Abstract
Accumulating evidence has demonstrated that long non-coding RNAs (lncRNAs) are key regulators of multiple biological processes by altering gene expression at various levels. Apoptosis in vascular endothelial cells (VECs) is closely linked to numerous cardiovascular diseases, such as arteriosclerosis, thrombus formation and plaque erosion. However, studies on lncRNAs in the cardiovascular system are just beginning. And thus far, no anti-apoptosis lncRNAs have been identified in VECs. Here, we focused on the anti-apoptosis roles of lncRNAs in the serum and FGF-2 starvation-induced apoptosis of VECs. Using microarray analysis, we found a novel lncRNA LOC100129973 which acted as an apoptosis inhibitor in VECs. Through sponging miR-4707-5p and miR-4767, lncRNA LOC100129973 upregulated the expression of two apoptosis repressors gene, Apoptosis Inhibitor 5 (API5) and BCL2 like 12 (BCL2L12) and thus alleviated the serum and FGF-2 starvation-induced apoptosis in VECs. This evidence suggests that lncRNA LOC100129973 is an attractive target to improve endothelial function and for therapy of apoptosis related cardiovascular diseases.
Similar content being viewed by others
Introduction
Long noncoding RNAs (lncRNAs) are defined as non-protein coding transcripts longer than 200 nucleotides without significant protein-coding potential. They constitute a large portion of mammalian transcriptome, since only ~2% of the mammalian genome is composed of genes that encode proteins1. LncRNAs could regulate the expression of genes at the epigenetic, transcriptional and post-transcriptional levels2,3,4. They play important roles in multiple physiological processes such as differentiation, proliferation, apoptosis, invasion and reprogramming of induced pluripotent stem cells5,6,7,8 by several regulatory mechanisms such as interacting with chromatin-modifying enzymes, RNA processing, structural scaffolds and so on9,10,11. In addition, the ability of lncRNAs to function as competing endogenous RNA (CeRNA) was first demonstrated in muscle differentiation5.
Vascular endothelial cells (VECs), which lie in the innermost of blood vessels, are vulnerable to stimulus. Apoptosis in VECs is closely linked to numerous cardiovascular diseases such as arteriosclerosis, thrombus formation and plaque erosion etc.12. Formerly, the investigation on the mechanisms of apoptosis mainly focused on the protein-coding genes. Recently, lncRNAs have attracted more and more interest13,14,15. Yet, there are no reports about apoptosis-related lncRNA in VECs.
Ischemia is a cardiovascular disease generally caused by atherosclerosis or thrombosis16,17 and is associated with apoptosis of VECs due to deficiency of survival growth factors18,19. In our previous work, human umbilical vein endothelial cells (HUVECs) were cultured under the serum and FGF-2-deprived condition to simulate the in vivo ischemic condition. We found that a small molecule, 6-amino-2,3-dihydro-3-hydroxymethyl-1,4-benzoxazine (ABO), elevated the viability of HUVECs in the absence of serum and FGF-220. Moreover, it was demonstrated that ABO effectively inhibited oxLDL-induced apoptosis of VECs21 and atherosclerosis in ApoE−/− mice22. These data suggest that ABO is an appropriate molecule for finding new factors that inhibit VEC apoptosis.
In this study, we aimed to find new factors which repress the serum and FGF-2 starvation-induced apoptosis of VECs by using ABO and microarray. Fortunately, we noticed that lncRNA LOC100129973 was significantly increased by ABO treatment. Furthermore, we demonstrated that through sponging miR-4707-5p and miR-4767, lncRNA LOC100129973 upregulated two apoptosis repressors, Apoptosis Inhibitor 5 (API5) and BCL2 like 2 (BCL2L12) and thus suppressed the serum and FGF-2 starvation-induced apoptosis in HUVECs.
Results
Long noncoding RNA LOC100129973 was upregulated by ABO treatment in HUVECs
Our previous data suggested that ABO is an appropriate molecule for finding new factors which could inhibit VEC apoptosis20,21,22,23,24. By morphological observation, AO staining and TUNEL assay, we confirmed that ABO efficiently inhibited the serum and FGF-2 starvation-induced apoptosis in HUVECs (Supplemental Fig. S1). To gain insights into the possible anti-apoptosis factors in the serum and FGF-2 starvation-induced apoptosis of VECs, we detected the changed transcripts by using ABO and microarray. The microarray assay revealed 22 genes with modified expression, including 6 upregulated genes and 16 downregulated genes in response to 50 μM ABO (Supplementary Table S1). The most significantly upregulated transcript was LOC100129973 (Gene ID: 100129973).
LOC100129973 is a validated long noncoding RNA (lncRNA) and the length of it is 1520 bp. This lncRNA is located in chromosome 15 (21.1) and antisense to guanine nucleotide binding protein, beta 5 (GNB5) and myosin VC (MYO5C) (Fig. 1A). We next examined the expression pattern of lncRNA LOC100129973 over a diverse panel of human cell types including HUVECs, hESCs, L-02 and human tumor cells such as A549, HeLa and PC3. Its expression was detected in all these human cells, while its expression is relative high in HUVECs (Supplemental Fig. S2). According to NCBI database, lncRNA LOC100129973 was only found in Homo sapiens, Rhinopithecus roxellana and Macaca nemestrina. Hence, HUVECs is the ideal model for studying the role of lncRNA LOC100129973. Then, we validated the up-regulation of lncRNA LOC100129973 using quantified real time RT-PCR (Fig. 1B,C). These results showed that in HUVECs, lncRNA LOC100129973 was upregulated by ABO treatment in a dose- and time -dependent manner.
LncRNA LOC100129973 was upregulated by ABO.
(A) The basic information of lncRNA LOC100129973 in human genome. (B) Quantified real-time PCR analysis of lncRNA LOC100129973 expression treated with different concentrations of ABO treatment for 6 h in HUVECs deprived of serum and FGF-2. (C) Quantified real-time PCR analysis of lncRNA LOC100129973 expression treated with 50 μM ABO for various times in HUVECs deprived of serum and FGF-2. Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr). n ≥ 3.
LncRNA LOC100129973 acted as an apoptosis repressor in HUVECs deprived of serum and FGF-2
To better understand the function of lncRNA LOC100129973 in VECs, the full-length lncRNA LOC100129973 was cloned into the pcDNA3.1 expression vector (pcDNA3.1- LOC100129973) and specific siRNAs against lncRNA LOC100129973 (siLOC100129973) were designed and synthesized. HUVECs were transfected with pcDNA3.1- LOC100129973 at 0.1, 0.2, 0.4 μg/mL or siLOC100129973 at 10, 20, 40 nM. The efficiency of overexpression or knockdown was detected by quantified real time RT-PCR (Fig. 2A).
LncRNA LOC100129973 suppressed the serum and FGF-2 starvation-induced apoptosis in HUVECs.
(A) Quantified real-time PCR analysis of lncRNA LOC100129973 overexpression and knock down efficiency. HUVECs were transfected with pcDNA3.1- LOC100129973 at 0.1, 0.2, 0.4 μg/mL or siLOC100129973 at 10, 20, 40 nM for 24 h and then starvation for another 24 h. (B) Hoechst 33258 staining of apoptotic HUVECs. Nor: M199 medium (with FGF-2 and serum); OE-Ctr: transfected with pcDNA3.1-empty vector; OE-LOC100129973: transfected with pcDNA3.1-LOC100129973 at 0.2 μg/mL; siCtr: transfected with scramble RNA for negative control; siLOC100129973: transfected with siLOC100129973 at 40 nM. After being transfected for 24 h, all the cells described above were deprived of serum and FGF-2 for another 12 or 24 h. Scale bar: 10 μm. Images are representative of at least 3 independent experiments. The percent of apoptosis was measured (%). (C,D) Western blot analysis of cleaved PARP protein level. HUVECs were transfected with pcDNA3.1-LOC100129973 at 0.1, 0.2, 0.4 μg/mL or siLOC100129973 at 10, 20, 40 nM for 24 h and starvation for another 24 h. The level of cleaved PARP was relative to that of β-actin. (Cropped, full-length blots are in Supplementary Fig. S4) Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr). n ≥ 3.
To clarify the roles of lncRNA LOC100129973 in the serum and FGF-2 starvation-induced apoptosis of HUVECs, we examined cell viability, nuclear DNA condensation and cleaved PARP by SRB assay, Hoechst 33258 staining and western blot. Cell viability assay showed that enhanced lncRNA LOC100129973 substantially increased cell viability, while the inhibition of lncRNA LOC100129973 decreased it (Supplemental Fig. S3). Furthermore, overexpression of lncRNA LOC100129973 inhibited the serum and FGF-2 starvation-induced apoptosis, whereas knockdown aggravated it (Fig. 2B). Moreover, lncRNA LOC100129973 overexpression efficiently decreased cleaved PARP in HUVECs (Fig. 2C). And for knockdown of lncRNA LOC100129973, the cleaved PARP was evidently promoted (Fig. 2D). Collectively, our data showed that lncRNA LOC100129973 acted as a repressor during the serum and FGF-2 starvation-induced apoptosis in HUVECs.
LncRNA LOC100129973 functioned as an endogenous sponge of miR-4707-5p and miR-4767
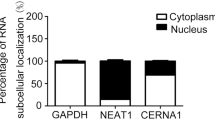
Recent studies have suggested that lncRNAs may act as endogenous sponge RNA to interact with miRNAs and influence the expression of these miRNAs5,25,26,27. To investigate the underlying mechanism of lncRNA LOC100129973 action, we analyzed whether lncRNA LOC100129973 could compete to bind with miRNAs as a miRNA-sponge by Micro Inspector (http://bioinfo.uni-plovdiv.bg/microinspector/). According to the analysis, there are two binding sites for miR-4707-5p and three for miR-4767 (Fig. 3A,B). Meanwhile, the binding free energy of miR-4707-5p is the lowest and miR-4767 is also very low: miR-4707-5p (−35.5) and miR-4767 (−31.4). It is known that miRNAs are present in the cytoplasm in the form of miRNA ribonucleoprotein complexes (miRNPs) that also contain Ago2, the core component of the RNA-induced silencing complex (RISC)28,29. By RNA-fluorescence in situ hybridization (RNA-FISH), lncRNA LOC100129973 was found to be distributed in both nucleus and cytoplasm of HUVECs (Fig. 3C). Moreover, lncRNA LOC100129973 was preferentially enriched in Ago2-containing miRNPs relative to control immunoglobulin G (IgG) immunoprecipitates (Fig. 3D). Therefore, we speculated that lncRNA LOC100129973 competed to bind with miR-4707-5p and miR-4767 as a miRNA-sponge and then regulated the expression of miRNA targets in HUVECs.
LncRNA LOC100129973 might function as a miRNA sponge.
(A,B) Potential sites targeted by miR-4707-5p and miR-4767 in lncRNA LOC100129973. (C) RNA Fluorescent in situ hybridization of the LncRNA LOC100129973 in HUVECs. The antisense probe was used as a negative control. Scale bar: 10 μm. (D) RNA-binding protein immunoprecipitation experiments were performed using Ago2 antibody in HUVECs. IgG was used as a negative control. Quantitative reverse transcription–polymerase chain reaction (qRT-PCR) was performed to detect pulled-down lncRNA LOC100129973. The lncRNA LOC100129973 RNA level of input was set as 100%.
After overexpression or knockdown of lncRNA LOC100129973 for 24 h and starvation for another 24 h, we evaluated the RNA levels of miR-4707-5p and miR-4767 in HUVECs. When lncRNA LOC100129973 expression was enhanced, the RNA levels of miR-4707-5p and miR-4767 were decreased in a dose-dependent manner (Fig. 4A,C); whereas, they were increased in a dose-dependent manner after lncRNA LOC100129973 RNA level was declined (Fig. 4B,D). So lncRNA LOC100129973 negatively regulated the RNA levels of miR-4707-5p and miR-4767 in HUVECs.
LncRNA LOC100129973 regulated the expression of miR-4707-5p and miR-4767 by directly binding with them.
(A,B) miRNA quantified real-time PCR analysis of miR-4707-5p expression after lncRNA LOC100129973 overexpression or knock down for 24 h and starvation for another 24 h in HUVECs. (C,D) miRNA quantified real-time PCR analysis of miR-4767 expression after lncRNA LOC100129973 overexpression or knock down for 24 h and starvation for another 24 h in HUVECs. (E) Luc-LOC100129973-WT or Luc-LOC100129973-ΔmiR-4707-5p plasmids were co-transfected into HEK293T cells with 20, 40, 80 nM miR-4707-5p mimics or scrambled miRNA for 24 h and luciferase activity was measured. (F) Luc-LOC100129973-WT or Luc-LOC100129973-ΔmiR-4767 plasmids were co-transfected into HEK293T cells with 20, 40, 80 nM miR-4767 mimics or scrambled miRNA for 24 h and luciferase activity was measured. Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr). n ≥ 3.
To verify whether lncRNA LOC100129973 act as a miRNA-sponge to direct-negatively regulate miR-4707-5p and miR-4767 RNA levels, we constructed three dual-luciferase miRNA target expression constructs: (1) Luc-LOC100129973-WT; (2) Luc-LOC100129973-ΔmiR-4707-5p (mutated on the putative miR-4707-5p sites); (3) Luc-LOC100129973-ΔmiR-4767 (mutated on the putative miR-4767 sites). Luciferase assays revealed that the mimics of miR-4707-5p and miR-4767 suppressed the luciferase activity of Luc-LOC100129973-WT in a dose-dependent manner, respectively (Fig. 4E,F). However, compared to Luc-LOC100129973-WT, miR-4707-5p mimics had less effect on Luc-LOC100129973-ΔmiR-4707-5p (Fig. 4E). Likewise, miR-4767 mimics had less effect on Luc-LOC100129973-ΔmiR-4767 (Fig. 4F). Taken together, it suggested that lncRNA LOC100129973 downregulated the RNA levels of miR-4707-5p and miR-4767 through directly binding to them.
miR-4707-5p and miR-4767 promoted apoptosis by targeting and downregulating two apoptosis inhibitors, API5 and BCL2L12, respectively
To address whether lncRNA LOC100129973 represses apoptosis though miR-4707-5p and miR-4767, we investigated the roles of miR-4707-5p and miR-4767 in apoptosis of HUVECs. HUVECs were transfected with the mimics or inhibitors of miR-4707-5p and miR-4767 at 25 or 50 nM for 24 h and starvation for another 24 h, the efficiency of overexpression or knockdown was detected (Supplemental Fig. S5). The western blot results showed that the mimics of miR-4707-5p or miR-4767 enhanced the cleaved PARP (Fig. 5A,C), while the inhibitors reduced the cleaved PARP (Fig. 5B,D). It indicated that miR-4707-5p and miR-4767 promoted the serum and FGF-2 starvation-induced apoptosis in HUVECs.
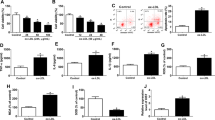
miR-4707-5p and miR-4767 promoted the serum and FGF-2 starvation-induced apoptosis and downregulated the protein of API5 and BCL2L12 in HUVECs.
(A,B) Western blot analysis of cleaved PARP and API5 protein levels after transfection with miR-4707-5p mimics (A) or inhibitor (B) for 24 h and serum starvation for another 24 h in HUVECs. (Cropped, full-length blots are in Supplementary Fig. S7) (C,D) Western blot analysis of cleaved PARP and BCL2L12 protein levels after transfection with miR-4767 mimics (C) or inhibitor (D) for 24 h and serum starvation for another 24 h in HUVECs. (Cropped, full-length blots are in Supplementary Fig. S8) Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr). n ≥ 3.
For further exploring the possible molecular mechanism of miR-4707-5p and miR-4767 in promoting apoptosis, we predicated the target genes of miR-4707-5p and miR-4767 by miRDB, Targetscan and DIANA LAB. Among these potential targets, we concentrated on Apoptosis Inhibitor 5 (API5) and BCL2 like 2 (BCL2L12), both of which inhibit cell apoptosis30,31,32,33. By using RNAhybrid software, we found that 3′UTR of API5 contained the miR-4707-5p-prediction binding sites with the highest score. Likewise, 3′UTR of BCL2L12 contains two miR-4767-prediction binding sites (Supplemental Fig. S6). Furthermore, we tested whether miR-4707-5p and miR-4767 could regulate the expression levels of API5 and BCL2L12. Enhanced RNA level of miR-4707-5p or miR-4767 led to a reduction on the RNA level (Fig. 6A,C) and protein level (Fig. 5A,C) of API5 or BCL2L12 in HUVECs. In contrast, knockdown of endogenous miR-4707-5p or miR-4767 induced an increase on the RNA level (Fig. 6B,D) and protein level (Fig. 5B,D) of API5 or BCL2L12 in HUVECs. These results showed that miR-4707-5p and miR-4767 downregulated the expression of two apoptosis inhibitors, API5 and BCL2L12, respectively.
miR-4707-5p and miR-4767 directly - targeted to API5 and BCL2L12 and regulated the expression of them in HUVECs, respectively.
(A–D) Quantified real-time PCR analysis of API5/BCL2L12 mRNA levels after transfection with miR-4707-5p/miR-4767 mimics (A,C) or inhibitor (B,D) for 24 h and starvation for another 24 h in HUVECs. (E) Luc-API5 3′UTR/Luc-API5 3′UTR -ΔmiR-4707-5p plasmid was co-transfected into HEK293T cells with 20, 40, 80 nM miR-4707-5p mimics or scrambled miRNA for 24 h and luciferase activity was measured. (F) Luc-BCL2L12 3′UTR/Luc-BCL2L12 3′UTR-ΔmiR-4767 plasmid was co-transfected into HEK293T cells with 20, 40, 80 nM miR-4767 mimics or scrambled miRNA for 24 h and luciferase activity was measured. Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr). n ≥ 3.
To verify whether miR-4707-5p and miR-4767 directly target to API5 and BCL2L12 respectively, we constructed four dual-luciferase miRNA target expression constructs: (1) Luc-API5 3′UTR; (2) Luc-API5 3′UTRmut (mutated on the putative miR-4707-5p sites); (3) Luc-BCL2L12 3′UTR; (4) Luc-BCL2L12 3′UTRmut (mutated on the putative miR-4767 sites). Luciferase assays showed that miR-4707-5p mimics suppressed luciferase activity of Luc-API5 3′UTR in a dose-dependent manner, but not Luc-API5 3′UTRmut (Fig. 6E); meanwhile, the mimics of miR-4767 also reduced luciferase activity of Luc-BCL2L12 3′UTR in a dose-dependent manner, but not Luc-BCL2L12 3′UTRmut (Fig. 6F). Collectively, these results suggested that miR-4707-5p and miR-4767 induced apoptosis in HUVECs by directly targeting and decreasing the expression of two apoptosis inhibitors, API5 and BCL2L12, respectively.
LncRNA LOC100129973 promoted the expression of two apoptosis inhibitors, API5 and BCL2L12, by sponging miR-4707-5p and miR-4767
Given that lncRNA LOC100129973 could function as an endogenous sponge of miR-4707-5p and miR-4767, which could reduce the expression of API5 and BCL2L12, we supposed that lncRNA LOC100129973 might regulate the expression of API5 and BCL2L12 through miR-4707-5p and miR-4767. We observed that the mRNA levels of two apoptosis inhibitors, API5 and BCL2L12, were increased after enforced lncRNA LOC100129973 expression in HUVECs (Fig. 7A,C); conversely, they were decreased by transfection with siLOC100129973 (Fig. 7B,D).
LncRNA LOC100129973 positively regulated API5 and BCL2L12 expression mediated by miR-4707-5p and miR-4767.
(A–D) After transfection with pcDNA3.1-LOC100129973 plasmid or siLOC100129973 for 24 h and serum starvation for another 24 h, quantified real-time PCR analysis of API5 (A,B)/BCL2L12 (C,D) expression in HUVECs were performed. (E–H) Quantified real-time PCR analysis of API5/BCL2L12 expression after being cotransfected with pcDNA3.1-LOC100129973 at 0.2 μg/mL and miR-4707-5p/miR-4767 mimics or mimics negative control at 50 nM (E,G); Being cotransfected with siLOC100129973 at 40 nM and miR-4707-5p/miR-4767 inhibitor or inhibitor negative control at 50 nM (F and H) for 24 h and followed with serum starvation for another 24 h. Data are mean ± SEM. of three independent experiments. *P < 0.05, **p < 0.01 vs. control (Ctr).
Furthermore, we investigated the roles of miR-4707-5p and miR-4767 in the lncRNA LOC100129973-induced upregulation of API5 and BCL2L12. Enhanced RNA levels of miR-4707-5p or miR-4767 in HUVECs with lncRNA LOC100129973 overexpression, led to decrease the expression of API5 or BCL2L12 (Fig. 7E,G). Conversely, by reducing RNA levels of miR-4707-5p or miR-4767 in HUVECs with lncRNA LOC100129973 knockdown, the expression level of API5 or BCL2L12 was increased (Fig. 7F,H). Taken together, our data demonstrated that, through sponging miR-4707-5p and miR-4767, lncRNA LOC100129973 was able to positively regulate the expression of API5 and BCL2L12 and thus alleviate the serum and FGF-2 starvation-induced apoptosis in HUVECs.
Discussion
In this study, we first found that lncRNA LOC100129973 was significantly increased by ABO treatment and demonstrated that lncRNA LOC100129973 was an apoptosis inhibitor. Furthermore, lncRNA LOC100129973 negatively regulated miR-4707-5p and miR-4767 by directly binding with them. The miR-4707-5p and miR-4767 aggravated apoptosis by targeting the 3′UTR of two apoptosis inhibitors, API5 and BCL2L12, respectively. Hence, lncRNA LOC100129973 increased the expression of API5 and BCL2L12 by sponging miR-4707-5p and miR-4767 and thus inhibited the serum and FGF-2 starvation-induced apoptosis in HUVECs (Fig. 8).
It has been well demonstrated that lncRNAs take part in multiple human diseases through altering gene expression at various levels. Currently uncovered functions of lncRNAs are classified as the followings: (1) Imprinting; (2) Scaffold/Guide for Epigenetic and Transcription Factors; (3) Enhancer Activation; (4) Molecular Sponges34. Obviously, lncRNA LOC100129973 is a distinct molecular sponge, which could sequester miR-4707-5p and miR-4767 and regulate the expression of their targets.
Cardiovascular diseases are one of the major causes of death worldwide and the morbidity is increasing year by year35. Although studies on lncRNAs in the cardiovascular system are just beginning, more and more reports have shown that lncRNAs are involved in multiple cardiovascular diseases34. For atherosclerosis, lncRNA H19 and ANRIL play roles in the pathologic mechanism of atherosclerosis40. BCL2L12 locates in both cytoplasm and nucleus41. Nuclear BCL2L12 physically interacts with p53 to form a complex and thus eliminates the tumor suppression of p5341,42. BCL2L12 inhibits post-mitochondrial apoptosis signaling - caspase-3/7 via distinct mechanisms43. For caspase 7, BCL2L12 directly interacts with it and then neutralizes its function44. For caspase 3, BCL2L12-induced transcriptional upregulation of the small heat shock protein αB-crystallin is instrumental to neutralization of caspase 3 activation43,45. As a CeRNA, lncRNA LOC100129973 could compete with miR-4767 to control BCL2L12 expression (Fig. 8). Therefore, in HUVECs, decreased apoptosis mediated by lncRNA LOC100129973 may be partially resulted from alleviating activation of caspase-3/7 and p53.
In summary, as shown in Fig. 8, with decoy activity of sequestering miR-4707-5p and miR-4767, lncRNA LOC100129973 promotes the expression of the API5 and BCL2L12 and thus suppresses apoptosis. Therefore, lncRNA LOC100129973 is an attractive target to improve endothelial function and for therapy of apoptosis related cardiovascular diseases.
Methods
Ethics statement
In this study, all subjects filled out a questionnaire which included their informed consent. The human subjects and all experimental procedures were performed in accordance with the ARRIVE guidelines46 and approved by the ethics committee in Shandong University, China.
Cell Culture
Human umbilical vein endothelial cells (HUVECs) were isolated from umbilical cords as described47, then cultured in M199 medium (Gibco, 31100-035) with 10% (v/v) fetal bovine serum (Hyclone, SV30087.02) and 10 IU/mL fibroblast growth factor 2 (FGF-2) in a humidified incubator at 37 °C with 5% CO2. Cells at not more than passage 10 were used for experiments. HEK293T cells were obtained from the Cell Bank of the Chinese Academy of Sciences (Shanghai, China) and were grown in DMEM (Gibco) with 8% fetal bovine serum (FBS; Gibco), penicillin (50 U/ml) and streptomycin (50 μg/ml) (Gibco, 10378-016).
Cell apoptosis assay
The cells treated as mentioned ways for 12 or 24 h were stained with acridin orange (AO, Fluka) for 5 min or Hochest33258 (10 μg/mL) for 15 min 37 °C. Stained cells were washed twice with PBS and then observed under a laser scanning confocal microscope (Leica, DMIRE2, Wetzlar, Germany) or an Olympus (Japan) BH-2 fluorescence microscope. Cells were regarded as apoptotic if nuclei were much brighter or showed condensed chromatin and nuclear fragmentation.
Terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling (TUNEL) was used to detect DNA fragmentation and to calculate apoptotic ratio. TUNEL was carried out following the manufacturer’s instructions (G3250 DeadEndTM Fluorometric TUNEL System, Promega, USA)48. For negative controls, TdT enzyme was not included in the incubation buffer. The apoptotic index was quantified by calculating the number of positive TUNEL cells from total 500 cells in five random microscopic fields.
LncRNA microarray analysis
HUVECs were treated with 50 μM ABO or 0.05% dimethyl sulfoxide (DMSO) in basal M199 medium for 6 h. Then total RNA samples were extracted with Trizol reagent (Invitrogen, 15596018). Agilent human long noncoding RNA 4 × 180K microarray platform was used in this study and lncRNA microarray analysis was performed by CapitalBio Corporation.
Quantitative real-time PCR
Total RNA was extracted from HUVECs by the Trizol reagent method (Invitrogen, Carlsbad, CA, USA) and underwent reverse transcription and quantitative RT-PCR (Roche, Light Cycler 2.0 system) with the primer pair sequences for genes (Supplementary Table S2). The reverse transcription step involved use of the PrimeScript RT reagent kit with gDNA Eraser (DRR047, TAKARA). For miRNAs, miR-4707-5p and miR-4767 were converted into cDNA with the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, PN4366596). The expression of mature miR-4707-5p and miR-4767 in HUVECs were quantified by the KAPA SYBR FAST qPCR Kit (Kapa Biosystems, KK4601). Small RNA U6 was used as internal control for small RNAs. Relative gene expression of miR-4707-5p or miR-4767 was normalized to U6. Quantitative RT-PCR reactions involved use of SYBR Premix Ex Taq (Tli RNaseH Plus) and were carried out in a 20 μl volume with 10 μl of 2X SYBR Green I, 0.4 μl sense primer, 0.4 μl antisense primer, 2 μl cDNA template and 7.2 μl distilled water. Relative gene expression was normalized to that of a housekee** gene (β - actin). The levels of expressed genes were measured by the 2−ΔΔCt method with MxPro 4.00 (Stratagene).
Transient transfection with plasmid in HUVECs
pcDNA3.1-LOC100129973 construct was generated by inserting the full length of lncRNA LOC100129973 into the pcDNA3.1 vector. HUVECs were cultured to full confluent prior to transfection in basic M199 medium and transient transfection was performed using Lipofectamine 2000 (Invitrogen, 11668-019) transfection reagent according to the manufacturer’s instructions. Cells were transfected with pcDNA3.1- empty as control.
Transient transfection with siRNA or miRNA mimics/inhibitor in HUVECs
Specific siRNAs against lncRNA LOC100129973 were synthesized by Invitrogen. Scramble siRNA was used as a control (Santa Cruz, sc-37007). RNA mimics and inhibitor for miR-4707-5p/miR-4767 were designed and purchased from Invitrogen. Corresponding Negative Control (NC) was designed and purchased from Invitrogen. Cells at 70% confluence were transfected with siRNA or miRNA mimics/inhibitor by Lipofectamine 2000 (Invitrogen, 11668–019) transfection reagent according to the manufacturer’s instructions.
Western blot analysis
Treated HUVECs were lysed in protein lysis buffer (Beyotime, P0013). Protein content was determined by use of the BCA Protein Assay Kit (Beyotime, P0011). Proteins were separated by 12% or 9% SDS-PAGE and transferred to PVDF membrane (Millipore, IPVH00010), which was incubated with primary antibodies for PARP (Cell Signaling, 9542L), Apoptosis Inhibitor 5 (API5; Abcam, ab65836), BCL2-like 12 (BCL2L12, Abcam, ab108346) and ACTB (Sigma, 122M4782) at 4 °C overnight and detected with corresponding horseradish peroxidase-conjugated secondary antibody (1:10000) at room temperature for 1 h. The membranes were incubated with Immobilon Western Chemiluminescent HRP Substrate (Millipore, WBKLS0500) for 5 min at room temperature and exposed to X-ray film (Kodak). The relative protein content was analyzed by ImageJ software and normalized to loading controls.
Luciferase reporter transfection and dual luciferase activity assay
HEK293T cells were seeded into 96-well plates at 8000 cells per well for cultured overnight, then co-transfected with plasmids of dual-luciferase (containing firefly and Renilla luciferase) reporters and miRNA mimics or NC (final concentration, 20, 40 or 80 nM) with Lipofectamine 2000 (Invitrogen, 11668–019) for 24 h. Dual luciferase activity was measured by the Dual-Glo Luciferase Assay System (Promega, E2920). After reagent was added, firefly luciferase or Renilla luciferase activity was measured by VICTOR X2 Multilabel Plate Reader (PerkinElmer, USA). Firefly luciferase activity was normalized to that of Renilla.
RNA fluorescent in situ hybridization (RNA-FISH)
LncRNA LOC100129973 subcellular localization in HUVECs was detected by use of a FISH kit (Roche Applied Science, Germany). Briefly, HUVECs were fixed in 4% paraformaldehyde, then prehybridized with hybridization solution and incubated with a digoxigenin-labeled LncRNA LOC100129973 probe. The antisense probe was used as a negative control. Cell nuclei were stained with 4′,6-diamidino-2-phenylindole (DAPI) for 5 min at room temperature. Fluorescence images were obtained by use of an LSM 700 Confocal Laser Scanning Microscope (Carl Zeiss).
RNA-binding protein immunoprecipitation (RIP) assay
RNA immunoprecipitation (RIP) experiments were performed with the Magna RIP RNA-Binding Protein Immunoprecipitation Kit (Millipore, 17–701) following the manufacturer’s instructions. The RIPAb+ Ago2 antibody (Millipore, 03–110) was used in RIP and mouse IgG was used as a negative control. The PCR primers for lncRNA LOC100129973 are listed in Supplementary Table S2.
Statistical analysis
Data are presented as mean ± SEM and analysis involved use of GraphPad Prism 5. Images were processed by Adobe Photoshop CS5 (Adobe, San Jose, USA). P < 0.05 was considered statistically significant.
Additional Information
How to cite this article: Lu, W. et al. Long Noncoding RNA LOC100129973 Suppresses Apoptosis by Targeting miR-4707-5p and miR-4767 in Vascular Endothelial Cells. Sci. Rep. 6, 21620; doi: 10.1038/srep21620 (2016).
References
Perkel, J. M. Visiting “noncodarnia”. Biotechniques 54, 301, 303–4 (2013).
Hung, T. & Chang, H. Y. Long noncoding RNA in genome regulation: prospects and mechanisms. RNA Biol 7, 582–5 (2010).
Yoon, J. H., Abdelmohsen, K. & Gorospe, M. Posttranscriptional gene regulation by long noncoding RNA. J Mol Biol 425, 3723–30 (2013).
Lee, J. T. Epigenetic regulation by long noncoding RNAs. Science 338, 1435–9 (2012).
Cesana, M. et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147, 358–69 (2011).
Keniry, A. et al. The H19 lincRNA is a developmental reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell Biol 14, 659–65 (2012).
Khaitan, D. et al. The melanoma-upregulated long noncoding RNA SPRY4-IT1 modulates apoptosis and invasion. Cancer Res 71, 3852–62 (2011).
Loewer, S. et al. Large intergenic non-coding RNA-RoR modulates reprogramming of human induced pluripotent stem cells. Nat Genet 42, 1113–7 (2010).
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′UTRs via Alu elements. Nature 470, 284–8 (2011).
Bertani, S., Sauer, S., Bolotin, E. & Sauer, F. The noncoding RNA Mistral activates Hoxa6 and Hoxa7 expression and stem cell differentiation by recruiting MLL1 to chromatin. Mol Cell 43, 1040–6 (2011).
Clemson, C. M. et al. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol Cell 33, 717–26 (2009).
Rajendran, P. et al. The vascular endothelium and human diseases. Int J Biol Sci 9, 1057–69 (2013).
Karreth, F. A. & Pandolfi, P. P. ceRNA cross-talk in cancer: when ce-bling rivalries go awry. Cancer Discov 3, 1113–21 (2013).
Tay, Y., Rinn, J. & Pandolfi, P. P. The multilayered complexity of ceRNA crosstalk and competition. Nature 505, 344–52 (2014).
Kartha, R. V. & Subramanian, S. Competing endogenous RNAs (ceRNAs): new entrants to the intricacies of gene regulation. Front Genet 5, 8 (2014).
Proctor, E. Assisted circulation in the experimental preparation-with special reference to pulmonary embolism and acute myocardial ischemia. Prog Cardiovasc Dis 12, 271–92 (1969).
Conti, C. R. & Mehta, J. L. Acute myocardial ischemia: role of atherosclerosis, thrombosis, platelet activation, coronary vasospasm and altered arachidonic acid metabolism. Circulation 75, V84–95 (1987).
Hogg, N. et al. Apoptosis in vascular endothelial cells caused by serum deprivation, oxidative stress and transforming growth factor-beta. Endothelium 7, 35–49 (1999).
Cho, S. W. et al. Delivery of small interfering RNA for inhibition of endothelial cell apoptosis by hypoxia and serum deprivation. Biochem Biophys Res Commun 376, 158–63 (2008).
Jiao, P. F. et al. Design, synthesis and preliminary biological evaluation of 2,3-dihydro-3-hydroxymethyl-1,4-benzoxazine derivatives. Bioorg Med Chem Lett 16, 2862–7 (2006).
Liu, X. et al. Protective effects of a benzoxazine derivative against oxidized LDL-induced apoptosis and the increases of integrin beta4, ROS, NF-kappaB and P53 in human umbilical vein endothelial cells. Bioorg Med Chem Lett 19, 2896–900 (2009).
Li, H. et al. Targeting annexin A7 by a small molecule suppressed the activity of phosphatidylcholine-specific phospholipase C in vascular endothelial cells and inhibited atherosclerosis in apolipoprotein E(−)/(−)mice. Cell Death Dis 4, e806 (2013).
Li, H. et al. Identification of a small molecule targeting annexin A7. Biochim Biophys Acta 1833, 2092–9 (2013).
Huang, S. et al. TIA1 interacts with annexin A7 in regulating vascular endothelial cell autophagy. Int J Biochem Cell Biol 57, 115–22 (2014).
Wang, J. et al. CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 38, 5366–83 (2010).
Cazalla, D., Yario, T. & Steitz, J. A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 328, 1563–6 (2010).
Ge, D. et al. Identification of a novel MTOR activator and discovery of a competing endogenous RNA regulating autophagy in vascular endothelial cells. Autophagy 10, 957–71 (2014).
Filipowicz, W., Bhattacharyya, S. N. & Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9, 102–14 (2008).
Izaurralde, E. Elucidating the temporal order of silencing. EMBO Rep 13, 662–3 (2012).
Morris, E. J. et al. Functional identification of Api5 as a suppressor of E2F-dependent apoptosis in vivo. PLoS Genet 2, e196 (2006).
Garcia-Jove Navarro, M. et al. Api5 contributes to E2F1 control of the G1/S cell cycle phase transition. PLoS One 8, e71443 (2013).
Noh, K. H. et al. API5 confers tumoral immune escape through FGF2-dependent cell survival pathway. Cancer Res 74, 3556–66 (2014).
Yang, M. C. et al. Bcl2L12 with a BH3-like domain in regulating apoptosis and TMZ-induced autophagy: a prospective combination of ABT-737 and TMZ for treating glioma. Int J Oncol 46, 1304–16 (2015).
Uchida, S. & Dimmeler, S. Long noncoding RNAs in cardiovascular diseases. Circ Res 116, 737–50 (2015).
Ounzain, S., Crippa, S. & Pedrazzini, T. Small and long non-coding RNAs in cardiac homeostasis and regeneration. Biochim Biophys Acta 1833, 923–33 (2013).
Li, L., **e, J., Zhang, M. & Wang, S. Homocysteine harasses the imprinting expression of IGF2 and H19 by demethylation of differentially methylated region between IGF2/H19 genes. Acta Biochim Biophys Sin (Shanghai) 41, 464–71 (2009).
Holdt, L. M. et al. Alu elements in ANRIL non-coding RNA at chromosome 9p21 modulate atherogenic cell functions through trans-regulation of gene networks. PLoS Genet 9, e1003588 (2013).
Tewari, M. et al. AAC-11, a novel cDNA that inhibits apoptosis after growth factor withdrawal. Cancer Res 57, 4063–9 (1997).
Van den Berghe, L. et al. FIF [fibroblast growth factor-2 (FGF-2)-interacting-factor], a nuclear putatively antiapoptotic factor, interacts specifically with FGF-2. Mol Endocrinol 14, 1709–24 (2000).
Scorilas, A. et al. Molecular cloning, physical map** and expression analysis of a novel gene, BCL2L12, encoding a proline-rich protein with a highly conserved BH2 domain of the Bcl-2 family. Genomics 72, 217–21 (2001).
Stegh, A. H. & DePinho, R. A. Beyond effector caspase inhibition: Bcl2L12 neutralizes p53 signaling in glioblastoma. Cell Cycle 10, 33–8 (2011).
Stegh, A. H. et al. Glioma oncoprotein Bcl2L12 inhibits the p53 tumor suppressor. Genes Dev 24, 2194–204 (2010).
Stegh, A. H. et al. Bcl2L12-mediated inhibition of effector caspase-3 and caspase-7 via distinct mechanisms in glioblastoma. Proc Natl Acad Sci USA 105, 10703–8 (2008).
Stegh, A. H. et al. Bcl2L12 inhibits post-mitochondrial apoptosis signaling in glioblastoma. Genes Dev 21, 98–111 (2007).
Stegh, A. H., Chin, L., Louis, D. N. & DePinho, R. A. What drives intense apoptosis resistance and propensity for necrosis in glioblastoma? A role for Bcl2L12 as a multifunctional cell death regulator. Cell Cycle 7, 2833–9 (2008).
Kilkenny, C., Browne, W. J., Cuthill, I. C., Emerson, M. & Altman, D. G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol 8, e1000412 (2010).
Jaffe, E. A., Nachman, R. L., Becker, C. G. & Minick, C. R. Culture of human endothelial cells derived from umbilical veins. Identification by morphologic and immunologic criteria. J Clin Invest 52, 2745–56 (1973).
Gavrieli, Y., Sherman, Y. & Ben-Sasson, S. A. Identification of programmed cell death in situ via specific labeling of nuclear DNA fragmentation. J Cell Biol 119, 493–501 (1992).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (No. 81321061, 91313303, 91539105, 31270877, 20972088 and 31070735) and the National 973 Research Project (No. 2011CB503906).
Author information
Authors and Affiliations
Contributions
J.Y.M. and B.X.Z. designed the research, W.L., S.Y.H. and L.S. performed the experiments. J.Y.M., B.X.Z. and W.L. analyzed the data and wrote the paper. All authors reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Lu, W., Huang, S., Su, L. et al. Long Noncoding RNA LOC100129973 Suppresses Apoptosis by Targeting miR-4707-5p and miR-4767 in Vascular Endothelial Cells. Sci Rep 6, 21620 (2016). https://doi.org/10.1038/srep21620
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep21620
- Springer Nature Limited
This article is cited by
-
Non-Coding RNA-Mediated Gene Regulation in Cardiovascular Disorders: Current Insights and Future Directions
Journal of Cardiovascular Translational Research (2023)
-
LncRNA NEAT-2 regulate the function of endothelial progenitor cells in experimental Sepsis model
Molecular Biology Reports (2023)
-
The Long Non-coding Road to Atherosclerosis
Current Atherosclerosis Reports (2020)
-
Long noncoding RNAs: emerging roles in pulmonary hypertension
Heart Failure Reviews (2020)
-
Long Noncoding RNA-CERNA1 Stabilized Atherosclerotic Plaques in apolipoprotein E−/− Mice
Journal of Cardiovascular Translational Research (2019)