Abstract
The diversity of cell types is a challenge for quantifying aging and its reversal. Here we develop ‘aging clocks’ based on single-cell transcriptomics to characterize cell-type-specific aging and rejuvenation. We generated single-cell transcriptomes from the subventricular zone neurogenic region of 28 mice, tiling ages from young to old. We trained single-cell-based regression models to predict chronological age and biological age (neural stem cell proliferation capacity). These aging clocks are generalizable to independent cohorts of mice, other regions of the brains, and other species. To determine if these aging clocks could quantify transcriptomic rejuvenation, we generated single-cell transcriptomic datasets of neurogenic regions for two interventions—heterochronic parabiosis and exercise. Aging clocks revealed that heterochronic parabiosis and exercise reverse transcriptomic aging in neurogenic regions, but in different ways. This study represents the first development of high-resolution aging clocks from single-cell transcriptomic data and demonstrates their application to quantify transcriptomic rejuvenation.
Similar content being viewed by others
Main
Aging is the progressive deterioration of cellular and organismal function. Age-dependent decline is linked in large part to the passage of time and therefore the chronological age of an individual. But such decline is not inexorable. At the same chronological age, some individuals have better organismal and tissue fitness (biological age) than others. Furthermore, aging trajectories can be slowed, and some aspects of aging can be reversed by specific interventions, including dietary restriction, exercise, reprogramming factors, senolytic compounds and young blood factors1,2,3,4,5,6. As aging is the primary risk factor for many diseases, particularly neurodegenerative diseases7,8, a better understanding of aging and ‘rejuvenation’ strategies could yield large benefits for a wide range of diseases.
Aging is complex and difficult to quantify. One quantification approach is to use machine learning to build age prediction models—‘aging clocks’—which can serve as integrative aging biomarkers. Such clocks should also accelerate our understanding of existing interventions and help identify new strategies to counter aging and age-related diseases. Machine learning models trained on high-dimensional datasets (for example, DNA methylation, transcriptomics and proteomics) can predict chronological age with remarkable accuracy. For example, regression-based aging clocks trained on DNA methylation profiles from multiple tissues (‘epigenetic aging clocks’)9,10,11,12,13 or blood plasma protein profiles14,15,16,17 have striking performance to predict chronological age in humans. Aging clocks directly optimized to predict biological age have also been developed on functional phenotypes12,13,18 or time remaining until death19,20. Interestingly, beneficial health interventions such as diet and exercise21,22,23 and genetic manipulations24,25,26 result in younger predictions from epigenetic aging clocks trained on chronological age. Thus, epigenetic aging clocks, despite being trained on chronological age, also capture dimensions of biological age.
So far, molecular aging clocks have largely relied on datasets built using bulk tissue input or purified cell populations9,10,11,12,13,27,28,29,30,31,32,33,34. Bulk tissue profiles (and even purified populations) average the molecular profiles from many cells, integrating tissue composition changes and cell-type-specific responses. Hence, the cell-type-specific contributions to aging and rejuvenation detected by these clocks remain unclear. While single-cell DNA methylation and transcriptomic data have started to be used to classify age35,36,37, cell-type-specific transcriptomic aging clocks have not yet been generated. Thus, it remains to be determined if aging clocks of different cell types ‘tick’ at different rates, which cell types predict age most accurately and how specific cell types respond to different interventions. The rapid advance of single-cell RNA-sequencing (RNA-seq) technologies provides a great opportunity to explore these unaddressed questions and identify new molecular aging clocks to study interventions to counter aging and age-related diseases.
Results
Cell-type-specific transcriptomic aging clocks
As a paradigm for tissue aging and functional decline in the brain, we focused on the neurogenic region located in the subventricular zone (SVZ) of the adult mammalian brain. The SVZ neurogenic region (or ‘niche’) contains neural stem cells (NSCs) that give rise to differentiated cells (neurons, astrocytes) that are important for olfactory discrimination and repair upon injury38,39,40,41,42,43,44,45. Importantly, this neurogenic region contains at least 11 different cell types and experiences age-related changes correlated with deterioration in tissue function42,46,47,48,11,21,22,23,24,31,32. Our results also highlight cell-type specificity for aging and possibly for rejuvenation interventions. This is unique to single-cell-based clocks and will allow a better understanding of cell heterogeneity in tissue aging and rejuvenation. Our data also reveal different potential for rejuvenation strategies, at least at the transcriptional level. These results raise the exciting possibility that aging clocks can serve to rapidly test the efficacy of rejuvenation interventions and to support combining specific interventions to counter aging and age-related diseases.
Methods
Our research complies with all relevant ethical regulations (AAALAC), under Institutional Animal Care and Use (IACUC) protocols 8661 and 16246 at Stanford University and VA Palo Alto Committee on Animal Research, ACORP LUO1736.
Animals
For aging cohorts and the exercise cohort, male C57BL/6 mice were obtained from the National Institute on Aging (NIA) Aged Rodent colony. For parabiosis cohort 1, old mice were male C57BL/6 mice from the NIA Aged Rodent colony and young mice were male B6.SJL-Ptprca Pepcb/BoyJ male (Pep boy) from the Jackson Laboratory. For parabiosis cohort 2, old mice were male C57BL/6J and young mice were male C57BL/6J or C57BL/6-Tg(UBC-GFP)30Scha/J from the Jackson Laboratory. Mice were housed in the Comparative Medicine Pavilion, ChemH/Neuroscience Vivarium or the SIM-1 Non-Barrier Rodent Facility at Stanford, or in the Veterinary Medical Unit at the Palo Alto VA. All these facilities provide equivalent standard conditions with a 12-h light–dark cycle, ad libitum food and water, ~21 °C temperature, and ~50% humidity. All mice were acclimated to their vivarium for at least 2 weeks before use in any experiment.
Tissue and cell collection for the subventricular zone neurogenic niche
For single-cell RNA-seq datasets, SVZ neurogenic niches were collected and processed as described in ref. 48. Briefly, mice were sedated with 1 ml of 2.5% vol/vol Avertin (Sigma-Aldrich, T48402-25G) and perfused with 15 ml of PBS (Corning, 21-040-CV) with heparin sodium salt (50 U ml−1; Sigma-Aldrich, H3149-50KU) to remove the blood, and brain collection was performed immediately. As previously described104, the SVZ from each hemisphere was microdissected and dissociated with enzymatic digestion with papain at a concentration of 14 U ml−1, rocking for 10 min at 37 °C. Note that the samples also contained some of the surrounding striatum, which contributed to the oligodendrocyte population in our study. The dissociated SVZ was triturated in a solution containing 0.7 mg ml−1 ovomucoid and 0.5 mg ml−1 DNase I (Sigma-Aldrich, DN25-100MG) in DMEM/F12 (Thermo Fisher, 11330032). The dissociated cells from the SVZ were centrifuged through 22% Percoll (Sigma-Aldrich, GE17-0891-01) in PBS to remove myelin debris. After centrifugation, cells were filtered through a 35-μm snap-cap filter (Corning, 352235), washed once with 1.5 ml of FACS buffer (HBSS (Thermo Fisher, 14175103), 1% BSA (Sigma, A7979) and 0.1% glucose (Sigma-Aldrich, G7021-1KG)) and spun down for 5 min at 300g. Cells were resuspended in 120 μl FACS buffer with live/dead staining performed using 1 μg ml−1 propidium iodide (BioLegend, 421301) and kept on ice until sorting. FACS sorting was performed on a BD FACS Aria II sorter, using a 100-μm nozzle at 13.1 PSI. Cells were sorted into low protein binding microcentrifuge tubes containing 750 μl of PBS with 1% BSA and 0.1% glucose. When not applying sample multiplexing (parabiosis cohort 1 and exercise cohort), cells were then centrifuged (300g for 5 min at 4 °C) and resuspended in 50 μl FACS buffer, counted and then immediately run on 10x Chromium to capture single-cell transcriptomes.
Cohorts of mice of different ages
To generate the single-cell RNA-seq dataset from mice of different ages and train aging clock models, we used four independent cohorts of aging mice. Each cohort had 4–8 male C57BL/6 mice from the NIA Aged Rodent colony, for a total of 28 mice. These 28 mice tiled 26 different ages (two pairs of mice had the same age), ranging from 3.3 months (young adult) to 29 months (geriatric adult).
Lipid-modified oligonucleotide multiplexing
Sample multiplexing was performed using LMOs, a method also known as MULTI-seq50. Lipid anchor and co-anchor reagents were kindly provided by the Gartner Laboratory at the University of California, San Francisco and custom oligonucleotides were ordered from Integrated DNA Technologies. We used MULTI-seq primer: 5′ CTTGGCACCCGAGAATTCC; and Universal.I5: 5′AATGATACGGCGACCACCGAGATCTACACTCTTTCCCTACACGACGCTCTTCCGATCT50.
We followed the exact protocol outlined by McGinnis et al.50 with the following modifications: (1) all labeling with LMOs was performed in a 4 °C cold room because, in our hands, the quality of labeling was very sensitive to temperature; (2) to avoid cell loss and cell clum**, cells were sorted into PBS with 2% BSA, and BSA was then removed using three PBS washes; (3) concentrations and volumes were adjusted to account for low cell numbers: 7.5 μl of 1 mM lipid anchor with oligonucleotide barcode mix was added to a 70 μl volume of resuspended cells followed by 7.5 μl of 1 mM lipid co-anchor; (4) labeling reactions were quenched with 2% BSA then samples were pooled before subsequent 1% BSA PBS washes to further reduce cell loss. The combined sample was resuspended at 50 μl for cell counting and single-cell RNA-seq.
Single-cell libraries and RNA sequencing
Single-cell RNA-seq was performed using a 10x Chromium machine and 10x Genomics V3.0 Transcriptomics kits (aging cohorts, parabiosis cohort 2 and exercise cohort) or a 10x Genomics V2 kit (parabiosis cohort 1). For sequencing, 10,000 cells per lane were targeted but typical yields were approximately 5,000 cells. Library preparation was done according to the manufacturer’s protocol (10x Genomics V3.0 or 10x Genomics V2 for parabiosis cohort 1). Sequencing was done to target a minimum of 25,000 reads per cell for transcriptome characterization and 5,000 reads per cell for LMO label recovery. The aging cohorts and the parabiosis cohort 2 samples were multiplexed with 4–8 samples per 10x Chromium lane. The parabiosis cohort 1 and the exercise samples were not multiplexed with LMO reagents. Sequencing was performed on either an Illumina HiSeq 4000 (aging cohorts and parabiosis cohort 1) or a NovoSeq using the 2 × 150-bp setting (parabiosis cohort 2 and exercise).
Analysis (quality control)
Cell Ranger (version 3.0.2) default settings were used to distinguish cells from background. Subsequent analysis was performed using R (version 3.6.3). Cells were filtered out in Seurat (version 3.2.3)105,106 if they contained fewer than 500 genes or greater than 10% mitochondrial reads. Small clusters of doublets that shared several marker genes from pure populations were identified and removed. LMO demultiplexing was performed using Seurat’s HTODemux function. A complete view of the data processing and quality-control parameters can be found at https://github.com/sunericd/svz_singlecell_aging_clocks.
Cell type annotation
Cell types in all datasets were manually annotated as described in ref. 48, and cross-referenced with annotations present in the single-cell database PanglaoDB107. Identification of major clusters was performed with the FindClusters() algorithm in the Seurat package, which uses a shared nearest-neighbor modularity optimization-based clustering algorithm106. Marker genes for each major cluster were found using the Seurat (version 4.1.1) function FindAllMarkers() using the Wilcoxon rank-sum test. Cell types were determined using marker genes identified from the literature and the marker genes were cross-referenced with annotations present in the single-cell database PanglaoDB107. This analysis identified ~11 clusters of cells (depending on the dataset), including astrocytes and qNSCs, aNSCs and NPCs, neuroblasts, neurons, oligodendrocyte progenitor cells, oligodendrocytes, endothelial cells, ‘mural’ cells (pericytes or smooth muscle) and microglia. The genes used for identification are included in Supplementary Table 2 and a clustering of a subset of these genes is presented in Extended Data Fig. 1c.
Consistent with our previous study48, we did not observe sufficient differences in transcriptomic signatures to separate astrocytes from qNSCs and aNSCs from NPCs. We have described these clusters as ‘astrocyte-qNSCs’ and ‘aNSC-NPCs’ throughout this study. Some cell types were not identified when using the LMO protocol (for example, T cells), probably because cells such as T cells are small and their membranes may not allow for efficient LMO labeling. We also identified only a few ependymal cells in several of our datasets, although these cells are known to be numerous in the SVZ neurogenic niche. This is probably because ependymal cells are too big to be efficiently uploaded in droplets and/or they are sheared in the 10x microfluidic device.
Cell cycle annotation and proliferative fraction
For cell cycle annotation (G1, S, G2/M) of cells in the SVZ neurogenic niche, we used Seurat’s CellCycleScoring function with default parameters. This annotation was used to calculate the ‘proliferative fraction’ in the SVZ neurogenic niche, that is, the percentage of cells predicted to be in S or G2/M phase. We used the proliferative fraction (ProliferativeFraction) as a functional metric of the SVZ neurogenic niche and used it to define ‘biological age’ in this study (‘Age prediction and validation strategy’).
To test the correlation between chronological age and proliferative fraction in the SVZ neurogenic niche, we used Pearson’s correlation. There was a negative correlation (Pearson R = −0.8) between chronological age and proliferative fraction in the SVZ.
Age prediction and validation strategy
Chronological or biological age (‘label’) was regressed onto all log-normalized gene expression values ln((gene transcripts / cell transcripts) × 10,000) (‘features’) in a particular cell type using the R package glmnet (version 4.0.2)51. To determine the most robust method to predict age from single-cell RNA-seq data, we tested various preprocessing approaches: SingleCell, Pseudobulk, BootstrapCell (‘BoostrapCell preprocessing’) and EnsembleCell (‘EnsembleCell preprocessing’). SingleCell uses bona fide single-cell transcriptomes with minimal processing as input to a lasso regression model to predict chronological or biological age. Pseudobulk involves naïve pseudobulking all cells from the same cell type and sample before using a lasso regression model to predict chronological or biological age. BoostrapCell uses lasso regression models and EnsembleCell uses elastic net models (described separately below)53. There was no manual filtering of genes. Both lasso regression and elastic net regression enforce sparsity in the model coefficients with tunable parameter such that only a subset of genes will have nonzero coefficients in the trained aging clock models.
Chronological age was defined as months since birth. Biological age was defined as 35 – (ProliferativeFraction × 100) where ProliferativeFraction was the number of cells predicted to be in S or G2/M phase divided by the total number of cells from that sample. The number 35 was selected to transform biological age into the same range as chronological age.
For validation, models were built on 3 of the 4 cohorts of mice, and validation was done on the remaining cohort (stringent ‘leave-one-cohort-out’ validation (cross-cohort validation)). For training of each model, hyperparameters were optimized with fivefold to tenfold cross validation. To quantify the performance of the models, the data were presented as a correlation between the actual chronological (or biological) age of the mouse from which the cell originated (x axis) and the median predicted chronological (or biological) age for that mouse (y axis). Density of cells is represented with graded colors and each mouse is represented as a dot. We fitted a linear model (black line) through the points as well as the 95% confidence interval (light gray) using geom_smooth (ggplot2). Pearson’s correlation (R) is indicated on the graph. In dot plots, both the R values and the MAE, that is, median absolute error across all the cells, are presented.
To test the correlation between chronological age and biological age, we used the Pearson correlation. There was a positive correlation (R = 0.84) between chronological age and biological age predictions.
BootstrapCell preprocessing
BootstrapCell uses a lasso model with the following characteristics: To generate a BootstrapCell, 15 single-cell transcriptomes were sampled without replacement from the pool of cells of a given cell type from a given animal (for example, oligodendrocytes from a single mouse). Gene counts were then summed. A BootstrapCell constructed from 15 cells was empirically found to balance the tradeoff between sample number and gene coverage per sample. This bootstrap** process was repeated 100 times for each cell type–animal combination. BootstrapCells were used as input into lasso regression models. This approach had the effect of normalizing the contribution of each animal rather than each single-cell transcriptome.
EnsembleCell preprocessing
We devised and evaluated a second preprocessing and age prediction technique to compare to our BootstrapCell approach and to test robustness to changes in preprocessing and model architecture. In the EnsembleCell approach, 20 elastic net models were trained for each cell type. For each model, gene expression data from cells were randomly partitioned into groups of 15 single-cell transcriptomes and the unique transcript counts for all cells in each group were summed to create ‘EnsembleCells’. To predict age from the gene expression profile of a cell, we used the weighted average of predictions across all 20 models, where weights were determined by the R2 (coefficient of determination) of the model on a held-out validation set (‘Age prediction and validation strategy’).
Use of aging clocks on independent mouse datasets
We determined if the single-cell-based models (‘aging clocks’) generated from our mouse SVZ neurogenic niche dataset could be applied to cells from an independent dataset and even to cells from another neurogenic region in the brain. To this end, we used a single-cell RNA-seq dataset of the SVZ neurogenic niche from young and old mice48 and a single-cell RNA-seq dataset of the dentate gyrus of the hippocampus from mice of three different ages60. These datasets were preprocessed as described above using the ‘BootstrapCell’ method. We examined the distribution of the predicted chronological or biological ages of each cell in these datasets, color coded by the age of the mouse of origin.
Use of aging clocks on human datasets
To determine if the single-cell-based aging clocks generated from the mouse SVZ neurogenic niche could apply to cells from other regions of the brain and in other species, we used a single-nucleus RNA-seq dataset of the middle temporal gyrus from humans of different ages61. The dataset was preprocessed using the ‘BootstrapCell’ method as described above. As oligodendrocytes and astrocytes were present both in the human dataset and our mouse SVZ neurogenic niche dataset, we applied our oligodendrocyte and astrocyte-qNSC chronological aging clocks to the corresponding cell types in the human dataset. We rescaled the raw predictions linearly to obtain rescaled predicted chronological ages for each human BootstrapCell (rescaled predicted age = m × raw predicted age + b, where m = 10 and b = 125.5 for oligodendrocytes; m = 5 and b = 32.75 for astrocytes). The linear rescaling did not change the reported correlation between predicted chronological age and actual chronological age. Correlation plots were generated as described in ‘Age prediction and validation strategy’.
Cell-type-specific aging clocks using Tabula Muris Senis
To determine whether the method we used to derive cell-type-specific aging clocks was generalizable to tissues other than neurogenic niches, we used the count matrices from the single-cell RNA-seq dataset of the multi-tissue aging atlas Tabula Muris Senis62. We chose three diverse cell types in different tissues: endothelial cells from limb muscle, mature natural killer T cells from spleen and podocytes from kidney. For each cell type, the data were preprocessed and aging clocks were trained using the BootstrapCell approach described above. The performance of these models was evaluated by iteratively training on all mice except for one mouse and obtaining predictions on the held-out mouse (‘leave-one-mouse-out’ cross validation (cross-mouse validation) instead of ‘leave-one-cohort-out’ cross validation (cross-cohort validation) because there were no distinct cohorts in this dataset).
Identification of genes that contribute to the aging clocks
Genes that contribute to each aging clock model were retrieved by selecting all genes from the clocks with nonzero coefficients (Supplementary Table 4). The weight of a gene on each clock model (that is, the level of contribution based on coefficient values) and the sign of the coefficient (positive, higher gene expression is associated with older age; negative, lower gene expression is associated with older age) are indicated using a donut plot, with sector size indicating the gene weight and color indicating coefficient sign. Genes with positive coefficient are mostly upregulated with age, and genes with negative coefficient are mostly downregulated with age. The regulation of each chronological and biological clock gene (compared to other genes) is presented using a volcano plot (Extended Data Fig. 4). Most genes selected by the clocks were differentially expressed during aging. Less than half of the genes selected by chronological and biological aging clocks in a particular cell type overlapped (Supplementary Table 4). To determine if chronological or biological clock genes were shared across cell types or specific to each cell types, we used UpSet plots. Most genes selected by chronological or biological clocks were cell-type specific. The ‘impact’ (sum of absolute values of coefficient) and ‘count’ (sum of gene number) of shared genes or specific genes are indicated as a stacked bar plot.
Properties of genes that contribute to the aging clocks
To determine if genes that contribute to the aging clocks have specific properties, we examined their variability by plotting the coefficient of variation as a function of mean expression. Genes used by the clocks were more highly expressed and, at a given level of expression, had a higher coefficient of variation (that is, were more variable) than genes not in the clock (Extended Data Fig. 3a).
We also verified that the increased variability of genes that contribute to the clocks was not merely due to sparsity in the single-cell RNA-seq dataset. On average, the majority of cells (for each cell type) express the genes that contribute to the clocks and this is higher than what was observed for genes that do not contribute to the clock (Extended Data Fig. 3b).
Gene-set enrichment analysis
GSEA was performed using Enrichr108 to query cell-type-specific clock genes for enrichment against GO biological process gene sets. Statistics were exported from the Enrichr web tool and processed and visualized in R with ggplot2 (version 3.3.3) package.
Parabiosis cohorts and single-cell RNA-seq dataset
Two independent cohorts of heterochronic parabiosis were generated (cohort 1 and cohort 2). Parabiosis cohort 1 involved six male mice across three pairings. We collected SVZ niches from one isochronic young mouse (5 months, control), one heterochronic young mouse (5 months, old blood), one heterochronic old mouse (26 months, young blood) and one isochronic old mouse (26 months, control), for a total of four SVZ niches (of six mice). Old parabionts were C57BL/6 male mice from the NIA Aged Rodent colony at Charles River. Young parabionts were B6.SJL-Ptprca Pepcb/BoyJ male (Pep boy) mice from The Jackson Laboratory and C57BL/6 male mice from the NIA. Of the young, only the Pep boy mice were used for transcriptomics. Congenic (rather than isogenic) pairings were performed to enable verification of blood chimerism by FACS with antibodies specific to CD45.1 (BioLegend, 110705; 1:100 dilution) or CD45.2 (BioLegend, 109814; 1:100 dilution) alleles. Mice were 4 and 25 months old at the start of the experiment, and parabiosis was conducted for 5 weeks until cell collection, when mice were 5 and 26 months old. Pairs were established as previously described69,75,80 by suturing the peritoneums of adjacent flanks and joining skin with surgical clips. Five weeks after the parabiosis surgery, mice were anaesthetized with 2.5% vol/vol avertin, euthanized by cardiac puncture and perfused with 15 ml PBS with heparin (50 U ml−1). SVZ dissection, digestion and FACS were performed as describe above. 10x Genomics single-cell transcriptome V2 libraries (one sample per 10x lane) were generated and sequenced on one Illumina HiSeq lane by the Stanford Function Genomics Facility. Animal care and parabiosis procedures were performed in accordance with Stanford University under IACUC protocols 8661 and 16246.
Parabiosis cohort 2 involved eighteen male mice across nine pairings. We collected SVZ niches from four isochronic young mice (5 months, control), four heterochronic young mice (5 months, old blood), four heterochronic old mice (21 months, young blood) and six isochronic old mice (21 months, control), for a total of eighteen SVZ niches (of eighteen mice). All mice in this cohort were sourced from the Jackson Laboratory and housed in the Veterinary Medical Unit at the Palo Alto VA77. Old mice were C57BL/6J and young were C57BL/6J or C57BL/6-Tg(UBC-GFP)30Scha/J. Mice were aged 4 and 19.5 months at the start of the experiment, and parabiosis proceeded for 5 weeks until cell collection, when mice were 5 and 21 months old. Surgeries were performed as described above. Five weeks after surgery, mice were anesthetized with 2.5% vol/vol avertin, euthanized by cardiac puncture and perfused with 15 ml PBS with heparin (50 U ml−1). SVZ dissection, digestion and FACS were performed as describe above. Tissue collection took place on three separate days and samples were multiplexed with LMOs. 10x Genomics single-cell transcriptome V3 libraries were generated in-house and sequenced by Novogene on an Illumina NovoSeq lane. Animal care and parabiosis procedures were approved by the VA Palo Alto Committee on Animal Research and listed on ACORP LUO1736.
Parabiosis cohort 1 and cohort 2 were generated in different animal facilities, by different surgeons, in different years, and they were analyzed with different versions of 10x Genomics single-cell transcriptomics kits. For visualization, data from the two independent cohorts were integrated on the cohort identity using the RunHarmony command from Harmony109. There were no statistically significant differences between young isochronic (control) predicted chronological ages across cohorts in all six cell-type-specific aging clocks (Wilcoxon rank-sum test for median predicted chronological ages), suggesting that there was not a major batch effect that could have influenced the age prediction.
Exercise cohort and single-cell RNA-seq dataset
C57BL/6 male mice from the NIA Aged Rodent colony at Charles River were housed in the Veterinary Medical Unit at the Palo Alto VA97. Young and old mice were aged 4.5 months and 21.5 months, respectively, at the start of the 5-week voluntary wheel running intervention, so they were 6 months and 23 months when tissues were collected. During the intervention period, mice (n = 4 for each age group) were singly housed in cages accommodating a running wheel. Control mice (n = 3–4 for each age group) had no access to a wheel. Running was verified by recording wheel revolutions. After 5 weeks, mice were anaesthetized with 2.5% v/v avertin, euthanized by cardiac puncture, perfused and cell suspensions from dissected SVZs generated as described in ‘Tissue and cell collection for the SVZ neurogenic niche’. Next, 10x Genomics V3.0 transcriptomics kits were used to generated libraries without upstream sample multiplexing. Tissue processing occurred across two separate mornings. SVZ libraries were pooled and sequenced on an Illumina NovoSeq.
Effect of rejuvenation interventions on the aging clocks
To measure the effect of heterochronic parabiosis and exercise on the aging clocks, we examined the distribution of predicted chronological or biological ages as described in ‘Use of aging clocks on independent mouse datasets’. We calculated the effect by the difference in median predicted chronological or biological age between intervention and control. In dot plots, these differences were represented as ‘effect’, using size and intensity of color, with blue indicating ‘rejuvenation’ and red indicating ‘aging’.
Comparison of heterochronic parabiosis and exercise effects
To compare the effect of heterochronic parabiosis and exercise, we calculated the mean of the difference between the median predicted chronological age for a mouse for each intervention (data from cohort 1 and cohort 2 for heterochronic parabiosis). Genes that were reversed by each intervention or by both, based on direction of average log fold change, were identified.
Differential expression analysis
To determine genes that were impacted by different interventions independently of the aging clocks, we used differential expression analysis, focusing on aNSC-NPCs (as this cell type is impacted by both interventions). MAST110 software was used to calculate differential expression statistics between three different conditions: age (young versus old), young blood (heterochronic parabiosis versus isochronic old control), exercise (exercise versus sedentary in old mice). To determine the DEGs between young and old, we defined ‘young’ as mice <7 months and ‘old’ as mice >20 months. Permissive cutoffs of 1.1-fold change and FDR < 0.1 were applied in each of the three different conditions. Overlap was presented as a Venn diagram.
Gene signature analysis
For specific gene signature analysis, we summed the expression of genes in one cell type from single-cell transcriptomic datasets within a specific gene signature defined by a specific GO term. Among the different signatures tested, we selected those that were significantly increased with age and reversed by at least one intervention. We focused on two signatures: the ‘interferon-γ response’ signature defined as the sum of all normalized expression values of genes in the interferon gene set defined by Dulken et al.48 and the ‘negative regulation of neurogenesis’ gene signature defined as the sum of all normalized expression values of genes in the GO term ‘negative regulation of neurogenesis’ gene set (v6.21)111. Data were presented as violin plots and statistical analyses were performed using the Wilcoxon rank-sum test at the cell level.
Intervention classification models
To evaluate the aging relevance of ‘rejuvenation’ interventions, we generated cell-type-specific models trained on the intervention rather than age as a label. We used classification models, based on logistic regression (cv.glmnet(type.measure = ‘mse’, family = ‘binomial’) using all log-normalized gene expression values ln((gene transcripts / cell transcripts) × 10,000) as features. These intervention classification models were trained on single-cell RNA-seq data from heterochronic parabiosis (young blood) versus isochronic parabiosis old (control) or from exercise versus sedentary old mice. The data were preprocessed using the same BootstrapCell approach as described above. For logistic regression, the label used corresponded to either the intervention (‘0’) or control (‘1’). Cross validation was performed on held-out cells (25% of the cells that were not used to build the models). After training and validating the intervention classification models, we applied these models to the single-cell RNA-seq dataset of the SVZ neurogenic niche from 28 mice, tiling 26 ages from young (3.3 months) to old (29 months). Data were plotted as described in ‘Age prediction and validation strategy’, with (log(p(control) / p(intervention))) as a function of the actual chronological age of aNSC-NPC BootstrapCell transcriptomes. Old mice were more likely to be classified as ‘isochronic old control’, whereas young mice were more likely to be classified as ‘heterochronic old’, indicating that the gene signature that distinguishes exposure to young and old blood is relevant to aging. R is the Pearson correlation. Higher correlation indicates that the main intervention signature overlaps with and reverses age-related changes. Correlations between intervention state prediction and chronological age across cell types and interventions were assessed, with a separate classifier built for each. The exercise classifiers were built to distinguish old sedentary from old exercised transcriptomes for each cell type. The lower correlation between intervention state predictions and age for the exercise samples implies that the signatures that distinguishes exercised and sedentary mice are less related to aging than those derived from parabiosis intervention classifiers.
Statistics and reproducibility
No statistical methods were used to predetermine sample sizes; we determined our sample sizes based on our previous analysis of similar types of datasets48. For study design, we used four independent cohorts of mice, each spanning different ages, to build the age prediction models. This design allows us to test the machine learning aging clock models with a robust cross-cohort validation (that is, ‘leave-one-cohort-out’ validation). Two independent experiments of heterochronic parabiosis were performed, involving 6 mice (4 collected, cohort 1) and 18 mice (cohort 2), with data collection spread across 4 d. One experiment of exercise (with controls lacking a running wheel) was performed, involving 15 mice processed across 2 d. Animals from group 3 from parabiosis cohort 2 were excluded because sample multiplexing failed and it was not possible to distinguish samples. The experiments were not randomized. Investigators were not blinded to allocation during experiments and outcome assessment, although the genomics analyses were performed in a systematic manner. To test correlations, we used Pearson’s correlation. To determine the statistical significance of the differences between intervention and control, we used the Wilcoxon rank-sum test (a non-parametric test).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All raw sequencing reads and key processed files are accessible at BioProject PRJNA795276 (aging, parabiosis) and the Gene Expression Omnibus under accession GSE196364 (exercise). Processed data files for the aging and parabiosis data can be found at https://doi.org/10.5281/zenodo.7145399. Processed data files for the exercise data can be found at https://doi.org/10.5281/zenodo.7338746. External raw sequencing reads for the mouse hippocampus dataset are accessible at the Gene Expression Omnibus under accession GSE159768. External data on human middle temporal gyrus are accessible at https://portal.brain-map.org/atlases-and-data/rnaseq/human-mtg-smart-seq/. External data from Tabula Muris Senis are accessible at https://figshare.com/projects/Tabula_Muris_Senis/64982/. PanglaoDB can be accessed at https://panglaodb.se/.
Code availability
The code used to analyze genomic data and generate aging clocks in the current study is available in the GitHub repository for this paper (https://github.com/sunericd/svz_singlecell_aging_clocks/). A frozen version of the code repository is available as Supplementary Software 1.
References
Ocampo, A., Reddy, P. & Belmonte, J. C. I. Anti-aging strategies based on cellular reprogramming. Trends Mol. Med. 22, 725–738 (2016).
Pluvinage, J. V. & Wyss-Coray, T. Systemic factors as mediators of brain homeostasis, ageing and neurodegeneration. Nat. Rev. Neurosci. 21, 93–102 (2020).
Mahmoudi, S., Xu, L. & Brunet, A. Turning back time with emerging rejuvenation strategies. Nat. Cell Biol. 21, 32–43 (2019).
Rando, T. A. & Chang, H. Y. Aging, rejuvenation, and epigenetic reprogramming: resetting the aging clock. Cell 148, 46–57 (2012).
Green, C. L., Lamming, D. W. & Fontana, L. Molecular mechanisms of dietary restriction promoting health and longevity. Nat. Rev. Mol. Cell Biol. 23, 56–73 (2022).
Campisi, J. et al. From discoveries in ageing research to therapeutics for healthy ageing. Nature 571, 183–192 (2019).
Niccoli, T. & Partridge, L. Ageing as a risk factor for disease. Curr. Biol. 22, R741–R752 (2012).
Partridge, L., Deelen, J. & Slagboom, P. E. Facing up to the global challenges of ageing. Nature 561, 45–56 (2018).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, 3156 (2013).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Simpson, D. J. & Chandra, T. Epigenetic age prediction. Aging Cell 20, e13452–e13452 (2021).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging 10, 573–591 (2018).
Belsky, D. W. et al. Quantification of the pace of biological aging in humans through a blood test, the DunedinPoAm DNA methylation algorithm. Elife 9, e54870 (2020).
Tanaka, T. et al. Plasma proteomic signature of age in healthy humans. Aging Cell 17, e12799 (2018).
Lehallier, B., Shokhirev, M. N., Wyss-Coray, T. & Johnson, A. A. Data mining of human plasma proteins generates a multitude of highly predictive aging clocks that reflect different aspects of aging. Aging Cell 19, e13256 (2020).
Menni, C. et al. Circulating proteomic signatures of chronological age. J. Gerontol. A Biol. Sci. Med. Sci. 70, 809–816 (2015).
Lehallier, B. et al. Undulating changes in human plasma proteome profiles across the lifespan. Nat. Med. 25, 1843–1850 (2019).
Sun, E. D. et al. Predicting physiological aging rates from a range of quantitative traits using machine learning. Aging 13, 23471–23516 (2021).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging 11, 303–327 (2019).
Schultz, M. B. et al. Age and life expectancy clocks based on machine learning analysis of mouse frailty. Nat. Commun. 11, 4618 (2020).
Fitzgerald, K. N. et al. Potential reversal of epigenetic age using a diet and lifestyle intervention: a pilot randomized clinical trial. Aging 13, 9419–9432 (2021).
Quach, A. et al. Epigenetic clock analysis of diet, exercise, education, and lifestyle factors. Aging 9, 419–446 (2017).
Thompson, M. J. et al. A multi-tissue full lifespan epigenetic clock for mice. Aging 10, 2832–2854 (2018).
Petkovich, D. A. et al. Using DNA methylation profiling to evaluate biological age and longevity interventions. Cell Metab 25, 954–960 (2017).
Lu, Y. et al. Reprogramming to recover youthful epigenetic information and restore vision. Nature 588, 124–129 (2020).
Martin-Herranz, D. E. et al. Screening for genes that accelerate the epigenetic aging clock in humans reveals a role for the H3K36 methyltransferase NSD1. Genome Biol. 20, 146 (2019).
Peters, M. J. et al. The transcriptional landscape of age in human peripheral blood. Nat. Commun. 6, 8570 (2015).
Fleischer, J. G. et al. Predicting age from the transcriptome of human dermal fibroblasts. Genome Biol. 19, 221 (2018).
Skene, N. G., Roy, M. & Grant, S. G. A genomic lifespan program that reorganises the young adult brain is targeted in schizophrenia. Elife 6, e17915 (2017).
Stubbs, T. M. et al. Multi-tissue DNA methylation age predictor in mouse. Genome Biol. 18, 68 (2017).
Meyer, D. H. & Schumacher, B. BiT age: a transcriptome-based aging clock near the theoretical limit of accuracy. Aging Cell 20, e13320 (2021).
Holzscheck, N. et al. Modeling transcriptomic age using knowledge-primed artificial neural networks. NPJ Aging Mech Dis. 7, 15 (2021).
Nie, C. et al. Distinct biological ages of organs and systems identified from a multi-omics study. Cell Rep. 38, 110459 (2022).
Gill, D. et al. Multi-omic rejuvenation of human cells by maturation phase transient reprogramming. Elife 11, e71624 (2022).
Singh, S. P. et al. Machine learning based classification of cells into chronological stages using single-cell transcriptomics. Sci. Rep. 8, 17156 (2018).
Trapp, A., Kerepesi, C. & Gladyshev, V. N. Profiling epigenetic age in single cells. Nature Aging 1, 1189–1201 (2021).
Lu, J. et al. Heterogeneity and transcriptome changes of human CD8+ T cells across nine decades of life. Nat. Commun. 13, 5128 (2022).
Faiz, M. et al. Adult neural stem cells from the subventricular zone give rise to reactive astrocytes in the cortex after stroke. Cell Stem Cell 17, 624–634 (2015).
Lim, D. A. & Alvarez-Buylla, A. The adult ventricular-subventricular zone (V-SVZ) and olfactory bulb (OB) neurogenesis. Cold Spring Harb. Perspect. Biol. 8, a018820 (2016).
Bond, A. M., Ming, G.-l & Song, H. Adult mammalian neural stem cells and neurogenesis: five decades later. Cell Stem Cell 17, 385–395 (2015).
Williamson, M. R., Jones, T. A. & Drew, M. R. Functions of subventricular zone neural precursor cells in stroke recovery. Behav. Brain Res. 376, 112209 (2019).
Navarro Negredo, P., Yeo, R. W. & Brunet, A. Aging and rejuvenation of neural stem cells and their niches. Cell Stem Cell 27, 202–223 (2020).
Gage, F. H. & Temple, S. Neural stem cells: generating and regenerating the brain. Neuron 80, 588–601 (2013).
Silva-Vargas, V., Crouch, E. E. & Doetsch, F. Adult neural stem cells and their niche: a dynamic duo during homeostasis, regeneration, and aging. Curr. Opin. Neurobiol. 23, 935–942 (2013).
Conover, J. C. & Todd, K. L. Development and aging of a brain neural stem cell niche. Exp. Gerontol. 94, 9–13 (2017).
Artegiani, B. et al. A single-cell RNA-sequencing study reveals cellular and molecular dynamics of the hippocampal neurogenic niche. Cell Rep. 21, 3271–3284 (2017).
Kalamakis, G. et al. Quiescence modulates stem cell maintenance and regenerative capacity in the aging brain. Cell 176, 1407–1419 (2019).
Dulken, B. W. et al. Single-cell analysis reveals T cell infiltration in old neurogenic niches. Nature 571, 205–210 (2019).
**e, X. P. et al. High-resolution mouse subventricular zone stem-cell niche transcriptome reveals features of lineage, anatomy, and aging. Proc. Natl Acad. Sci. USA 117, 31448 (2020).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
Friedman, J., Hastie, T. & Tibshirani, R. Regularization paths for generalized linear models via coordinate descent. J. Stat. Softw. 33, 1–22 (2010).
Tibshirani, R. Regression shrinkage and selection via the lasso. J. R. Stat. Soc. Series B 58, 267–288 (1996).
Zou, H. & Hastie, T. Regularization and variable selection via the elastic net. J. R. Stat. Soc. Series B 67, 301–320 (2005).
Levine, M. E. et al. DNA methylation age of blood predicts future onset of lung cancer in the women’s health initiative. Aging 7, 690–700 (2015).
Marioni, R. E. et al. DNA methylation age of blood predicts all-cause mortality in later life. Genome Biol. 16, 25 (2015).
Maslov, A. Y., Barone, T. A., Plunkett, R. J. & Pruitt, S. C. Neural stem cell detection, characterization, and age-related changes in the subventricular zone of mice. J. Neurosci. 24, 1726–1733 (2004).
Enwere, E. et al. Aging results in reduced epidermal growth factor receptor signaling, diminished olfactory neurogenesis, and deficits in fine olfactory discrimination. J. Neurosci. 24, 8354–8365 (2004).
Tropepe, V., Craig, C. G., Morshead, C. M. & van der Kooy, D. Transforming growth factor-alpha null and senescent mice show decreased neural progenitor cell proliferation in the forebrain subependyma. J. Neurosci. 17, 7850–7859 (1997).
Shook, B. A., Manz, D. H., Peters, J. J., Kang, S. & Conover, J. C. Spatiotemporal changes to the subventricular zone stem cell pool through aging. J. Neurosci. 32, 6947–6956 (2012).
Harris, L. et al. Coordinated changes in cellular behavior ensure the lifelong maintenance of the hippocampal stem cell population. Cell Stem Cell 28, 863–876 (2021).
Hodge, R. D. et al. Conserved cell types with divergent features in human versus mouse cortex. Nature 573, 61–68 (2019).
Tabula Muris, C. A single-cell transcriptomic atlas characterizes ageing tissues in the mouse. Nature 583, 590–595 (2020).
Zhao, T. et al. PISD is a mitochondrial disease gene causing skeletal dysplasia, cataracts, and white matter changes. Life Sci. Alliance 2, e201900353 (2019).
Thomas, H. E. et al. Mitochondrial complex I activity is required for maximal autophagy. Cell Rep. 24, 2404–2417 (2018).
Cheriyath, V., Leaman, D. W. & Borden, E. C. Emerging roles of FAM14 family members (G1P3/ISG 6-16 and ISG12/IFI27) in innate immunity and cancer. J. Interferon Cytokine Res. 31, 173–181 (2011).
Ruckh, J. M. et al. Rejuvenation of regeneration in the aging central nervous system. Cell Stem Cell 10, 96–103 (2012).
Salpeter, S. J. et al. Systemic regulation of the age-related decline of pancreatic beta cell replication. Diabetes 62, 2843–2848 (2013).
Loffredo, F. S. et al. Growth differentiation factor 11 is a circulating factor that reverses age-related cardiac hypertrophy. Cell 153, 828–839 (2013).
Villeda, S. A. et al. Young blood reverses age-related impairments in cognitive function and synaptic plasticity in mice. Nat. Med. 20, 659–663 (2014).
Sinha, M. et al. Restoring systemic GDF11 levels reverses age-related dysfunction in mouse skeletal muscle. Science 344, 649–652 (2014).
Baht, G. S. et al. Exposure to a youthful circulaton rejuvenates bone repair through modulation of beta-catenin. Nat. Commun. 6, 7131 (2015).
Gontier, G. et al. Tet2 rescues age-related regenerative decline and enhances cognitive function in the adult mouse brain. Cell Rep. 22, 1974–1981 (2018).
Sousa-Victor, P. et al. MANF regulates metabolic and immune homeostasis in ageing and protects against liver damage. Nat. Metab. 1, 276–290 (2019).
Brack, A. S. et al. Increased Wnt signaling during aging alters muscle stem cell fate and increases fibrosis. Science 317, 807–810 (2007).
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005).
Ma, S. et al. Heterochronic parabiosis induces stem cell revitalization and systemic rejuvenation across aged tissues. Cell Stem Cell. 29, 990–1005 (2022).
Palovics, R. et al. Molecular hallmarks of heterochronic parabiosis at single-cell resolution. Nature 603, 309–314 (2022).
Katsimpardi, L. et al. Vascular and neurogenic rejuvenation of the aging mouse brain by young systemic factors. Science 344, 630–634 (2014).
Castellano, J. M., Kirby, E. D. & Wyss-Coray, T. Blood-borne revitalization of the aged brain. JAMA Neurol. 72, 1191–1194 (2015).
Villeda, S. A. et al. The ageing systemic milieu negatively regulates neurogenesis and cognitive function. Nature 477, 90–94 (2011).
Smith, L. K. et al. beta2-microglobulin is a systemic pro-aging factor that impairs cognitive function and neurogenesis. Nat. Med. 21, 932–937 (2015).
Rebo, J. et al. A single heterochronic blood exchange reveals rapid inhibition of multiple tissues by old blood. Nat. Commun. 7, 13363 (2016).
Yousef, H. et al. Aged blood impairs hippocampal neural precursor activity and activates microglia via brain endothelial cell VCAM1. Nat. Med. 25, 988–1000 (2019).
Gonzalez-Armenta, J. L., Li, N., Lee, R. L., Lu, B. & Molina, A. J. A. Heterochronic parabiosis: old blood induces changes in mitochondrial structure and function of young mice. J. Gerontol. A Biol. Sci. Med. Sci. 76, 434–439 (2021).
van Praag, H., Christie, B. R., Sejnowski, T. J. & Gage, F. H. Running enhances neurogenesis, learning, and long-term potentiation in mice. Proc. Natl Acad. Sci. USA 96, 13427–13431 (1999).
van Praag, H., Shubert, T., Zhao, C. & Gage, F. H. Exercise enhances learning and hippocampal neurogenesis in aged mice. J. Neurosci. 25, 8680–8685 (2005).
Nokia, M. S. et al. Physical exercise increases adult hippocampal neurogenesis in male rats provided it is aerobic and sustained. J. Physiol. 594, 1855–1873 (2016).
Blackmore, D. G. et al. An exercise ‘sweet spot’ reverses cognitive deficits of aging by growth-hormone-induced neurogenesis. iScience 24, 103275 (2021).
Kannangara, T. S. et al. Running reduces stress and enhances cell genesis in aged mice. Neurobiol. Aging 32, 2279–2286 (2011).
Marlatt, M. W., Potter, M. C., Lucassen, P. J. & van Praag, H. Running throughout middle-age improves memory function, hippocampal neurogenesis, and BDNF levels in female C57BL/6J mice. Dev. Neurobiol. 72, 943–952 (2012).
Horowitz, A. M. et al. Blood factors transfer beneficial effects of exercise on neurogenesis and cognition to the aged brain. Science 369, 167–173 (2020).
Brown, J. et al. Enriched environment and physical activity stimulate hippocampal but not olfactory bulb neurogenesis. Eur. J. Neurosci. 17, 2042–2046 (2003).
Lee, J. C., Yau, S. Y., Lee, T. M. C., Lau, B. W. & So, K. F. Voluntary wheel running reverses the decrease in subventricular zone neurogenesis caused by corticosterone. Cell Transplant. 25, 1979–1986 (2016).
Bednarczyk, M. R., Aumont, A., Decary, S., Bergeron, R. & Fernandes, K. J. Prolonged voluntary wheel-running stimulates neural precursors in the hippocampus and forebrain of adult CD1 mice. Hippocampus 19, 913–927 (2009).
Mastrorilli, V. et al. Physical exercise rescues defective neural stem cells and neurogenesis in the adult subventricular zone of Btg1 knockout mice. Brain Struct. Funct. 222, 2855–2876 (2017).
Lupo, G. et al. Molecular profiling of aged neural progenitors identifies Dbx2 as a candidate regulator of age-associated neurogenic decline. Aging Cell 17, e12745 (2018).
Liu, L. et al. Exercise reprograms the inflammatory landscape of multiple stem cell compartments during mammalian aging. Preprint at bioRxiv https://doi.org/10.1101/2022.01.12.475145 (2022).
Slavov, N. Driving single cell proteomics forward with innovation. J. Proteome Res. 20, 4915–4918 (2021).
Bahar, R. et al. Increased cell-to-cell variation in gene expression in ageing mouse heart. Nature 441, 1011–1014 (2006).
Enge, M. et al. Single-cell analysis of human pancreas reveals transcriptional signatures of aging and somatic mutation patterns. Cell 171, 321–330 (2017).
Martinez-Jimenez, C. P. et al. Aging increases cell-to-cell transcriptional variability upon immune stimulation. Science 355, 1433–1436 (2017).
Salzer, M. C. et al. Identity noise and adipogenic traits characterize dermal fibroblast aging. Cell 175, 1575–1590 (2018).
Levine, M. et al. A rat epigenetic clock recapitulates phenotypic aging and co-localizes with heterochromatin. Elife 9, e59201 (2020).
Codega, P. et al. Prospective identification and purification of quiescent adult neural stem cells from their in vivo niche. Neuron 82, 545–559 (2014).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Satija, R., Farrell, J. A., Gennert, D., Schier, A. F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Franzen, O., Gan, L. M. & Bjorkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA-sequencing data. Database 2019, baz046 (2019).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene-set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Finak, G. et al. MAST: a flexible statistical framework for assessing transcriptional changes and characterizing heterogeneity in single-cell RNA-sequencing data. Genome Biol. 16, 278 (2015).
Bult, C. J. et al. Mouse Genome Database (MGD) 2019. Nucleic Acids Res. 47, D801–D806 (2019).
Acknowledgements
We thank members of the laboratory of A.B., especially C. Bedbrook, J. Chen, P. Navarro, A. Reeves, X. Zhao and R. Yeo for reading the manuscript and critical discussion. We thank C. Bedbrook, P. Navarro and P. P. Singh for independent checking of the code. We thank C. S. McGinnis from the Gartner Laboratory at the University of California, San Francisco for providing MULTI-seq reagents. We thank A. Kundaje for fruitful discussion. We thank L. Bonanno and J. Luo for conducting surgeries in Parabiosis cohort 2, and A. Yang and T. Iram from the T.W.-C. laboratory for facilitating SVZ collection in Parabiosis cohort 2. We thank the Stanford Shared FACS Facility for FACS use and technical support. Elements of Figures 1a,e,g, 4a, 5a and 7a were created with BioRender.com. Support was provided by NSF Graduate Research Fellowship (to M.T.B. and E.D.S.), P.D. Soros Fellowship for New Americans (to E.D.S.), Knight-Hennessy Scholars Program (to E.D.S.), HHMI James H. Gilliam Fellowship (to J.M.R.), P01AG036695 (to A.B., T.A.R. and M.A.G.), the Milky Way Research Foundation (to A.B.), Chan Zuckerberg Initiative award (to A.B.), Simons Foundation grant (to A.B.), R01AG072255 (to T.W.-C), and a generous gift from M. and T. Barakett (to A.B.).
Author information
Authors and Affiliations
Contributions
M.T.B., E.D.S. and A.B. planned the study. M.T.B. generated all datasets and performed all analyses, except for those indicated below. E.D.S. performed analyses with EnsembleCell clocks, external clock validation and generalization in mouse and human, derivation of Tabula Muris Senis clocks, gene overlap, testing of characteristics of clock genes, statistical analysis of intervention effect and gene signatures and independent code checking. B.M.G. and N.S. performed heterochronic parabiosis pairings, under the supervision of I.L.W. and T.W.-C., respectively. L.L. set up the exercise intervention under the supervision of T.A.R. J.M.R. and L.L. provided help in the analysis of the exercise data, under the supervision of M.A.G. and T.A.R., respectively. L.X. helped with dissection of neurogenic regions for single-cell RNA-seq and independent code checking. M.T.B., E.D.S. and A.B. wrote the manuscript, and all authors provided comments.
Corresponding author
Ethics declarations
Competing interests
M.T.B. is a cofounder of Retro Biosciences. The remaining authors declare no competing interests.
Peer review
Peer review information
Nature Aging thanks the anonymous reviewers for the contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
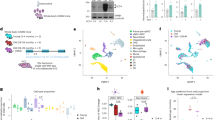
Extended Data Fig. 1 Characteristics of the SVZ single-cell transcriptomic data and generalization performance of cell-type-specific aging clocks to humans and generalization of the framework to other cell types and tissues.
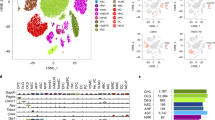
a, Lipid-modified oligonucleotide cell barcodes detected from 8 SVZ samples multiplexed in one 10x lane. b, Same as in (a) but visualized using tSNE. Samples 5 and 6 were from mice of the same age (and colored in the same color in Fig. 1c). c, Heatmap of single cell gene expression for top 5 cell type markers used for annotation of cell type clusters. Colored bar on top indicates the various cell type clusters. d, Overview of Pearson correlation coefficients (R) and median absolute error (MAE) values for tested methods of predicting chronological age across cell types from single-cell transcriptomic data, including both full distribution (Full) and median metrics only (Median). Performance is based on cross-cohort-validation. e, Correlation plot to assess the generalizability of chronological aging clocks (BoostrapCell) to a human dataset from Hodge et al61. the single-nucleus RNA-seq dataset of the middle temporal gyrus of human patients of different ages (Hodge et al.61). Density of BootstrapCell predictions is depicted in color and overlaid black dots represent the median prediction for each sample. R values are Pearson’s correlation coefficients at the sample level. Bands correspond to 95% confidence interval. f, Performance of chronological aging clocks (BootstrapCell) derived from the single cell RNA-seq multi-tissue atlas Tabula Muris Senis, 2020. Predicted chronological age of endothelial cells from limb muscle, mature natural killer (NK) T cells from spleen, and podocyte cells from kidney from aging clocks built on Tabula Muris Senis as a function of actual chronological age for several mice of different ages. Density of BootstrapCell predictions is depicted in color and overlaid black dots represent the median prediction for each sample. Performance is based on cross-mouse validation. R values are Pearson’s correlation coefficients at the sample level.
Extended Data Fig. 2 Genes that contribute to the chronological aging clocks and biological aging clocks.
a, Contribution of individual genes to the chronological aging clocks (BootstrapCell) (see Fig. 3a for aNSC-NPCs). Donut plots, with sector size denoting gene weight in the model and color indicating sign of expression change with age. Total number of genes used by the clock is provided in the center of each donut plot. Positive coefficients (orange) indicate increased gene expression is associated with older age. Negative coefficients (blue) indicate decreased gene expression is associated with older age. b, As in (a) but for biological aging clocks (BootstrapCell) and their coefficients. c, Upset plot illustrating the intersection of gene sets used by cell-type-specific biological aging clocks. No genes were used in all 6 biological aging clocks. d, Count and coefficient impact of shared and cell-type-specific clock genes for chronological aging clocks. Shared is defined as present in at least one of the other five clocks (see Fig. 3c for aNSC-NPCs). e, As in (d) but for biological aging clocks. f, Top enriched Gene Ontology Biological Process terms from gene set enrichment analysis of genes used in biological aging clocks. Shared genes (present in two or more clocks) are enriched for cytokine-mediated signaling pathway and cellular response to type I interferon. The aNSC-NPC biological aging clock genes are enriched for cell cycle pathways.
Extended Data Fig. 3 Variability and mean expression of genes in the chronological aging clocks.
a, Scatter plots of the log2 coefficient of variation (CV) of the normalized BootstrapCell gene expression as a function of the log2 mean normalized BootstrapCell gene expression for all identified genes in the six different cell types. Red dots correspond to genes that contribute to the chronological aging clocks for each cell type (selected by clock) and black dots correspond to genes that do not contribute to the chronological aging clocks (not selected by clock). b, As in (a) but for the fraction of cells with nonzero counts in the dataset as a function of the log2 coefficient of variation (CV) of the normalized gene expression.
Extended Data Fig. 4 Comparison of genes in cell-type-specific aging clocks and differentially expressed genes with age.
a, Volcano plots of negative log10 false discovery rate (FDR) from differential gene expression analysis using MAST for the young and old mouse groups. We defined ‘young’ as mice <7 months old and ‘old’ as mice >20 months old. Colored dots correspond to genes that contribute to the chronological aging clocks for each cell type (selected by clock) (orange representing genes with positive clock coefficients and blue representing genes with negative clock coefficients) and gray dots correspond to that do not contribute to the chronological aging clocks (not selected by clock). b, As in (a) but for the biological aging clocks.
Extended Data Fig. 5 Gene detection rate in cell-type-specific aging clocks.
a, Unique genes detected in single cell transcriptomes from the subventricular zone as a function of gene detection rate. Red dots indicate unique genes detected at 2%, 20%, and 80% detection rates. b, Upset plots showing transcriptome overlaps between cell types at different levels of expression detection. Most genes are shared if a very low threshold of detection is used. Above 80% detection rate, transcriptomes are very cell type specific. However, the shared core of easily detected genes in transcriptomes (70 genes) is much larger than the shared core of genes selected by clocks (~1).
Extended Data Fig. 6 Application of cell-type-specific aging clocks to heterochronic parabiosis cohorts.
a, UMAP projection of single-cell transcriptomes labeled by cell type from both parabiosis cohorts. b, Scatter plot of log2 mean BootstrapCell expression of all genes in the parabiosis cohort 1 data compared to the same genes in the parabiosis cohort 2 data for aNSC-NPCs. Red dots correspond to genes that contribute to the aNSC-NPC chronological aging clock (selected by clock). Gray dots correspond to genes that do not contribute to the aNSC-NPC chronological aging clock (not selected by clock). c, Density plots for prediction of chronological age using chronological aging clocks for Parabiosis cohort 1 (BootstrapCell). d, Density plots for prediction of chronological age using chronological aging clocks for Parabiosis cohort 2 (BoostrapCell). e, Density plots for prediction of chronological age using chronological aging clocks for both cohorts separated by mouse (BootstrapCell). f, Density plots for prediction of biological age using biological aging clocks for Parabiosis cohort 1 (BootstrapCell). g, Density plots for prediction of biological age using biological aging clocks for Parabiosis cohort 2 (BoostrapCell). h, Density plots for prediction of biological age using biological aging clocks for both cohorts separated by mouse (BoostrapCell).
Extended Data Fig. 7 Statistical comparison of the rejuvenation effect size for heterochronic parabiosis and exercise at the mouse level.
a, Violin plots of the median predicted chronological ages for all mice in parabiosis cohort 2 (BootstrapCell). Each dot correspond to the median predicted chronological age of an individual mouse. P-values at the mouse level obtained from the two-sided Wilcoxon rank-sum test. b, As in (a) but for the median predicted biological ages. c, As in (a) but for the exercise cohort. d, As in (a) but for the median predicted biological age in the exercise cohort.
Extended Data Fig. 8 Application of cell-type-specific aging clocks to exercise cohort.
a, UMAP projection of single-cell transcriptomes labeled by cell type from the exercise intervention cohort. b, Density plots to predict chronological age using chronological aging clocks (BootstrapCell) for young and old, exercise and control samples. c, Density plots to predict biological age using biological aging clocks (BoostrapCell) for young and old, exercise and control samples.
Extended Data Fig. 9 Comparison of genes in cell-type-specific aging clocks impacted by heterochronic parabiosis and by exercise.
a, Dot plot summarizing and comparing intervention effects across cell types. Effect sizes for parabiosis were determined by averaging cohort 1 and cohort 2. Exposure to young blood via heterochronic parabiosis has a stronger rejuvenation effect than exercise, and the impact is strongest in aNSC-NPCs. b, Pie charts indicating the overlap and directional effects of different interventions on genes selected by chronological aging clocks (BootstrapCell). Top: selected clock genes increase with age: Bottom: selected clock genes decrease with age. c, Barplots showing the proportion of genes that are differentially expressed age which are reversed by intervention, cell type, and whether the genes increase or decrease with age (abs(ln(fold change)) > 0.1, or approximately greater than a 1.1 fold change with age, FDR < 0.1). Parabiosis is effective at shifting differentially expressed genes during aging towards a more young-associated expression levels (more green in ‘Parabiosis’ column). Reduction of expression of genes that increase with age is larger than the induction of expression of genes that decrease with age (more green in ‘Age Increased’ rows).
Extended Data Fig. 10 Comparison of mean expression of genes in the aging clocks and genes impacted by rejuvenation in the heterochronic parabiosis and exercise datasets.
a, Scatter plots of the log2 mean normalized BootstrapCell gene expression in the exercise single-cell data as a function of the log2 mean normalized BootstrapCell gene expression in the parabiosis (cohorts 1 and 2 combined) single-cell data for the six main cell types. Red dots correspond to genes that contribute to the chronological aging clocks (selected by clock) and gray dots correspond to genes that do not contribute to the chronological aging clocks (not selected by clock). Genes that contribute to the clock are both highly expressed in the parabiosis and exercise datasets. b, As in (a) but colored dots correspond to genes identified as differentially expressed by parabiosis (green) and exercise (blue). Gray dots correspond to genes that are affected neither by parabiosis nor by exercise (neither). Permissive 1.1-fold change and FDR < 0.1 cutoffs were applied in each of the 2 different conditions.
Supplementary information
Supplementary Table 1
Mice metadata including age and multiplexing oligonucleotide sequence.
Supplementary Table 2
Gene markers for cell-type identification along with statistical significance and effect size measures from Seurat.
Supplementary Table 3
Prediction performance summary table for all models.
Supplementary Table 4
Clock genes and coefficients for all models.
Supplementary Software 1
Zip file containing a frozen version of the code used in this paper.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Buckley, M.T., Sun, E.D., George, B.M. et al. Cell-type-specific aging clocks to quantify aging and rejuvenation in neurogenic regions of the brain. Nat Aging 3, 121–137 (2023). https://doi.org/10.1038/s43587-022-00335-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s43587-022-00335-4
- Springer Nature America, Inc.
This article is cited by
-
Emerging role of senescent microglia in brain aging-related neurodegenerative diseases
Translational Neurodegeneration (2024)
-
Aging atlas reveals cell-type-specific effects of pro-longevity strategies
Nature Aging (2024)
-
Restoration of neuronal progenitors by partial reprogramming in the aged neurogenic niche
Nature Aging (2024)
-
Partial reprogramming of the mammalian brain
Nature Aging (2024)
-
TISSUE: uncertainty-calibrated prediction of single-cell spatial transcriptomics improves downstream analyses
Nature Methods (2024)