Abstract
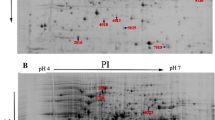
The quorum sensing (QS) system SprIR of the psychrotrophic strain Serratia proteamaculans 94 was investigated. A mutant was constructed with the inactivated sprR gene encoding the regulatory receptor protein SprR. Inactivation of this gene was shown to affect the composition of fatty acids synthesized by S. proteamaculans 94 and did not affect the synthesis of N-acyl-L-homoserine-lactones (AHL); the activities of extracellular proteases, chitinases, and hemolysins; the swimming motility of cells; and the suppression of mycelium growth of fungal plant pathogens by volatile compounds emitted by this strain. Inactivation of the sprI gene (but not the sprR gene) reduced the biofilm formation, which increased when exogenous AHL was added to the culture. The comparative proteomic analysis of cell of the parent strain and mutant strains with inactivated sprI and sprR genes showed that the expression of 30 proteins in S. proteamaculans 94 is affected by the SprIR quorum sensing system.



Similar content being viewed by others
REFERENCES
Fuqua, C., Winans, S.C., and Greenberg, E.P., Census and consensus in bacterial ecosystems: the LuxR-LuxI family of quorum-sensing transcriptional regulators, Annu. Rev. Microbiol., 1996, vol. 50, pp. 727–751. https://doi.org/10.1146/annurev.micro.50.1.727
Waters, C.M. and Bassler, B.L., Quorum sensing: cell-to-cell communication in bacteria, Annu. Rev. Cell. Dev. Biol., 2005, vol. 21, pp. 319–346. https://doi.org/10.1146/annurev.cellbio.21.012704.131001
Khmel, I.A. and Metlitskaya, A.Z., Quorum sensing regulation of gene expression: a promising target for drugs against bacterial pathogenicity, Mol. Biol. (Moscow), 2006, vol. 40, pp. 195–210.
Veselova, M.A. Quorum sensing regulation in Pseudomonas, Russ. J. Genet., 2010, vol. 46, no. 2, pp. 129–137. https://doi.org/10.1134/S1022795410020018
Zaitseva, Y.V., Popova, A.A., and Khmel, I.A., Quorum sensing regulation in bacteria of the family Enterobacteriaceae, Russ. J. Genet., 2014, vol. 50, no. 4, pp. 323–340. https://doi.org/10.1134/S1022795414030120
Papenford, K. and Bassler, B., Quorum-sensing signal-response systems in gram-negative bacteria, Nat. Rev. Microbiol., 2016, vol. 14, pp. 576–588. https://doi.org/10.1038/nrmicro.2016.89
Whiteley, M., Diggle, S.P., and Greenberg, E.P., Bacterial quorum sensing: the progress and promise of an emerging research area, Nature, 2017, vol. 551, no. 7680, pp. 313–320. https://doi.org/10.1038/natur.24624
Beck von Bodman, S.B., Majerczak, D.R., and Coplin, D.L., A negative regulator mediates quorum-sensing control of exopolysaccharide production in Pantoea stewartia subsp. stewartii, Proc. Natl. Acad. Sci. U.S.A., 1998, vol. 95, pp. 7687–7692. https://doi.org/10.1073/pnas.95.13.7687
Horng, Y.T., Deng, S.C., Daykin, M., et al., The LuxR family protein SpnR functions as a negative regulator of N-acylhomoserine lactone-dependent quorum sensing in Serratia marcescens, Mol. Microbiol., 2002, vol. 45, pp. 1655–1671. https://doi.org/10.1046/j.1365-2958.2002.03117.x
Minogue, T.D., Wehland-von Trebra, M., Bernhard, F., and von Bodman, S.B., The autoregulatory role of EsaR, a quorum-sensing regulator in Pantoea stewartia ssp. stewartii: evidence for a repressor function, Mol. Microbiol., 2002, vol. 44, no. 6, pp. 1625–1635. https://doi.org/10.1046/j.1365-2958.2002.02987.x
Christensen, A.B., Riedel, K., Eberl, L., et al., Quorum-sensing-directed protein expression in Serratia proteamaculans B5a, Microbiology, 2003, vol. 149, pp. 471–483. https://doi.org/10.1099/mic.0.25575-0
Van Houdt, R., Givskov, M., and Michiels, C.W., Quorum sensing in Serratia, FEMS Microbiol. Rev., 2007, vol. 31, pp. 407–424. https://doi.org/10.1111/j.1574-6976.2007.00071.x
Demidyuk, I.V., Kalashnikov, A.E., Gromova, T.Y., et al., Cloning, sequencing, expression, and characterization of protealysin, a novel neutral proteinase from Serratia proteamaculans representing a new group of thermolysin-like proteases with short N-terminal region of precursor, Protein Expression Purif., 2006, vol. 47, pp. 551–561. https://doi.org/10.1016/j.pep.2005.12.005
Zaitseva, Y.V., Koksharova, O.A., Lipasova, V.A., et al., SprI/SprR quorum sensing system of Serratia proteamaculans 94, BioMed. Res. Int., 2019, vol. 2019, article ID 3865780. https://doi.org/10.1155/2019/3865780
Zaitseva, Y.V., Lipasova, V.A., Plyuta, V.A., et al., Effect of inactivation of luxS gene on the properties of Serratia proteamaculans 94 strain, Folia Microbiol. (Praha), 2019, vol. 64, pp. 265–272. https://doi.org/10.1007/s12223-018-0657-5
Miller, J.H., Experiments in Molecular Genetics, Cold Spring Harbor, N.Y.: Cold Spring Harbor Laboratory, 1972.
McClean, K.H., Winson, M.K., Fish, L., et al., Quorum sensing in Chromobacterium violaceum: exploitation of violacein production and inhibition for the detection of N-acylhomoserine lactones, Microbiology, 1997, vol. 143, pp. 3703–3711. https://doi.org/10.1099/00221287-143-12-3703
Shaw, P.D., **, G., Daly, S.L., et al., Detecting and characterizing N-acyl-homoserine lactone signal molecules by thin-layer chromatography, Proc. Natl. Acad. Sci. U.S.A., 1997, vol. 94, pp. 6036–6041. https://doi.org/10.1073/pnas.94.12.6036
Current Protocols in Molecular Biology, Ausubel, F.M., Brendt, R., and Kingston, R.E., Eds., New York: Wiley, 1994.
Hoang, T.T., Karkhoff-Schweizer, R.R., Kutchma, A.J., and Schweizer, H.P., A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants, Gene, 1998, vol. 212, pp. 77–86. https://doi.org/10.1016/s0378-1119(98)00130-9
Dennis, J.J. and Zylstra, G.J., Plasposons: modular self-cloning minitransposon derivatives for rapid genetic analysis of gram-negative bacterial genomes, Appl. Environ. Microbiol., 1998, vol. 64, no. 7, pp. 2710—2715. https://doi.org/10.1128/AEM.64.7.2710-2715.1998
Popova, A.A., Koksharova, O.A., Lipasova, V.A., et al., Inhibitory and toxic effects of volatiles emitted by strains of Pseudomonas and Serratia on growth and survival of selected microorganisms, Caenorhabditis elegans and Drosophila melanogaster, BioMed Res. Int., 2014, vol. 2014, article ID 125704. https://doi.org/10.1155/2014/125704
Microbial Identification System Operational Manual, Version 4, Newark, DE: Microbial ID, 1992.
Stead, D.E., Sellwood, J.E., Wilson, J., and Viney, I., Evaluation of a commercial microbial identification system based on fatty acid profiles for rapid, accurate identification of plant pathogenic bacteria, J. Appl. Bacteriol., 1992, vol. 72, pp. 315–321.
Yan, J.X., Wait, R., Berkelman, T., et al., A modified silver staining protocol for visualization of proteins compatible with matrix-assisted laser desorption/ionization and electrospray ionization-mass spectrometry, Electrophoresis, 2000, vol. 21, pp. 3666–3672. https://doi.org/10.1002/1522-2683(200011)21:17<3666::AID-ELPS3666>3.0.CO;2-6
Shevchenko, A., Chernushevich, I., Wilm, M., and Mann, M., De novo peptide sequencing by nanoelectrospray tandem mass spectrometry using triple quadrupole and quadrupole/time-of-flight instruments, Methods Mol. Biol., 2000, vol. 146, pp. 1–16. https://doi.org/10.1385/1-59259-045-4:1
Davies, D.G., Parsek, M.R., Pearson, J.P., et al., The involvement of cell-to-cell signals in the development of a bacterial biofilm, Science, 1998, vol. 280, pp. 295–298.
Labbate, M., Queck, S.Y., Koh, K.S., et al., Quorum sensing-controlled biofilm development in Serratia liquefaciens MG1, J. Bacteriol., 2004, vol. 186, pp. 692–698. https://doi.org/10.1128/jb.186.3.692-698.2004
Ercolini, D., Russo, F., Nasi, A., et al., Mesophilic and psychrotrophic bacteria from meat and their spoilage potential in vitro and in beef, Appl. Environ. Microbiol., 2009, vol. 75, pp. 1990–2001. https://doi.org/10.1128/AEM.02762-08
Rice, S.A., Koh, K.S., Queck, S.Y., et al., Biofilm formation and sloughing in Serratia marcescens are controlled by quorum sensing and nutrient cues, J. Bacteriol., 2005, vol. 187, pp. 3477–3485. https://doi.org/10.1128/JB.187.10.3477-3485.2005
Zaitseva, Yu.V., Voloshina, P.V., Liu, Kh., et al., Involvement of the global regulators GrrS, RpoS, and SplIR in formation of biofilms in Serratia plymuthica, Russ. J. Genet., 2010, vol. 46, no. 5, pp. 541–545. https://doi.org/10.1134/S1022795410050054
Sinensky, M., Homeoviscous adaptation—a homeostatic process that regulates the viscosity of membrane lipids in Escherichia coli, Proc. Natl. Acad. Sci. U.S.A., 1974, vol. 71, pp. 522–525. https://doi.org/10.1073/pnas.71.2.522
Cybulski, L.E., Albanesi, D., Mansilla, M.C., et al., Mechanism of membrane fluidity optimization: isothermal control of the Bacillus subtilis acyl-lipid desaturase, Mol. Microbiol., 2002, vol. 45, pp. 1379–1388. https://doi.org/10.1046/j.1365-2958.2002.03103.x
Funding
This work was partially supported by the Russian Foundation for Basic Research, projects no. 18-04-00375 and no. 19-04-00756, and by a grant of the Ministry of Education and Sciences of the Russian Federation, State Order for Yaroslavl State University, project no. 0856-2020-0008.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by E. Makeeva
Rights and permissions
About this article
Cite this article
Zaitseva, Y.V., Lipasova, V.A., Koksharova, O.A. et al. Peculiarities of the SprIR Quorum Sensing System of Serratia proteamaculans 94 and Its Involvement in Regulation of Cellular Processes. Russ J Genet 57, 161–172 (2021). https://doi.org/10.1134/S1022795421020149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795421020149