraflagellar transport
- INV:
-
Invasive cancer
- LUM A:
-
Luminal A subtype
- LUM B:
-
Luminal B subtype
- Q1:
-
Quartile 1
- Q4:
-
Quartile 4
- RM:
-
Reduction mammoplasties
- str:
-
Stroma
- TN:
-
Triple negative subtype.
References
Goetz SC, Anderson KV: The primary cilium: a signalling centre during vertebrate development. Nat Rev Genet. 2010, 11: 331-344. 10.1038/nrg2774.
Hildebrandt F, Benzing T, Katsanis N: Ciliopathies. N Engl J Med. 2011, 364: 1533-1543. 10.1056/NEJMra1010172.
Hassounah NB, Bunch TA, McDermott KM: Molecular pathways: the role of primary cilia in cancer progression and therapeutics with a focus on Hedgehog signaling. Clin Cancer Res. 2012, 18: 2429-2435. 10.1158/1078-0432.CCR-11-0755.
Basten SG, Giles RH: Functional aspects of primary cilia in signaling, cell cycle and tumorigenesis. Cilia. 2013, 2: 6-10.1186/2046-2530-2-6.
Wong SY, Seol AD, So PL, Ermilov AN, Bichakjian CK, Epstein EH, Dlugosz AA, Reiter JF: Primary cilia can both mediate and suppress Hedgehog pathway-dependent tumorigenesis. Nat Med. 2009, 15: 1055-1061. 10.1038/nm.2011.
Han YG, Kim HJ, Dlugosz AA, Ellison DW, Gilbertson RJ, Alvarez-Buylla A: Dual and opposing roles of primary cilia in medulloblastoma development. Nat Med. 2009, 15: 1062-1065. 10.1038/nm.2020.
Barakat MT, Humke EW, Scott MP: Kif3a is necessary for initiation and maintenance of medulloblastoma. Carcinogenesis. 2013, 34: 1382-1392. 10.1093/carcin/bgt041.
Schraml P, Frew IJ, Thoma CR, Boysen G, Struckmann K, Krek W, Moch H: Sporadic clear cell renal cell carcinoma but not the papillary type is characterized by severely reduced frequency of primary cilia. Mod Pathol. 2009, 22: 31-36. 10.1038/modpathol.2008.132.
Kim J, Dabiri S, Seeley ES: Primary cilium depletion typifies cutaneous melanoma in situ and malignant melanoma. PLoS ONE. 2011, 6: e27410-10.1371/journal.pone.0027410.
Seeley ES, Carriere C, Goetze T, Longnecker DS, Korc M: Pancreatic cancer and precursor pancreatic intraepithelial neoplasia lesions are devoid of primary cilia. Cancer Res. 2009, 69: 422-430. 10.1158/0008-5472.CAN-08-1290.
Gradilone SA, Radtke BN, Bogert PS, Huang BQ, Gajdos GB, LaRusso NF: HDAC6 inhibition restores ciliary expression and decreases tumor growth. Cancer Res. 2013, 73: 2259-2270. 10.1158/0008-5472.CAN-12-2938.
Egeberg DL, Lethan M, Manguso R, Schneider L, Awan A, Jorgensen TS, Byskov AG, Pedersen LB, Christensen ST: Primary cilia and aberrant cell signaling in epithelial ovarian cancer. Cilia. 2013, 1: 15-
Hassounah NB, Nagle R, Saboda K, Roe DJ, Dalkin BL, McDermott KM: Primary cilia are lost in preinvasive and invasive prostate cancer. PLoS ONE. 2013, 8: e68521-10.1371/journal.pone.0068521.
Basten SG, Willekers S, Vermaat JS, Slaats GG, Voest EE, van Diest PJ, Giles RH: Reduced cilia frequencies in human renal cell carcinomas versus neighboring parenchymal tissue. Cilia. 2013, 2: 2-10.1186/2046-2530-2-2.
Yuan K, Frolova N, **e Y, Wang D, Cook L, Kwon YJ, Steg AD, Serra R, Frost AR: Primary cilia are decreased in breast cancer: analysis of a collection of human breast cancer cell lines and tissues. J Histochem Cytochem. 2010, 58: 857-870. 10.1369/jhc.2010.955856.
Nobutani K, Shimono Y, Yoshida M, Mizutani K, Minami A, Kono S, Mukohara T, Yamasaki T, Itoh T, Takao S, Minami H, Azuma T, Takai Y: Absence of primary cilia in cell cycle-arrested human breast cancer cells. Genes Cells. 2014, 19: 141-152. 10.1111/gtc.12122.
Wettenhall JM, Smyth GK: LimmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics. 2004, 20: 3705-3706. 10.1093/bioinformatics/bth449.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, Barrette T, Pandey A, Chinnaiyan AM: ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004, 6: 1-6. 10.1016/S1476-5586(04)80047-2.
Visvader JE: Kee** abreast of the mammary epithelial hierarchy and breast tumorigenesis. Genes Dev. 2009, 23: 2563-2577. 10.1101/gad.1849509.
McDermott KM, Liu BY, Tlsty TD, Pazour GJ: Primary cilia regulate branching morphogenesis during mammary gland development. Curr Biol. 2010, 20: 731-737. 10.1016/j.cub.2010.02.048.
Pan J, Snell W: The primary cilium: keeper of the key to cell division. Cell. 2007, 129: 1255-1257. 10.1016/j.cell.2007.06.018.
Scholzen T, Gerdes J: The Ki-67 protein: from the known and the unknown. J Cell Physiol. 2000, 182: 311-322. 10.1002/(SICI)1097-4652(200003)182:3<311::AID-JCP1>3.0.CO;2-9.
Lingle WL, Barrett SL, Negron VC, D’Assoro AB, Boeneman K, Liu W, Whitehead CM, Reynolds C, Salisbury JL: Centrosome amplification drives chromosomal instability in breast tumor development. Proc Natl Acad Sci U S A. 2002, 99: 1978-1983. 10.1073/pnas.032479999.
Axlund SD, Yoo BH, Rosen RB, Schaack J, Kabos P, Labarbera DV, Sartorius CA: Progesterone-inducible cytokeratin 5-positive cells in luminal breast cancer exhibit progenitor properties. Horm Cancer. 2013, 4: 36-49. 10.1007/s12672-012-0127-5.
Inanc M, Ozkan M, Karaca H, Berk V, Bozkurt O, Duran AO, Ozaslan E, Akgun H, Tekelioglu F, Elmali F: Cytokeratin 5/6, c-Met expressions, and PTEN loss prognostic indicators in triple-negative breast cancer. Med Oncol. 2014, 31: 801-
Hanahan D, Weinberg RA: Hallmarks of cancer: the next generation. Cell. 2011, 144: 646-674. 10.1016/j.cell.2011.02.013.
Broekhuis JR, Leong WY, Jansen G: Regulation of cilium length and intraflagellar transport. Int Rev Cell Mol Biol. 2013, 303: 101-138.
Bhogaraju S, Engel BD, Lorentzen E: Intraflagellar transport complex structure and cargo interactions. Cilia. 2013, 2: 10-10.1186/2046-2530-2-10.
Richardson AL, Wang ZC, De Nicolo A, Lu X, Brown M, Miron A, Liao X, Iglehart JD, Livingston DM, Ganesan S: X chromosomal abnormalities in basal-like human breast cancer. Cancer Cell. 2006, 9: 121-132. 10.1016/j.ccr.2006.01.013.
Schmidts M, Arts HH, Bongers EM, Yap Z, Oud MM, Antony D, Duijkers L, Emes RD, Stalker J, Yntema JB, Plagnol V, Hoischen A, Gilissen C, Forsythe E, Lausch E, Veltman JA, Roeleveld N, Superti-Furga A, Kutkowska-Kazmierczak A, Kamsteeg EJ, Elcioglu N, van Maarle MC, Graul-Neumann LM, Devriendt K, Smithson SF, Wellesley D, Verbeek NE, Hennekam RC, Kayserili H, Scambler PJ, et al: Exome sequencing identifies DYNC2H1 mutations as a common cause of asphyxiating thoracic dystrophy (Jeune syndrome) without major polydactyly, renal or retinal involvement. J Med Genet. 2013, 50: 309-323. 10.1136/jmedgenet-2012-101284.
Dagoneau N, Goulet M, Genevieve D, Sznajer Y, Martinovic J, Smithson S, Huber C, Baujat G, Flori E, Tecco L, Cavalcanti D, Delezoide AL, Serre V, Le Merrer M, Munnich A, Cormier-Daire V: DYNC2H1 mutations cause asphyxiating thoracic dystrophy and short rib-polydactyly syndrome, type III. Am J Hum Genet. 2009, 84: 706-711. 10.1016/j.ajhg.2009.04.016.
Yoon JW, Gallant M, Lamm ML, Iannaccone S, Vieux KF, Proytcheva M, Hyjek E, Iannaccone P, Walterhouse D: Noncanonical regulation of the Hedgehog mediator GLI1 by c-MYC in Burkitt lymphoma. Mol Cancer Res. 2013, 11: 604-615. 10.1158/1541-7786.MCR-12-0441.
Corbit KC, Shyer AE, Dowdle WE, Gaulden J, Singla V, Chen MH, Chuang PT, Reiter JF: Kif3a constrains beta-catenin-dependent Wnt signalling through dual ciliary and non-ciliary mechanisms. Nat Cell Biol. 2008, 10: 70-76. 10.1038/ncb1670.
Lancaster MA, Schroth J, Gleeson JG: Subcellular spatial regulation of canonical Wnt signalling at the primary cilium. Nat Cell Biol. 2011, 13: 700-707.
Benson JR, Wishart GC: Predictors of recurrence for ductal carcinoma in situ after breast-conserving surgery. Lancet Oncol. 2013, 14: e348-e357. 10.1016/S1470-2045(13)70135-9.
Pazour GJ, Rosenbaum JL: Intraflagellar transport and cilia-dependent diseases. Trends Cell Biol. 2002, 12: 551-555. 10.1016/S0962-8924(02)02410-8.
Ostrowski LE, Blackburn K, Radde KM, Moyer MB, Schlatzer DM, Moseley A, Boucher RC: A proteomic analysis of human cilia: identification of novel components. Mol Cell Proteomics. 2002, 1: 451-465. 10.1074/mcp.M200037-MCP200.
Li JB, Gerdes JM, Haycraft CJ, Fan Y, Teslovich TM, May-Simera H, Li H, Blacque OE, Li L, Leitch CC, Lewis RA, Green JS, Parfrey PS, Leroux MR, Davidson WS, Beales PL, Guay-Woodford LM, Yoder BK, Stormo GD, Katsanis N, Dutcher SK: Comparative genomics identifies a flagellar and basal body proteome that includes the BBS5 human disease gene. Cell. 2004, 117: 541-552. 10.1016/S0092-8674(04)00450-7.
Smith JC, Northey JG, Garg J, Pearlman RE, Siu KW: Robust method for proteome analysis by MS/MS using an entire translated genome: demonstration on the ciliome of Tetrahymena thermophila. J Proteome Res. 2005, 4: 909-919. 10.1021/pr050013h.
Pazour GJ, Agrin N, Leszyk J, Witman GB: Proteomic analysis of a eukaryotic cilium. J Cell Biol. 2005, 170: 103-113. 10.1083/jcb.200504008.
Gherman A, Davis EE, Katsanis N: The ciliary proteome database: an integrated community resource for the genetic and functional dissection of cilia. Nat Genet. 2006, 38: 961-962. 10.1038/ng0906-961.
Inglis PN, Boroevich KA, Leroux MR: Piecing together a ciliome. Trends Genet. 2006, 22: 491-500. 10.1016/j.tig.2006.07.006.
Wagner V, Gessner G, Heiland I, Kaminski M, Hawat S, Scheffler K, Mittag M: Analysis of the phosphoproteome of Chlamydomonas reinhardtii provides new insights into various cellular pathways. Eukaryot Cell. 2006, 5: 457-468. 10.1128/EC.5.3.457-468.2006.
Liu Q, Tan G, Levenkova N, Li T, Pugh EN, Rux JJ, Speicher DW, Pierce EA: The proteome of the mouse photoreceptor sensory cilium complex. Mol Cell Proteomics. 2007, 6: 1299-1317. 10.1074/mcp.M700054-MCP200.
Mayer U, Kuller A, Daiber PC, Neudorf I, Warnken U, Schnolzer M, Frings S, Mohrlen F: The proteome of rat olfactory sensory cilia. Proteomics. 2009, 9: 322-334. 10.1002/pmic.200800149.
Rosenbaum JL, Witman GB: Intraflagellar transport. Nat Rev Mol Cell Biol. 2002, 3: 813-825. 10.1038/nrm952.
Scholey JM: Intraflagellar transport. Annu Rev Cell Dev Biol. 2003, 19: 423-443. 10.1146/annurev.cellbio.19.111401.091318.
Pazour GJ: Intraflagellar transport and cilia-dependent renal disease: the ciliary hypothesis of polycystic kidney disease. J Am Soc Nephrol. 2004, 15: 2528-2536. 10.1097/01.ASN.0000141055.57643.E0.
Nachury MV, Seeley ES, ** H: Trafficking to the ciliary membrane: how to get across the periciliary diffusion barrier?. Annu Rev Cell Dev Biol. 2010, 26: 59-87. 10.1146/annurev.cellbio.042308.113337.
Ware SM, Aygun MG, Hildebrandt F: Spectrum of clinical diseases caused by disorders of primary cilia. Proc Am Thorac Soc. 2011, 8: 444-450. 10.1513/pats.201103-025SD.
Verghese E, Weidenfeld R, Bertram JF, Ricardo SD, Deane JA: Renal cilia display length alterations following tubular injury and are present early in epithelial repair. Nephrol Dial Transplant. 2008, 23: 834-841.
Kim J, Lee JE, Heynen-Genel S, Suyama E, Ono K, Lee K, Ideker T, Aza-Blanc P, Gleeson JG: Functional genomic screen for modulators of ciliogenesis and cilium length. Nature. 2010, 464: 1048-1051. 10.1038/nature08895.
Tammachote R, Hommerding CJ, Sinders RM, Miller CA, Czarnecki PG, Leightner AC, Salisbury JL, Ward CJ, Torres VE, Gattone VH, Harris PC: Ciliary and centrosomal defects associated with mutation and depletion of the Meckel syndrome genes MKS1 and MKS3. Hum Mol Genet. 2009, 18: 3311-3323. 10.1093/hmg/ddp272.
Rios AC, Fu NY, Lindeman GJ, Visvader JE: In situ identification of bipotent stem cells in the mammary gland. Nature. 2014, 506: 322-327. 10.1038/nature12948.
Perou CM, Sorlie T, Eisen MB, van de Rijn M, Jeffrey SS, Rees CA, Pollack JR, Ross DT, Johnsen H, Akslen LA, Fluge O, Pergamenschikov A, Williams C, Zhu SX, Lonning PE, Borresen-Dale AL, Brown PO, Botstein D: Molecular portraits of human breast tumours. Nature. 2000, 406: 747-752. 10.1038/35021093.
Kabos P, Haughian JM, Wang X, Dye WW, Finlayson C, Elias A, Horwitz KB, Sartorius CA: Cytokeratin 5 positive cells represent a steroid receptor negative and therapy resistant subpopulation in luminal breast cancers. Breast Cancer Res Treat. 2011, 128: 45-55. 10.1007/s10549-010-1078-6.
Lim E, Vaillant F, Wu D, Forrest NC, Pal B, Hart AH, Asselin-Labat ML, Gyorki DE, Ward T, Partanen A, Feleppa F, Huschtscha LI, Thorne HK, kConFab , Fox SB, Yan M, French JD, Brown MA, Smyth GK, Visvader JE, Lindeman GJ: Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat Med. 2009, 15: 907-913. 10.1038/nm.2000.
Boyd NF, Dite GS, Stone J, Gunasekara A, English DR, McCredie MR, Giles GG, Tritchler D, Chiarelli A, Yaffe MJ, Hopper JL: Heritability of mammographic density, a risk factor for breast cancer. N Engl J Med. 2002, 347: 886-894. 10.1056/NEJMoa013390.
Pavelka N, Rancati G, Li R: Dr Jekyll and Mr Hyde: role of aneuploidy in cellular adaptation and cancer. Curr Opin Cell Biol. 2011, 22: 809-815.
Dumont N, Liu B, Defilippis RA, Chang H, Rabban JT, Karnezis AN, Tjoe JA, Marx J, Parvin B, Tlsty TD: Breast fibroblasts modulate early dissemination, tumorigenesis, and metastasis through alteration of extracellular matrix characteristics. Neoplasia. 2013, 15: 249-262.
Acknowledgements
The authors would like to acknowledge Tom Bunch, and Martha Nunez for feedback and scientific discussions. We would like to thank the TACMASS members for tissue processing and H&E staining. We would also like to thank Lynn Rychter and Robert Hers off in the Department of Pathology for digitally scanning tissue slides. Finally, we would also like to acknowledge Vern Harts, Denise Roe, and Kathy Lynn Abode for assistance with statistical analysis. Funding was provided by the Cancer Center Support Grant, National Cancer Institute, NIH, P30CA023074 (K. M. McDermott); Cancer Biology Training Grant, National Cancer Institute, NIH, T32CA009213 (N. B. Mascoutah); R00, National Institute of Child Health and Human Development, NIH, R00HD056965 (K. M. McDermott). Funding sources did not play a role in the design, in the collection, analysis, interpretation of data, writing of the manuscript, or decision to submit for publication.
Author information
Authors and Affiliations
Corresponding author
Additional information
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
IM and NBH carried out the staining and imaging of tissues, and drafted the methods and figures. LL and RN analyzed all normal and cancer pathology. RP and FWL analyzed microarray dataset and performed statistical analysis. KW assembled the breast cancer cohort. KMM conceived and designed the study and participated in writing of the manuscript. All authors read and approved the final manuscript.
Electronic supplementary material
Authors’ original submitted files for images
Below are the links to the authors’ original submitted files for images.
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Menzl, I., Lebeau, L., Pandey, R. et al. Loss of primary cilia occurs early in breast cancer development. Cilia 3, 7 (2014). https://doi.org/10.1186/2046-2530-3-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/2046-2530-3-7