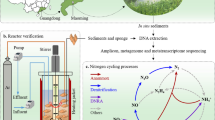
Study site, plastic preparation, and experimental design
In this study, the in situ incubations were conducted in an estuary of **amen, China (118°07’E, 24°61’N–118°09E’, 24°59’N) during April and May 2021. This region possesses a subtropical maritime monsoon climate, and has an average temperature of 21 °C and receives 1100 mm of rain68. Owing to human activities, pollutants in rainwater runoff and a portion of wastewater have been recently transported to the estuary, leading to reduced water quality and slight eutrophication. We sampled the estuarine water at 4 m depth and determined its characteristics. The water (pH 6.5, 20 °C) contained 1.19 mg L–1 dissolved oxygen, 21.2–25.7 mg L−1 total organic carbon, 0.72~0.99 mg L−1 nitrate, 0.19–0.43 mg L−1 ammonium, 0.09~0.17 mg L−1 nitrite, and 2.89~7.95 mg L−1 iron.
Four types of common commercial plastics, i.e., polyethylene (PE), polystyrene (PS), polypropylene (PP), and polyvinylchloride (PVC), were selected. They have a low bioreactivity with densities of 0.88–0.97 g cm−31. The PE-type food bags were from Cleanwrap Co., China. PS was obtained from transparent plastic cups (Chuanguan Co., China). PP was from sealable plastic bags from **ngmin Co., China. The PVC-type cling films were donated by Jusu Co., China. The four types of plastics were cut into a size of ~10 × 10 cm for subsequent experiments.
To investigate the denitrifying capability of the plastisphere, we performed (1) an in situ incubation for 30 days (30d), and (2) a subsequent denitrification experiment for 24 h (24 h). The detailed experimental design is illustrated in Supplementary Fig. 1. At the initial stage of in situ incubation, the obtained 10 × 10-cm plastic debris was sterilized using 70% ethanol and connected with cotton cords, and we used stone debris placed under the plastics to maintain buoyancy. This led to a ~4 m water depth for the plastics suspended in estuarine water. After 30 days, the plastic debris and surrounding bulk water (10 L) were harvested with a polymethyl-methacrylate sampler (Haifuda, China). A portion of plastics and bulk water was used for water quality analysis, SEM, and D2O-labeled single-cell Raman spectroscopy. The remaining material was used for the preparation of plastisphere suspensions and the subsequent 24 h experiment. Methods for water quality detections and SEM are listed in Supplementary Methods 1 and 2.
Prior to the denitrification experiment, a portion of bulk water was first filtered through a 0.22-μm polycarbonate membrane to remove microorganisms and then transferred into 250-mL sterile beakers. Thereafter, the collected plastic debris was placed in these beakers, and biofilms were repeatedly shaken, agitated, and washed with the sterile bulk water until a large proportion of the biofilms were detached69. Plastisphere suspensions were then collected for the determinations of denitrification rate, N2O reduction ratio (fR), EPS and c-di-GMP levels, and further experiments (Supplementary Fig. 1). During the 24 h denitrification experiment, we established plastisphere groups (i.e., plastisphere suspensions of PE, PS, PP, and PVC) and a bulk water group (estuarine water). Each plastisphere group was presented in triplicate, and the bulk water group (in quintuplicate) was set as control. Forty milliliters of plastisphere suspensions and bulk water were transferred into 120-mL serum bottles, respectively. Next, these bottles were sealed, purged with high-purity He (>99.99%) for 5 min, and incubated at 20 °C for 24 h. No additional carbon or nitrogen source was added, aiming to explore the basal capacities of denitrification in the plastisphere and bulk water. Each lab-scale experiment was conducted in parallel, and variations in the concentrations of organic matter during the lab-scale experiment are shown in Supplementary Fig. 10. After 24 h, N2O concentration and isotopocules, denitrifier community, and gene abundance in the plastisphere and bulk water were measured.
In this study, we prepared the plastisphere suspensions, aiming to explore the biofilms on the plastics under more controlled conditions during the lab-scale experiments. To examine if the growth pattern (attached or detached biofilms) has significant impacts on microorganisms during the 24 h experiment, we further investigated N2O production, anoxic microbial activities, and community structure of the attached and detached biofilms (Supplementary Fig. 11). These results showed no significant differences between attached and detached biofilms, suggesting that denitrifier activities during 24 h were not likely to be significantly impacted by growth patterns. In addition, we selected the water volume of 40 mL to conduct the 24 h experiment, because it is the minimum volume for a large proportion of biofilm detachment from plastic debris. We also conducted the same experiments with different volumes (80 and 120 mL) to avoid bias induced by incubation water volume (Supplementary Fig. 12). The results showed that N2O production in the plastisphere was higher than that in bulk water, regardless of water volumes.
In addition, it should be noted that marine particle debris is mainly in the form of plastics, occupying up to 60–80%70. Moreover, other particle fragments such as glass, metal, or stone, could be deposited to marine sediments, and the direct effects of these particles on marine waters are minor. Therefore, we only consider the plastic debris in this study.
EPS and c-di-GMP detections
After the 30-d incubation, the EPS and c-di-GMP concentrations in the plastisphere and bulk water were measured. Extractions of EPS were following the formaldehyde-NaOH method71. Plastisphere suspension (3 mL) or bulk water (3 mL) together with 2 mL sterile phosphate buffer were transferred into a 15-mL centrifuge tube. After centrifugation (5000×g, 4 °C, 5 min) and removal of the supernatant, 5 mL of sterile phosphate buffer and 50 μL of 37% formaldehyde were added to re-suspend the pellet. The tube was shaken for 5 min and stored at 4 °C for 60 min, and then 2 mL of 1 M NaOH was added. After being stored at 4 °C for 3 h, the supernatant was harvested by centrifugation (5000×g, 4 °C, 10 min) and used for PS, PN, and eDNA detections after filtering through a 0.22-μm polycarbonate membrane. PS concentration was measured at λ = 625 nm following the Anthrone method using glucose as standard. PN concentration was determined at λ = 562 nm by a Protein Assay Kit (Beyotime Biotechnology, China) using BSA as standard. eDNA concentration was quantified at λ = 260 nm by a DNA Detection Kit (QuantiFluor, dsDNA system, USA).
Microbial c-di-GMP in the plastisphere and bulk water were extracted and measured based on previous studies72,73. The extraction solvent consisted of acetonitrile/methanol (50/50, v/v). Cells in the plastisphere suspension and bulk water were disrupted using ultra-sonication for 5 min (120 W, 4 °C), and then transferred into a 15 mL centrifuge tube. After adding 8 mL extraction solvent and vortexing for 60 s, the suspensions were maintained at −80 °C for 2 h, and the supernatants were collected by centrifugation (14,000×g, 15 min). Thereafter, a pressured gas blowing concentrator (HGC-12A. Hengao, China) was used to dry the supernatants under continuous flows of N2. The residues were resuspended in 500 µL of ultrapure water for subsequent detection of c-di-GMP. The concentration of c-di-GMP was detected with an LC-MS/MS equipped with an electrospray ionization source (Applied Biosystems 6500, USA) at a reversed-phase. The m/z + 689/150 and + 689/343 quantifiers were c-di-GMP transitions. Declustering potential (DP) was −185 and collision energy (CE) was −45 and −48. The obtained peak areas were compared with a calibration curve generated with a c-di-GMP standard (Sigma-Aldrich, Germany). Concentrations of intracellular proteins were measured using a Protein Assay Kit (Beyotime Biotechnology, China).
Denitrification rate and N2O isotope measurement
After the 30-d incubation, denitrification rates of the plastisphere and bulk water were measured using the 15N isotope-pairing technique21,74. Five milliliters of plastisphere suspension or bulk water were prepared and transferred to a 12 mL gas-tight vial (Labco Exetainer, UK). The vials were purged with high-purity He (99.99%) for 5 min to replace the headspace air. The samples were first pre-incubated at 20 °C for 24 h to remove residual nitrate. After the removal, the vials were spiked with K15NO3 solution (99.8% atom, Macklin Co. China) to reach a concentration of 200 μM 15NO3– on the basis of in situ concentrations, and then re-purged with high-purity He. The samples were incubated in dark for 8 h at 20 °C. After the incubation, 2 mL of headspace gas was taken by a syringe and transferred to a new 12 mL vial pre-filled with He gas. The concentrations of 28N2, 29N2, and 30N2 in the headspace were quantified with a GasBench-II/isotope ratio mass spectrometry (Thermal Delta V Advantage, Germany). The detection limit is 0.1 μM. The denitrification rate was calculated as follows:
$${{{{{\rm{Denitrification}}}}}}\,{{{{{\rm{rate}}}}}}\,({{{{{{\rm{mg}}}}}}}^{15}\,N\,{L}^{-1}{h}^{-1})=\frac{{T}_{30}\times 2\times (1-{F}_{n})}{{F}_{n}}+2\times {T}_{30}$$
(1)
where T30 (mg 30N2 L−1 h−1) represents the production rates of 30N2 during the 8 h. Fn (99%) is the 15N fraction in added 15NO3− to residual 15NO3− after pre-incubation.
After the 24-h denitrification experiment, the headspace gas from the 120-mL serum bottle was taken to measure N2O concentrations and isotopic compositions. Three milliliters of headspace gas was used to analyze N2O concentration with a gas chromatograph (7890A, Agilent Technologies, USA), and 1 mL of headspace gas was used to detect N2O isotope compositions, including δ15Nbulk, δ15Nα, δ15Nβ, and δ18O, with a Precon+Gasbench/isotope ratio mass spectrometer (Delta V plus, Thermal, USA)75. Of these,15Nα (central,14N–15N–16O) and 15Nβ (terminal, 15N–14N–16O) are the isotopocules76. N2O gas in samples was first enriched in liquid N2 trap, purified, and then separated by a gas chromatography column (30-m long, 0.32-mm i.d.). Next, high-purity He flowing at 2 mL min−1 was applied to transport the N2O gas to the mass spectrometer. N2O isotopocules were detected by simultaneously capturing NO+ (m/z 30, 31) and N2O+ (m/z 44, 45, 46). The scrambling factor was determined as 0.085. In this study, the isotopocule values of the internal reference gas (high-purity N2O (>99.99%)) were pre-analyzed in Thünen Institute of Climate-Smart Agriculture, Germany. Two standard gases provided by Dr. Anette Goeske and Dr. Reinhard Well were applied to perform two-point calibrations for values of N2O site preference (N2O-SP, AK1 N2O-SP = −2‰, Mix 1 N2O-SP = 15‰, 5ppm). Values of δ15Nbulk, δ15Nα, δ15Nβ, δ18O, and SP of N2O were calculated as follow:77,78
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{\updelta }}}{\!\,}^{15}{{{{{{\rm{N}}}}}}}{\!\,}^{{{{{{\rm{i}}}}}}}(\permil )=({\!\,}^{15}{{{{{{\rm{N}}}}}}}_{{{{{{\rm{sample}}}}}}}{\!\,}^{{{{{{\rm{i}}}}}}}{-}{\!\,}^{15}{{{{{{\rm{N}}}}}}}_{{{{{{\rm{standard}}}}}}})/{\!\,}^{15}{{{{{{\rm{N}}}}}}}_{{{{{{\rm{standard}}}}}}}({{{{{\rm{i}}}}}}=\alpha \,{{{{{\rm{or}}}}}}\,\beta )$$
(2)
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{\updelta }}}{\!\,}^{18}{{{{{\rm{O}}}}}}(\permil )=({\!\,}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{sample}}}}}}}{{-}}{\!\,}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{standard}}}}}}}){/}{\!\,}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{standard}}}}}}}$$
(3)
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{\updelta }}}{\!\,}^{15}{{{{{{\rm{N}}}}}}}{\!\,}^{{{{{{\rm{bulk}}}}}}}(\permil )=({{{\updelta }}}{\!\,}^{15}{{{{{{\rm{N}}}}}}}{\!\,}^{{{\alpha }}}+{{{\updelta }}}{\!\,}^{15}{{{{{{\rm{N}}}}}}}^{{{\upbeta }}})/2$$
(4)
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}({{\permil}})={\updelta }^{15}{{{{{{\rm{N}}}}}}}^{{{\alpha }}}-{{{\updelta }}}^{15}{{{{{{\rm{N}}}}}}}^{{{\upbeta }}}$$
(5)
where δ18O and δ15Ni are the ratios of 18O/16O and 15N/14N, respectively. δ15Nα and δ15Nβ denote the ratios of 14N–15N–16O and 15N–14N–16O, respectively. All values were presented as ‰ relative to atmospheric N2 (15N) or Vienna Standard Mean Ocean Water (18O). The typical detection precisions are 0.3‰ (N2O-δ15Nbulk), 0.9‰ (N2O-δ15Nα), 0.9‰ (N2O-δ15Nβ) and 0.6‰ (N2O-δ18O).
Correction of N2O-o18O and N2O isotope mass balance model
During denitrification, the incorporation of the oxygen atom from H2O into N2O or nitrate could affect the measured δ18O values of N2O (N2O-δ18O)75,79,80. Previous studies conducted in soils suggest that an almost oxygen exchange occurred during denitrification79. Therefore, assuming a complete O exchange δ18O values of the ambient water (H2O-δ18O) could be used for interpretation. As the variation of H2O-δ18O values in global surface seawater is minor81, we used a theoretical H2O-δ18O value (0.91‰, between −2.98 and 1‰)81,82 reported in the literature for seawater to correct the measured N2O-δ18O values (subtracting H2O-δ18O value).
After correction, the multiple-isotope relationships between N2O-SP and N2O-δ18O were plotted based on the map** approach83, which was first applied by Lewicka-Szczebak et al. Then, we further established a N2O isotope mass balance28 to explore the relative fractions of N2O-producing sources in the plastisphere and bulk water. As the denitrification experiment was anaerobically executed, N2O productions via BD, FD, and CD processes were taken into account. Fractions of BD (fBD), FD (fFD), and CD (fCD) pathways to total N2O flux were expressed as follow:
$${{{{{\rm{Total}}}}}}\,{{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}\,{{{{{\rm{production}}}}}}={{{{{{\rm{f}}}}}}}_{{{{{{\rm{BD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{FD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{CD}}}}}}}=1$$
(6)
For N2O-SP values, N2O isotope mass contributions of BD, FD, and CD processes to the total N2O-SPNR values production were expressed as:
$${{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{NR}}}}}}}={{{{{{\rm{f}}}}}}}_{{{{{{\rm{BD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{BD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{FD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{FD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{CD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{CD}}}}}}}$$
(7)
where N2O-SPNR refers to the N2O-SP value before reduction. N2O-SPBD, N2O-SPFD, and N2O-SPCD are the N2O-SP values of BD (−7.5–3.7‰)65,76,84, FD (27.2–39.9‰)75,76,85,86,87, and CD (10–22‰)29,32,76,88, respectively. Similarly, the contributions of the three endmembers to N2O-δ18O values of the three processes were expressed as:
$${{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{NR}}}}}}}={{{{{{\rm{f}}}}}}}_{{{{{{\rm{BD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{BD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{FD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{FD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{CD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{CD}}}}}}}$$
(8)
where N2O-δ18ONR refers to the N2O-δ18O value before reduction. N2O-δ18OBD, N2O-δ18OFD, and N2O-δ18OCD are the N2O-δ18O values of BD (16.4–23.3‰)65,76,84, FD (42.0–55.1‰)75,76,85,86,87, and CD (24.9–37.6‰)29,32,76,88, respectively.
Nitrogen and oxygen isotope fractionations during N2O reduction such as BD can increase the measured N2O-SP and N2O-δ18O values of remaining N2O. Fractionation-derived shifts in N2O-SP and N2O-δ18O values during N2O reduction were thus estimated according to a previous study and presented as:
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}={{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{NR}}}}}}}{-{{{{{\rm{f}}}}}}}_{{{{{{\rm{R}}}}}}}\times {{{{{{\rm{SP}}}}}}}_{{{{{{\rm{\varepsilon }}}}}}{{{{{\rm{N2O}}}}}}}$$
(9)
$${{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{\rm{O}}}}}}={{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{NR}}}}}}}{-{{{{{\rm{f}}}}}}}_{{{{{{\rm{R}}}}}}}\times {{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{\varepsilon }}}}}}{{{{{\rm{N2O}}}}}}}$$
(10)
where SP and δ18O are the N2O-SP and N2O-δ18O values after N2O reduction, respectively, which are the measured SP and δ18O values here; fR denotes the N2O reduction ratio (0.75 in this study). This was estimated using the C2H2 approach41, i.e., the proportion of the residual (without C2H2; residual N2O after N2O reduction) to total (with C2H2; residual N2O after N2O reduction) N2O concentrations. SPεN2O (−6‰) and δ18OεN2O (−25‰) were assumed to be the fractionation effect during N2O reduction76,89. Given the N2O reduction effect during experiments, Eqs. (7) and (8) were modified as:
$${{{{{{\rm{N}}}}}}}_{2}{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}={{{{{{\rm{f}}}}}}}_{{{{{{\rm{BD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{BD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{FD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{FD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{CD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{SP}}}}}}}_{{{{{{\rm{CD}}}}}}}{-{{{{{\rm{f}}}}}}}_{{{{{{\rm{R}}}}}}}\times {{{{{{\rm{SP}}}}}}}_{{{{{{\rm{\varepsilon }}}}}}{{{{{\rm{N2O}}}}}}}$$
(11)
$${{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{\rm{O}}}}}}={{{{{{\rm{f}}}}}}}_{{{{{{\rm{BD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{BD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{FD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{FD}}}}}}}+{{{{{{\rm{f}}}}}}}_{{{{{{\rm{CD}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{CD}}}}}}}{-{{{{{\rm{f}}}}}}}_{{{{{{\rm{R}}}}}}}\times {{{{{{\rm{N}}}}}}}_{2}{{{{{{\rm{O}}}}}}-{{{{{\rm{\delta }}}}}}}^{18}{{{{{{\rm{O}}}}}}}_{{{{{{\rm{\varepsilon }}}}}}{{{{{\rm{N2O}}}}}}}$$
(12)
We further applied the Gaussian estimation and Monte Carlo sampling method to obtain the mean values and standard errors of N2O-SP and N2O-δ18O for each endmember, and assess the errors of the mass balance model. The detailed MATLAB codes and Monte Carlo stimulations (10,000 sampling) are provided in Supplementary Table 2 and Supplementary Fig. 13. The values of SP and δ18O-(N2O/H2O) used in the model for BD were −1.5‰ and 19‰, respectively; for FD were 37‰ and 47‰; for CD were 16‰ and 30‰. Combining Eqs. (6), (11), and (12), the relative contributions of BD, FD, and CD to N2O productions are acquired in the plastisphere and bulk water. Proportions estimated by the equations are considered as zero if yielding negative values, and the other two proportions sum up to 100% accordingly.
D2O-labeled single-cell Raman spectroscopy
To compare microbial phenotypic activity and to explore intracellular lipid/fatty acid levels between plastisphere and bulk water, a D2O-labeled single-cell Raman spectroscopy experiment was performed15,90. After the 30-d incubation, the four types of plastics were cut to produce ~1.5 × 1.5 cm plastic fragments. For the plastisphere, 1 mL of sterilized bulk water and the plastic fragments were transferred to a 12-well plate and mixed with 1 mL D2O, finally reaching a concentration of 50% D2O. For bulk water, 1 mL of in situ bulk water and 1 mL D2O were added to another 12-well plate. Each group was conducted in triplicate. The 12-well plates were anaerobically incubated at 300 rpm and 20 °C for 24 h. Next, plastic fragments were transferred to 5 mL phosphate buffer with 0.5% Tween-20, and attached microorganisms were separated by vortexing for 10 min. After washing with sterilized water twice (4000×g, 5 min), 2 μL of microorganisms from bulk water and plastisphere were spotted on a tin foil-coated microslide and dried at 25 °C. Raman spectroscopy was acquired with a LabRAM Aramis (HORIBA Jobin-Yvon) confocal micro-Raman system, including a 300-g/mm diffraction grating, a 532-nm Nd:YAG laser, and a ×100 objective (Olympus, Japan). We used the Raman characteristic band of a silicon wafer (520.6 cm−1) to calibrate the Raman spectrometer. The generated spectra were subsequently analyzed in LabSpec-5 software (HORIBA Jobin-Yvon) with baseline correction and normalization. Peak intensities of Raman bands assigned to C–D (2040–2300 cm−1) and C–H (2800–3100 cm−1) were obtained to estimate (C–D)/(C–D + C–H) ratios42.
Denitrifier community
After the 24 h denitrification experiment, 2 mL of plastisphere suspension was taken for DNA extraction with a FastDNA Kit (MP, SA, USA). In situ bulk water samples were first filtered through sterilized 0.22μm polycarbonate filters that were then used for DNA extraction15. Bacterial nirS gene (bnirS-type denitrifier), bacterial nirK gene (bnirK-type denitrifier), and Internally Transcribed Spacer (ITS) gene (fungal community) were used for amplicon sequencing with the primer sets (Supplementary Table 3). The obtained raw data were quality-filtered, chimera checked and then grouped at 97% similarity. Taxonomy was compared with GeneBank and UNITE databases using QIIME. Information of fungal nirK denitrifiers (fnirK-type denitrifiers) was screened from fungal communities based on previous studies and NR database36,91. The coverage of screened operational taxonomic unit (OTU) to total OTUs of ITS data was >85%, which was used for subsequent analysis. The α-diversities including abundance-based coverage estimation (ACE), Chao1, Shannon, and Simpson indexes, were estimated in R. Principal coordinates analysis coupled with analysis of similarities (ANOSIM) test was applied to explore β-diversities of the three types of denitrifier communities. Co-occurrence network analysis was performed in R92 (nodes R2 > 0.90 and P < 0.05) and visualized by Gephi (Version 0.9.2). Sequences will be submitted under accession number SUB10278422.
We investigated denitrifiers-based niche breadth and overlap according to Levins’ niche breadth method and asymmetric α-model in R14,46,47. These two indicators reflect the potential of denitrifiers adapting to environments and competing for resources in the plastisphere and bulk water. The formulas were as follow:
$${{{{{{\rm{B}}}}}}}_{{{{{{\rm{i}}}}}}}=\frac{1}{\mathop{\sum}\nolimits_{{{{{{\rm{i}}}}}}=1}^{{N}}\times {{{{{{\rm{P}}}}}}}_{{{{{{\rm{ij}}}}}}}^{2}}$$
(13)
and
$${{{{{{\rm{\alpha }}}}}}}_{{{{{{\rm{ij}}}}}}}=\frac{\mathop{\sum}\nolimits_{{{{{{\rm{i}}}}}}=1}^{N}\times {{{{{{\rm{P}}}}}}}_{{{{{{\rm{i}}}}}}{{{{{\rm{\alpha }}}}}}}\times {{{{{{\rm{P}}}}}}}_{{{{{{\rm{j}}}}}}{{{{{\rm{\alpha }}}}}}}}{\mathop{\sum}\nolimits_{{{{{{\rm{i}}}}}}=1}^{{N}}\times {{{{{{\rm{P}}}}}}}_{{{{{{\rm{i}}}}}}{{{{{\rm{\alpha }}}}}}}^{2}}$$
(14)
where Bi and αij are the niche breadth and overlap of denitrifier i in the plastisphere or bulk water, respectively; N denotes the total number of denitrifiers in each type of denitrifier community; Pij denotes the proportion of denitrifier i in communities.
Gene abundance
The abundances of bacteria (16S rRNA) and fungi (ITS) in the plastisphere and bulk water were quantified with qPCR using the universal primers 515F/907R and ITS1F/ITS2R, respectively68,92. The standard curves were obtained by a serial dilution of the plasmids with 16S rRNA and ITS genes. The mixing system (20 μL) included 2 μL of DNA sample, 10 μL of Mix-enzyme (LightCycler SYBR Green I), 0.8 μL of 515 F/907 R or ITS1F/ITS2R primers, and 7.2 μL of H2O. The thermal cycling conditions of 16 S rRNA: 95 °C (3 min), 40 cycles at 95 °C (30 s), 55 °C (30 s) and 72 °C (45 s), and 72 °C (10 min). The thermal cycling conditions of ITS: 94 °C (10 min), 40 cycles at 94 °C (30 s), 56 °C (30 s) and 72 °C (45 s), and 72 °C (7 min). Samples with sterile pure water were regarded as negative control. Amplification efficiencies were 106.8–115.5% with R2 > 0.995. Based on the Ribosomal RNA Operon Copy Number Database, microbial cell numbers could be assessed by the absolute abundances of 16S rRNA/4.193,94.
High-throughput qPCR-based SmartChip technique95 was applied to detect denitrifying genes (narG, nirS, nirK, nosZ) abundances in the plastisphere and bulk water. The mixing system included 3.1 μL of DNA sample, 24.8 μL of Mix-enzyme (LightCycler SYBR Green I), and 3.1 μL of denitrifying gene primers. They were dispensed into a SmartChip via a MultiSample NanoDispenser, and then quantified with a Real-Time PCR system (WaferGen, Biosystems, USA). The bacterial 16 S rRNA (515 F/907 R) gene was set as a reference. The thermal cycle of qPCR conditions included heating at 95 °C (10 min), followed by 40 cycles at 95 °C (30 s), annealing at 58 °C (30 s), and extension at 72 °C (30 s). The threshold cycle of 31 was regarded as the detection limit. Information of primer pairs and other procedures are listed in our previous study95 and Supplementary Table 4. Relative copy numbers of denitrifying gene and16S rRNA gene were estimated as follows:
$${{{{{{\rm{Relative}}}}}}\;{{{{{\rm{gene}}}}}}\;{{{{{\rm{copy}}}}}}\;{{{{{\rm{numbers}}}}}}=10}^{(31-{{{{{\rm{threshold}}}}}}\;{{{{{\rm{cycle}}}}}})}/(3.33)$$
(15)
Normalized relative denitrifying gene abundance was the proportion of the relative functional gene copy number to the 16S rRNA gene copy number.
Statistical analysis
In this study, the significance analyses on raw data were performed in IBM SPSS (Version 22.0). The significant differences in microbial activity, EPS concentration, c-di-GMP, and lipid/fatty acid level, N2O concentration, denitrification rate, the proportion of denitrification pathways to N2O production, functional gene abundance, denitrifiers-based niche breadth and overlap between plastisphere and bulk water were tested with one-way ANOVA method followed by a post-hoc Tukey test. P value < 0.05 was considered as significant.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.