Abstract
The MIR663AHG gene encodes both miR663AHG and miR663a. While miR663a contributes to the defense of host cells against inflammation and inhibits colon cancer development, the biological function of lncRNA miR663AHG has not been previously reported. In this study, the subcellular localization of lncRNA miR663AHG was determined by RNA-FISH. miR663AHG and miR663a were measured by qRT-PCR. The effects of miR663AHG on the growth and metastasis of colon cancer cells were investigated in vitro and in vivo. CRISPR/Cas9, RNA pulldown, and other biological assays were used to explore the underlying mechanism of miR663AHG. We found that miR663AHG was mainly distributed in the nucleus of Caco2 and HCT116 cells and the cytoplasm of SW480 cells. The expression level of miR663AHG was positively correlated with the level of miR663a (r = 0.179, P = 0.015) and significantly downregulated in colon cancer tissues relative to paired normal tissues from 119 patients (P < 0.008). Colon cancers with low miR663AHG expression were associated with advanced pTNM stage (P = 0.021), lymph metastasis (P = 0.041), and shorter overall survival (hazard ratio = 2.026; P = 0.021). Experimentally, miR663AHG inhibited colon cancer cell proliferation, migration, and invasion. The growth of xenografts from RKO cells overexpressing miR663AHG was slower than that of xenografts from vector control cells in BALB/c nude mice (P = 0.007). Interestingly, either RNA-interfering or resveratrol-inducing expression changes of miR663AHG or miR663a can trigger negative feedback regulation of transcription of the MIR663AHG gene. Mechanistically, miR663AHG could bind to miR663a and its precursor pre-miR663a, and prevent the degradation of miR663a target mRNAs. Disruption of the negative feedback by knockout of the MIR663AHG promoter, exon-1, and pri-miR663A-coding sequence entirely blocked these effects of miR663AHG, which was restored in cells transfected with miR663a expression vector in rescue experiment. In conclusion, miR663AHG functions as a tumor suppressor that inhibits the development of colon cancer through its cis-binding to miR663a/pre-miR663a. The cross talk between miR663AHG and miR663a expression may play dominant roles in maintaining the functions of miR663AHG in colon cancer development.
Similar content being viewed by others
Introduction
Colon cancer is a malignant disease with a mortality rate second to that of lung cancer worldwide. In 2022, it is estimated that there will be ~100,000 newly diagnosed colon cancer patients in the United States, and the mortality rate will reach ~8.6% [1]. Although the popularization of DNA methylation marker screening, colonoscopy examination, and resection of colorectal mucosal polyps have made considerable progress in early diagnosis and prevention, the incidence of colon cancer in people under 50 years old is still steadily increasing [1]. In-depth investigations of the mechanisms may hasten elucidation of the development of colon cancer.
With the development of high throughput RNA sequencing (RNA-seq) technology, it has been found that there are a large number of noncoding RNA (ncRNA) molecules transcribed from the human genome [2]. Long noncoding RNAs (lncRNAs) may regulate chromatin conformation, gene transcription, hnRNA splicing, and other biological processes such as cancer development [3,4,5,6,7,8]. The expression status of many lncRNAs is abnormal in cancer tissues. Some factors directly affect cancer cell proliferation, apoptosis, metastasis, angiogenesis, drug resistance, and stem cell stemness [9,10,11,12]. In addition, ncRNAs have tissue-specific and tumor-specific expression patterns and are expected to be potential screening, diagnostic, and prognostic biomarkers for colorectal cancer in stool, serum/plasma, and tissue samples of patients [13,14,15].
MIR663AHG is a rare human-specific gene (Entrez Gene: 284801) located in the centromere of chromosome 20. The primary transcript (hnRNA) of this gene can be simultaneously spliced into microRNA miR663a and lncRNA miR663AHG. We have previously reported that miR663a expression is downregulated in colon cancer tissues and that miR663a inhibits the growth and metastasis of colon cancer cells [7, 16]. A correlation investigation suggests that changes in miR663AHG expression are associated with the occurrence of intervertebral disc degeneration (IDD) [17]. Another study revealed that miR663AHG is downregulated in psoriatic tissues [18]. According to the Genotype-Tissue Expression Project (GTEx) project database [19], miR663AHG is ubiquitously expressed in various human normal tissues with relative abundance (Fig. S1). However, the biological functions of miR663AHG have not been previously reported. In this study, we studied, for the first time, the roles of miR663AHG in colon cancer development and its molecular mechanisms.
Results
Characterization of miR663AHG and its subcellular distribution
Compared with protein-coding genes, the transcription start sites (TSSs) of ncRNA genes are more diverse [20, 21]. MIR663AHG is a multiexon gene. miR663a and miR663AHG are spliced from the same hnRNA. The values of miR663AHG were less than zero in PhyloCSF analysis (Fig. 1A), suggesting that this lncRNA does not encode a protein [22].
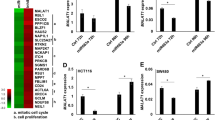
A The prediction of the protein-coding ability of miR663AHG, as analyzed with the PhyloCSF online tool. The genomic locations of miR663AHG exons, miR663a, and CpG islands in human chromosome 20 are also illustrated using the adapted images downloaded from the UCSC website. B Comparisons of the levels of miR663AHG and miR663a in cancer cell lines subclassified into MIR663AHG DNA methylation-high, -moderate, and -low groups according to the CCLE databases. */***: P < 0.05/0.001 and NS: P > 0.05 in Mann–Whitney U-test. C The baseline level of miR663AHG expression in six colon cancer cell lines, as determined by qRT-PCR. D Subcellular distribution of endogenous miR663AHG in various colon cancer cells in the RNA-FISH analysis; 18 S rRNA and U6 were used as cytoplasmic and nuclear RNA controls, respectively.
It is well-known that gene transcription is regulated by the methylation status of CpG islands around transcription start sites (TSSs). A CpG island is located around the MIR663AHG TSS (Fig. 1A). According to the results of mining CCLE DNA methylation and RNA-seq databases [23], the levels of miR663AHG and miR663a were significantly lower in cell lines with high methylation levels than in those with moderate or low methylation levels (P < 0.05; Fig. 1B). This suggests that the transcription of the MIR663AHG gene is regulated by DNA methylation.
In eukaryotic cells, the functions of lncRNAs are closely related to their subcellular localization. To characterize the distribution pattern of miR663AHG in colon cancer cells, its baseline expression status in six colon cancer cell lines was initially determined by qRT-PCR. The level of miR663AHG was higher in Caco2 or SW620 cells than in HCT116 or RKO cells and lowest in SW480 and LoVo cells (Fig. 1C). Similarly, the level of miR663a was highest in SW620 cells and lowest in SW480 cells (Data not shown). The results of RNA-FISH analysis showed that miR663AHG was located in both the cytoplasm and nucleus of colon cancer cells: mainly in the nucleus of Caco2 and HCT116 cells and in the cytoplasm of SW480 cells (Fig. 1D). No miR663AHG hybridization signal could be detected in LoVo cells.
Downregulation of miR663AHG in colon cancer is associated with poor prognosis
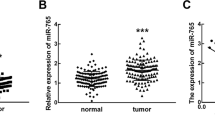
We previously reported that miR663a expression was significantly downregulated in colon cancer tissues (CCs) relative to matched surgical margin controls (SMs) from patients [7]. Patients (n = 119) for whom enough RNA samples were available for miR663AHG detection were re-enrolled in the present study. The results of the qRT-PCR analysis showed that miR663AHG expression was significantly downregulated in CCs compared with SMs, especially in CCs with lymph metastasis or at advanced pTNM stages or with good differentiation (Fig. 2A, B; Table S1).
A The level of miR663AHG expression in colon cancer (CC) and paired surgical margin (SM) samples from 119 patients, as measured by qRT-PCR. The P-value in the Mann–Whitney U test and the ratio of CCs with miR663AHG downregulation are labeled. B Comparisons of the level of miR663AHG expression in CCs at different pTNM stages, lymph metastasis, and differentiation states. Detailed information is listed in Table S1. C Overall survival curves for patients with colon cancer with high and low miR663AHG expression (according to the median value). D The coexpression status of miR663AHG and miR663a in 182 (CC and SM) tissue samples. E The coexpression status of miR663AHG and miR663a in 353 ovarian cancer tissue samples. The Spearman correlation coefficient (r) is labeled.
In the Kaplan-Meier analysis, the overall survival (OS) of patients with colon cancer with low miR663AHG expression (below the median) was significantly shorter than that of patients with high miR663AHG expression (P = 0.018; Fig. 2C). Univariate analysis showed that four risk factors were significantly associated with the OS of these patients (Table S3). These factors included low miR663 AHG expression (hazard ratio [HR], 2.026; 95% confidence interval [CI], 1.113–3.689), pTNM stage, lymph metastasis, and distant metastasis. In multivariate analysis, however, only the metastasis status, but not the miR663AHG level, was significantly associated with patients’ OS.
In addition, combined with our previously reported miR663a data [7], we analyzed the correlation between the levels of miR663AHG and miR663a in these colon tissues (n = 182; including CCs and SMs). A positive correlation was observed (r = 0.179, P = 0.015), implying a possible role of miR663a in the biological function of miR663AHG (Fig. 2D). Furthermore, we also analyzed the correlation using public microarray datasets [24] and found a similar correlation among 353 ovarian cancer samples (Spearman r = 0.168, P = 0.002; Fig. 2E).
miR663AHG inhibits the growth and metastasis of colon cancer cells in vitro and in vivo
To investigate the biological functions of miR663AHG, two human colon cancer cell lines, SW480 and LoVo, with a low baseline level of miR663AHG expression, were stably transfected with the lentiviral miR663AHG expression vector, while the other two cell lines SW620 and HCT116, with a high baseline level of miR663AHG expression, were transfected with siRNAs against miR663AHG (si663AHG). The proliferation of SW480 and LoVo cells was significantly inhibited by miR663AHG overexpression, whereas the proliferation of SW620 and HCT116 cells was significantly promoted by miR663AHG knockdown in the long-term dynamic observation analysis (Fig. 3A, B). Similar differences in the colony formation rate were also induced by both miR663AHG overexpression and knockdown (Fig. 3C, D). These results were confirmed in the animal model. The growth of xenografts derived from RKO cells stably overexpressing miR663AHG was significantly slower than that of empty control cells (P = 0.009; Fig. 3E).
A, B Impact of miR663AHG overexpression or knockdown on the proliferation of colon cancer cells, as measured by long-term dynamic observation. The transfection efficiency of the miR663AHG expression vector was monitored by examination of the level of ZsGreen expression 48 h post-transfection. The level of miR663AHG is presented as the mean ± SD. C, D Effect of miR663AHG overexpression or knockdown on the colony formation of colon cancer cells. E Images and tumor weights of subcutaneous xenografts derived from miR663AHG stably overexpressing RKO cells in BALB/c nude mice. */**/***: P < 0.05/0.01/0.001 for (A–D) by Student’s t-test, for (E) by the Mann–Whitney U-test.
Furthermore, the results of wound healing assays revealed that the migration of SW480 and LoVo cells was impeded by stable miR663AHG overexpression (Fig. 4A), while that of SW620 and HCT116 cells was enhanced by miR663AHG knockdown (Fig. 4B). The results of transwell assays also showed that miR663AHG overexpression or knockdown significantly decreased or increased the migration and invasion of these cells, respectively (Fig. 4C, D). Although the difference in the average number of lung surface metastatic nodules was not statistically significant (Fig. 4E left, P = 0.286; Fig. S3), a significant decrease in the weight of lungs was observed between NOD-SCID mice injected with LoVo cells with and without stable miR663AHG overexpression (P = 0.004; Fig. 4E right).
A, B Effects of miR663AHG overexpression or siRNA knockdown on the migration of colon cancer cells in wound-healing analysis. C, D Effects of miR663AHG overexpression or siRNA knockdown on the migration and invasion of colon cancer cells in transwell analyses. (E) Effect of miR663AHG overexpression on the experimental lung metastasis of LoVo cells in NOD/SCID mice. Both the average number of metastatic lesions and lung weight are displayed. */**/***: P < 0.05/0.01/0.001 for (A–D) by Student’s t-test, for (E) by the Mann–Whitney U-test.
Collectively, these results indicate that miR663AHG suppresses colon cancer cell proliferation, migration, and invasion in vitro and suppresses tumor formation and lung metastasis in vivo.
Negative feedback of the MIR663AHG gene by its miR663AHG and miR663a products
The cis-regulation between miRNAs and their host genes provides a novel layer for ncRNA-mediated gene regulation [25,26,27]. qRT-PCR analysis revealed that miR663AHG overexpression or knockdown significantly reversed endogenous miR663a expression in various colon cancer cell lines (Fig. 5A, B). Similarly, transfection of miR663a mimic or antisense/inhibitor also reversed the level of endogenous miR663AHG in these cell lines (Fig. 5C, D).
A Downregulation of miR663a by transient miR663AHG overexpression. B Upregulation of miR663a by siRNA knockdown of miR663AHG. C, D Downregulation or upregulation of miR663AHG by transfection of miR663a mimic or antisense/inhibitor. E, F Effect of resveratrol treatment on miR663AHG and miR663a expression in colon cancer cells. */**/***: P < 0.05/0.01/0.001 by Student’s t-test.
miR663a is a well-recognized miRNA involved in host responses to stress and inflammatory factors, including reactive oxygen species (ROS) [28]. Resveratrol (RSV) is a well-known antioxidant. RSV treatment increases the level of miR663a in various cells [29, 30]. Here, we found that RSV treatment (final concentration 6.25 μM for 12 h) not only significantly increased the miR663a level but also decreased the miR663AHG level in various colon cancer cell lines (Fig. 5E, F). These phenomena suggest that expression changes of miR663AHG or miR663a, whether induced by transfection or antioxidant treatment, could result in negative expression feedback of the MIR663AHG gene.
To evaluate the importance of the negative feedback in miR663AHG function, we knocked out the 751-bp genomic sequence from the MIR663AHG promoter to the pri-miR663a-coding fragment in RKO cells with CRISPR/Cas9 (MIR663A/HG-KO; Fig. S2A–C). The level of miRNA663AHG expression was decreased by ~80% in MIR663A/HG-KO cells relative to wild-type control cells (MIR663A/HG-WT; Fig. S2D). While enforced miR663AHG expression changes significantly enhanced or inhibited the proliferation of MIR663A/HG-WT RKO cells, such effects could not be observed in MIR663A/HG-KO cells in the long observation (Fig. 6A). In the rescue experiment, the inhibitory effect of miR663AHG on the proliferation was restored in MIR663A/HG-KO cells transfected with the miR663a expression vector (Fig. 6B). Similarly, miR663AHG knockdown significantly enhanced the migration and invasion of MIR663A/HG-WT cells but did not affect those of MIR663A/HG-KO cells in the wound healing, transwell migration and invasion analyses (Fig. 6C, D). No effect of miR663AHG overexpression on the migration and invasion of MIR663A/HG-KO cells was observed. These results demonstrate that these effects of miR663AHG are dependent on the negative feedback of the MIR663AHG gene and consequent changes in miR663a expression.
A The proliferation curves of MIR663A-WT and MIR663A/HG-KO cells with and without miR663AHG overexpression or knockdown in long-term live cell dynamic observation. B Effect of miR663AHG overexpression on the proliferation of MIR663A/HG-KO cells with and without miR663a expression in the rescue experiment. C Effect of miR663AHG knockdown on the migration of MIR663A-WT and MIR663A/HG-KO cells, as determined with a wound healing assay. D Effect of miR663AHG overexpression and knockdown on the migration and invasion of MIR663A-WT and MIR663A/HG-KO cells, as determined with Transwell assays (n = 5). */**/***: P < 0.05/0.01/0.001 and NS, P > 0.05 by Student’s t-test.
miR663AHG binds to miR663a and its precursors and protects miR663a targets
To explore the potential underlying mechanism of the inhibitory roles of miR663AHG in colon cancer development, we performed bioinformatics analysis using the StarBase v3.0 database. We found that miR663AHG might directly bind in cis to the miR663a precursors pre-miR663a (and pri-miR663a) (Fig. S4). As expected, the results of RNA pulldown analysis showed that biotin-labeled miR663AHG indeed bound to pre-miR663a (and pri-miR663a) in RKO cells and that biotin-labeled pre-miR663a also bound to miR663AHG (Fig. 7A). In addition, miR663AHG was also significantly enriched by biotin-labeled miR663a in RKO cells (Fig. 7B). These results confirmed that miR663AHG could directly bind to miR663a and its precursors.
A PAGE gel images of RNA pulldown-PCR analysis to detect miR663AHG cis-binding to pre- and pri-miR663a. Starbase v3.0-predicted sequences for miR663AHG-pre-miR663a cis-binding are illustrated on the top. Blue letters represent the sequence of mature miR663a. B The results of RNA pulldown-qRT-PCR analysis to detect the enrichment status of miR663AHG by biotin-labeled miR663a in RKO cells. Starbase v3.0-predicted sequences for miR663AHG-miR663a cis-binding are included under the chart. C Effect of transient miR663AHG overexpression on the level of pre-miR663a expression in colon cancer cells. D Effects of miR663AHG overexpression on the mRNA levels of miR663a target genes in colon cancer cells, with wild-type (WT) or knockout (KO) of MIR663A/HG alleles. */**/***: P < 0.05/0.01/0.001, NS: P > 0.05 by Student’s t-test.
To understand the significance of miR663AHG-pre-miR663a and miR663AHG-miR663a cis-binding, we further analyzed the effects of miR663AHG overexpression on the pre-miR663a level in colon cancer SW480 and LoVo cells. Similar to miR663a (Fig. 5A), the level of pre-miR663a was significantly decreased by miR663AHG overexpression in these cells (Fig. 7C). These results indicate that miR663AHG-pre-miR663a cis-binding may contribute to expression changes of miR663a and its precursor.
In addition, we also determined the expression changes of a set of miR663a targets. We found that the mRNA levels of the TP53, JUNB, JUND, PIK3CD, P21, and TGFB1 genes were significantly elevated by miR663AHG overexpression in SW480, LoVo, and MIR663A/HG-WT RKO cells. However, miR663AHG overexpression did not affect the mRNA levels of these targets in MIR663A/HG-KO RKO cells (Fig. 7D).
Discussion
Primary transcripts of the MIR663AHG gene can be spliced into both miR663a and miR663AHG. While miR663a contributes to host defense responses to inflammatory factors such as reactive oxygen species (ROS) in cancer development [7, 16, 31,32,33,34,35], the biological function of miR663AHG has not been previously reported. In this study, we reported for the first time that miR663AHG inhibits the development and progression of human colon cancer in a miR663a-dependent manner.
According to the GTEx databases for 570 donors [36], three main miR663AHG isoforms (ENST00000608521, ENST00000608487, and ENST00000601079) are prevalently expressed in various human normal tissues with a similar expression pattern, and some rare miR663AHG isoforms are specifically expressed in the normal testis (Data file 1). Most miR663AHG isoforms are significantly downregulated in testicular germ cell tumors (TGCT) and acute myeloid leukemia (LAML), according to the results of The Gene Expression Profiling Interactive Analysis (GEPIA2) using Cancer Genome Atlas (TCGA) RNA-seq datasets (Figure S5) [37, 38]. Using a sensitive and quantitative transcript-specific RT-PCR assay, we found that miR663AHG (ENST00000608521) was significantly downregulated in colon cancer tissues and associated with colon cancer metastasis. Our in vitro and in vivo experiments consistently support the hypothesis that miR663AHG functions as a tumor suppressor. The roles of other miR663AHG isoforms in cancer development are worthy of further study.
The functions of most biological macromolecules depend on their subcellular distributions. In our RNA-FISH experiment, a set of fluorescently labeled RNA probes targeting various miR663AHG isoforms were used. We found that miR663AHG was distributed in the cytoplasm and nucleus of colon cancer cell lines. No miR663AHG signal was detected in LoVo cells with a low, if any, baseline level of miR663AHG expression, indicating high accuracy of the RNA-FISH assay. However, cytoplasmic miR663AHG signals were detected in SW480 cells with a low level of baseline miR663AHG expression, suggesting the possibility of other miR663AHG isoforms in these cells, which may not be covered by the miR663AHG/ENST00000608521-specific RT-PCR primer set used in the present study. To confirm this, we mined CCLE RNA-seq datasets and found that the expression levels of two miR663AHG isoforms ENST00000615682 and ENST00000665857, which are most frequently expressed (~10%) in cancer cell lines (n = 1406), are detectable in SW480 cells (Data file 2) [23]. The RT-PCR primer set indeed does not cover ENST00000615682 and ENST00000665857. This may account for the difference in the expression level and subcellular location of miR663AHG between SW480 and HCT116 or Caco2 cells.
It is well recognized that pre-miRNAs are nuclear-cytoplasmic transportation forms, and mature miRNAs are mainly distributed in the cytoplasm. That miR663AHG is located in both the nucleus and cytoplasm is consistent with the observation of miR663AHG cis-binding to both pre-miR663a and miR663a. Interestingly, we found that miR663AHG could bind to pri-miR663a, suggesting an additional regulatory machine in the nucleus. miR663AHG may regulate the functions of miR663a at multiple levels.
Our and others’ works indicate that miR663a inhibits the development and metastasis of colon cancer and that the miR663a target genes MALAT1 and TTC22 promote colon cancer metastasis [7, 16, 39, 40]. Here, we further found that miR663AHG could inhibit the proliferation, migration/invasion of colon cancer cells in vitro and the growth and lung metastasis of colon cancer cells in vivo.
As described above, while miR663AHG prevents miR663a targets from degradation, both miR663AHG and miR663a inhibit the development and metastasis of colon cancer. In addition, the expression levels of miR663AHG and miR663a are positively correlated with each other not only in colon tissues from cancer patients but also in ovarian cancer tissues. Notably, disruption of the negative feedback of the MIR663AHG gene entirely abolished the effects of miR663AHG on the biological behaviors of cancer cells. These phenomena suggest that the functions of endogenous miR663AHG and miR663a may be intrinsically connected: they form the miR663AHG-miR663a axis and play the same role in maintaining homeostasis of defense responses of host cells to inflammatory factors and cancer development.
Mechanistically, we found that alterations of miR663AHG (or miR663a) expression, whether induced by expression vectors or the antioxidant resveratrol treatment, could reverse alterations of endogenous miR663a (or miR663AHG) expression, demonstrating the presence of a negative feedback loop of the expression of the MIR663AHG gene. However, both miR663AHG and miR663a are spliced from the same primary transcript. The abovementioned positive correlation of miR663AHG and miR663a expression in human tissues suggests that the transcriptional activity of the MIR663AHG gene should be a dominant factor in determining the expression levels of miR663AHG and miR663a in human cells, although their exact expression levels and biological functions are inversely affected by each other.
In conclusion, our study demonstrates that lncRNA miR663AHG binds to miR663a and its precursor pre-miR663a and induces the negative feedback of miR663a expression. miR663AHG may play an important role in the host defense responses to inflammation and in the inhibition of colon cancer development through cis-binding to miR663a and its precursors.
Methods
Clinical sample collection
Colon cancer and paired normal surgical margin tissue samples (SMs, >5 cm from cancer lesions) were collected from 119 patients (Table S1) at Peking University Cancer Hospital from 2004 to 2011 and stored at −80 °C. These patients were enrolled in our previous studies [7, 16]. This study complied with the 1964 Declaration of Helsinki and was approved by the Institutional Review Board of the Peking University Cancer Hospital. All patients provided informed consent before sample collection.
Cell lines and culture
The colon cancer cell lines HCT116 and SW480 were kindly provided by Professor Yuanjia Chen at Peking Union Medical College Hospital; SW620 was kindly provided by Professor Chengchao Shou at Peking University Cancer Hospital; RKO was kindly provided by Dr. Guoren Deng at the University of California; Caco2 was purchased from National Infrastructure of Cell Line Resource (Bei**g, China); and LoVo was purchased from the American Type Culture Collection (ATCC, Manassas, USA). The human cell line HEK293FT was kindly provided by Professor Yasuhito Yuasa at Tokyo Medical and Dental University. HCT116, SW480, RKO, and SW620 cells were cultured in RPMI-1640 medium containing 10% FBS. HEK293FT cells were cultured in DMEM with 10% FBS. Caco2 cells were cultured in MEM with 10% FBS and 1% NEAA (Invitrogen, Gibco, USA), and LoVo cells were cultured in F12K medium with 10% FBS. All media were supplemented with 100 U/mL penicillin/streptomycin (Life Technologies, Carlsbad, CA, USA). All cell lines were tested and authenticated by Bei**g JianLian Gene Technology Co. before they were used. Short tandem repeat (STR) patterns were analyzed using the Goldeneye 20A STR Identifier PCR Amplification Kit.
Plasmid construction and transfection
Full-length miR663AHG (ENST00000608521.6) was synthesized by GenScript (Nan**g, China) and then inserted into the pHBLV-CMV-MCS-EF1-ZsGreen-T2A-puro lentiviral expression vector (Hanbio, Shanghai, China). The empty control and miR663AHG expression vectors were generated with a lentiviral packaging kit (Syngentech Co., Ltd., Bei**g, China) according to the manufacturer’s manual in HEK293FT cells [6]. The lentivirus particle-containing culture medium was collected 48 h after transfection, filtered with a 0.45-μm filter, and added to colon cancer cells. The stably infected cells were selected for three days with 1 μg/mL puromycin (Sigma, St. Louis, MO, USA). X-tremeGENE HP DNA Transfection Reagent (Roche, Mannheim, Germany) was used in transient transfection with the miR663AHG expression vector following the manufacturer’s instructions. Transfection efficiency was monitored with the density of the ZsGreen tag integrated within these vectors and RT-PCR for target genes in the cells.
miR663a mimic and antisense/inhibitor were purchased from RiboBio Co., Ltd. (Guangzhou, China) [7, 16]. siRNAs against miR663AHG (si663AHG#1: sense 5’-gaggugcuuugccucugaatt-3’ and antisense 5’-uucagaggcaaagcaccuctt-3’; si663AHG#2: sense 5’-gcaugcaaugggcaaucuatt-3’ and antisense 5’-uagauugcccauugcaugctt-3’) were synthesized by Genepharma Co. (Shanghai, China). A mixture of siR663AHG #1 and #2 was used to knock down miR663AHG expression (siR663AHG). X-tremeGENE siRNA Transfection Reagent (Roche, Mannheim, Germany) was used in transfection with these siRNAs above. Transfection efficiency was verified using RT-PCR for miR663a or miR663AHG in the cells.
RNA extraction and quantitative RT-PCR assays (qRT-PCR)
Total RNA was isolated using TRIzol Reagent (ZYMO Research, Bei**g, China) according to the manufacturer’s instructions. First-strand cDNA was generated using the TransScript First-Strand cDNA Synthesis Kit (TransGen Biotech, Bei**g, China). Exon-exon qRT-PCR was performed using FastStart Universal SYBR Green Master Mix (Roche, Mannheim, Germany). U6 small nuclear RNA was used as an internal control for miR663a; the level of miR663AHG was normalized to that of GAPDH (for cultured cells) or Alu (for tissues). Similarly, the levels of TP53, JUNB, JUND, PIK3CD, P21, and TGFB1 mRNAs in cell lines were normalized to that of GAPDH. The relative mRNA levels further normalized to the control were further calculated using the typical 2-ΔΔCT method [7]. The relative level of pre-miR663a was calculated according to the formula [41]: pre-miR663a = 2-CT(pri-miR663a+pre-miR663a) − 2-CT(pri-miR663a). Sequences of these PCR primers are listed in Table S2.
RNA-FISH assay
Fluorescence-conjugated miR663AHG probes for miR663AHG were purchased from RiboBio Co., Ltd (Guangzhou, China). According to the manufacturer’s instructions, cells were pretreated with 4% paraformaldehyde, hybridized with miR663AHG probes labeled with Cy3, and stained with DAPI using the Ribo Fluorescent In Situ Hybridization kit (RiboBio Co., Ltd., Guangzhou, China). Probes for 18S rRNA and U6 RNA were used as cytoplasmic and nuclear RNA controls. Images were obtained with a Leica SP5 Laser Scanning Confocal Microscope (Leica, Germany).
Cell proliferation assay using IncuCyte
Long-term dynamic observation assays were used to detect cell proliferation [6]. Briefly, all cells were seeded in 96-well plates at a density of 2 × 103 cells per well. To measure proliferation, cells were photographed every 12 h in the long-term dynamic observation platform (IncuCyte, Essen, MI, USA) for at least 96 h. IncuCyte ZOOM software (Essen, Ann Arbor, MI, USA) was used to analyze cell confluence.
Wound healing assay
All cells were seeded in 6-well plates at 95–100% confluence before the experiment. A pipette tip was used to scratch the cell monolayer gently. After washing with 1 × PBS twice, the cells were cultured with RPMI-1640 medium with 1% FBS. Images of wound healing were captured at different times by a microscope (Nikon, Japan).
Transwell assays
For the migration assay, cells (2 × 104 cells per chamber) were separately seeded in the upper chambers (8 μm pores; Corning Inc., Corning, NY). For the invasion assay, the upper chambers were precoated with Matrigel (BD Biosciences, Franklin Lakes, NJ, USA). Then, cells (4 × 104 cells per chamber) were separately seeded in the upper chambers with the same aperture. All cells were resuspended in 180 μL of serum-free RPMI-1640 medium before seeding. After a 24–48 h incubation, the chambers were fixed with 4% paraformaldehyde for 30 min and then stained with 0.1% crystal violet. Nonmigrated/noninvasive cells on the upper surface of the insert were wiped with a cotton swab. Images of migrating and invading cells were captured using a microscope (Leica DMI4000B, Milton Keynes, Bucks, UK).
Animal experiments [42]
Six-week-old female BALB/c mice (purchased from Bei**g Huafukang Biotech) were used for the subcutaneous xenograft experiment. RKO cells were resuspended in 0.10 mL 1× PBS (1 × 107 cells/mL) and inoculated subcutaneously into the bilateral inguinal region of mice (6 mice per group, 1 × 106 cells per injection). Mice were sacrificed on the 21st inoculation day. All xenografts were separated, weighed, and photographed. For the experimental lung metastatic model, LoVo cells were resuspended in 0.10 mL PBS (1 × 107 cells/mL) and then injected into the tail vein of NOD/SCID mice (7 mice per group, 1 × 106 cells per injection). The mice were sacrificed eight weeks post-injection. All lung tissues were excised, weighed, and photographed. No randomization was used and no blinding was done. This study was approved by the Institute’s animal ethics committee.
Knockout of the genomic MIR663A/HG sequence by CRISPR/Cas9 [43, 44]
A dual gRNA approach was used to knock out the DNA sequence from the MIR663AHG promoter to miR663a by the CRISPR/Cas9 system. The oligonucleotides used for single guide RNA (sgRNA) construction were individually designed for the target sequence (Fig. S2 and Table S2). They were synthesized by Thermo Scientific, Inc. (Rockford, IL, USA). These gRNAs were cloned into the PX458 vector (Plasmid #48138, Addgene, Inc.) and transfected into RKO cells. Then, a flow sorting assay was performed by green fluorescence with a FACSCalibur flow cytometer (BD Biosciences, Franklin Lakes, USA) 48 h post-transfection. Two weeks later, monoclonal cells in good growth conditions were selected, and the knockout status of the target sequence in subclones was identified by PCR (Fig. S2B). The results of PCR sequencing revealed that a 751-bp genomic sequence (including the MIR663AHG promoter, exon-1, and pri-miR663a-encoding fragment) was knocked out (MIR663A/HG-KO) (Fig. S2C, gray line polygon).
RNA pulldown assay [6]
The full-length sequences of miR663AHG and pre-miR663a were amplified from the above vector and genomic DNA with a primer set (forward 5’-gtcccacggtgggggcg-3’ and reverse 5’-agttactttaaggctttatttgt-3’) and a primer set (forward 5’-ccttccggcgtcccagg-3’ and reverse 5’-catggccgggccacca-3’) and inserted into the pGEM-T easy vector [45]. These vectors were cleaved using the restriction endonuclease SpeI, and the in vitro transcription experiment followed the instructions of the Riboprobe In Vitro Transcription Systems (Promega, Madison, WI, USA). Then, the transcribed miR663AHG or pre-miR663a RNA products were labeled by biotin in vitro using the Pierce RNA 3′ End Desthiobiotinylation Kit (Thermo Scientific, Rockford, IL, USA). The biotin-labeled products were bound to streptavidin magnetic beads and incubated with separated lysates from MIR663A/HG-WT or MIR663A/HG-KO RKO cells according to the instructions of the Pierce Magnetic RNA-Protein Pull-Down Kit (Thermo Scientific, Rockford, IL, USA). RNA from the pulldown components was separated, reverse transcribed, and analyzed with quantitative PCR assays. The PCR products were analyzed using 10% nondenaturing PAGE gel electrophoresis.
The download of Cancer Cell Line Encyclopedia (CCLE) and The Cancer Genome Atlas (TCGA) datasets
MIR663AHG promoter DNA methylation data (1 kb upstream of the promoter) were extracted from the CCLE DNA methylation RRBS data file (promoter methylation -1 kb upstream of the promoter). These cell lines were subclassified into three methylation groups (methylation high, moderate, and low) with equal size. miR663AHG expression data were extracted from the RNA-seq data file (CCLE RNAseq gene expression data for 1019 cell lines, RPKM); miR663a expression data were extracted from the RNA-seq miRNA expression data file (CCLE miRNA expression data). These datasets [23] were downloaded from the CCLE official website (https://sites.broadinstitute.org/ccle/datasets). The expression levels of miR663AHG and miR663A were extracted from miRNA and lncRNA array datasets for 352 ovarian carcinoma samples in a previous study (Data file 3) [24].
Statistical analyses
Data analysis and processing software were mainly GraphPad Prism 5.0 software (San Diego, CA, USA) and IBM SPSS 23.0 software (SPSS Inc., Chicago, IL, USA). The Kolmogorov-Smirnov test or Shapiro-Wilk test was used to test whether the data conformed to a normal distribution. Student’s t-test and Pearson correlation analysis were used for data conforming to a normal distribution, while the Mann–Whitney U-test and Spearman correlation analysis were used for nonnormally distributed data. Survival analysis of patients with colon cancer was performed using the Kaplan-Meier method. Cox risk proportional regression models were used in the univariate and multifactor survival analyses. P < 0.05 (two-sided) was considered a statistically significant difference. All biological experiments were repeated one or two times.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A. Cancer statistics, 2022. CA: Cancer J Clin. 2022;72:7–33.
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, et al. Landscape of transcription in human cells. Nature 2012;489:101–8.
Qian X, Yang J, Qiu Q, Li X, Jiang C, Li J, et al. LCAT3, a novel m6A-regulated long non-coding RNA, plays an oncogenic role in lung cancer via binding with FUBP1 to activate c-MYC. J Hematol Oncol. 2021;14:112.
Liu Z, Chen Z, Fan R, Jiang B, Chen X, Chen Q, et al. Over-expressed long noncoding RNA HOXA11-AS promotes cell cycle progression and metastasis in gastric cancer. Mol Cancer. 2017;16:82.
Liu Y, Zhang P, Wu Q, Fang H, Wang Y, **ao Y, et al. Long non-coding RNA NR2F1-AS1 induces breast cancer lung metastatic dormancy by regulating NR2F1 and ΔNp63. Nat Commun. 2021;12:5232.
Ma W, Qiao J, Zhou J, Gu L, Deng D. Characterization of novel LncRNA P14AS as a protector of ANRIL through AUF1 binding in human cells. Mol Cancer. 2020;19:42.
Tian W, Du Y, Ma Y, Gu L, Zhou J, Deng D. MALAT1-miR663a negative feedback loop in colon cancer cell functions through direct miRNA-lncRNA binding. Cell Death Dis. 2018;9:857.
Liu L, Chen Y, Huang Y, Cao K, Liu T, Shen H, et al. Long non-coding RNA ANRIL promotes homologous recombination-mediated DNA repair by maintaining ATR protein stability to enhance cancer resistance. Mol Cancer. 2021;20:94.
Chen S, Shen X. Long noncoding RNAs: functions and mechanisms in colon cancer. Mol Cancer. 2020;19:167.
Liu S, Sun Y, Hou Y, Yang L, Wan X, Qin Y, et al. A novel lncRNA ROPM-mediated lipid metabolism governs breast cancer stem cell properties. J Hematol Oncol. 2021;14:178.
Tian C, Abudoureyimu M, Lin X, Chu X, Wang R. Linc-ROR facilitates progression and angiogenesis of hepatocellular carcinoma by modulating DEPDC1 expression. Cell Death Dis. 2021;12:1047.
Liu S, Bu X, Kan A, Luo L, Xu Y, Chen H, et al. SP1-induced lncRNA DUBR promotes stemness and oxaliplatin resistance of hepatocellular carcinoma via E2F1-CIP2A feedback. Cancer Lett. 2022;528:16–30.
Svoboda M, Slyskova J, Schneiderova M, Makovicky P, Bielik L, Levy M, et al. HOTAIR long non-coding RNA is a negative prognostic factor not only in primary tumors, but also in the blood of colorectal cancer patients. Carcinogenesis 2014;35:1510–5.
Zhang JX, Song W, Chen ZH, Wei JH, Liao YJ, Lei J, et al. Prognostic and predictive value of a microRNA signature in stage II colon cancer: a microRNA expression analysis. Lancet Oncol. 2013;14:1295–306.
Saus E, Brunet-Vega A, Iraola-Guzmán S, Pegueroles C, Gabaldón T, Pericay C. Long non-coding RNAs as potential novel prognostic biomarkers in colorectal cancer. Front Genet. 2016;7:54.
Tian W, Du Y, Ma Y, Zhang B, Gu L, Zhou J, et al. miR663a‑TTC22V1 axis inhibits colon cancer metastasis. Oncol Rep. 2019;41:1718–28.
Huang J, Li Y, Ye Z, Cheng Z, Huang J, Lu S, et al. Prediction of a potential mechanism of intervertebral disc degeneration based on a novel competitive endogenous RNA network. BioMed Res Int. 2021;2021:6618834.
Zhi Y, Du J, Qian M, Song N. Long non-coding RNA RP11-342L8.2, derived from RNA sequencing and validated via RT-qPCR, is upregulated and correlates with disease severity in psoriasis patients. Ir J Med Sci. 2022;191:2643–9.
The GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science. 2020;369:1318–30.
Beermann J, Piccoli MT, Viereck J, Thum T. Non-coding RNAs in development and disease: background, mechanisms, and therapeutic approaches. Physiol Rev. 2016;96:1297–325.
Schanen BC, Li X. Transcriptional regulation of mammalian miRNA genes. Genomics 2011;97:1–6.
Wang Y, He L, Du Y, Zhu P, Huang G, Luo J, et al. The long noncoding RNA lncTCF7 promotes self-renewal of human liver cancer stem cells through activation of Wnt signaling. Cell Stem Cell. 2015;16:413–25.
Ghandi M, Huang FW, Jané-Valbuena J, Kryukov GV, Lo CC, McDonald ER 3rd, et al. Next-generation characterization of the Cancer Cell Line Encyclopedia. Nature 2019;569:503–8.
Wu M, Shang X, Sun Y, Wu J, Liu G. Integrated analysis of lymphocyte infiltration-associated lncRNA for ovarian cancer via TCGA, GTEx and GEO datasets. PeerJ 2020;8:e8961.
Yu Y, Nangia-Makker P, Farhana L, Majumdar APN. A novel mechanism of lncRNA and miRNA interaction: CCAT2 regulates miR-145 expression by suppressing its maturation process in colon cancer cells. Mol Cancer. 2017;16:155.
Cai X, Cullen BR. The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA. 2007;13:313–6.
Lu Y, Zhao X, Liu Q, Li C, Graves-Deal R, Cao Z, et al. lncRNA MIR100HG-derived miR-100 and miR-125b mediate cetuximab resistance via Wnt/β-catenin signaling. Nat Med. 2017;23:1331–41.
Michaille JJ, Piurowski V, Rigot B, Kelani H, Fortman EC, Tili E. MiR-663, a MicroRNA Linked with Inflammation and Cancer That Is under the Influence of Resveratrol. Medicine. 2018;5:74.
Tili E, Michaille JJ, Alder H, Volinia S, Delmas D, Latruffe N, et al. Resveratrol modulates the levels of microRNAs targeting genes encoding tumor-suppressors and effectors of TGFβ signaling pathway in SW480 cells. Biochem Pharmacol. 2010;80:2057–65.
Tili E, Michaille JJ, Adair B, Alder H, Limagne E, Taccioli C, et al. Resveratrol decreases the levels of miR-155 by upregulating miR-663, a microRNA targeting JunB and JunD. Carcinogenesis 2010;31:1561–6.
Jiao L, Deng Z, Xu C, Yu Y, Li Y, Yang C, et al. miR-663 induces castration-resistant prostate cancer transformation and predicts clinical recurrence. J Cell Physiol. 2014;229:834–44.
Yi C, Wang Q, Wang L, Huang Y, Li L, Liu L, et al. MiR-663, a microRNA targeting p21 (WAF1/CIP1), promotes the proliferation and tumorigenesis of nasopharyngeal carcinoma. Oncogene 2012;31:4421–33.
Shi Y, Chen C, Yu SZ, Liu Q, Rao J, Zhang HR, et al. miR-663 suppresses oncogenic function of CXCR4 in glioblastoma. Clin Cancer Res. 2015;21:4004–13.
Zhang C, Chen B, Jiao A, Li F, Sun N, Zhang G, et al. miR-663a inhibits tumor growth and invasion by regulating TGF-β1 in hepatocellular carcinoma. BMC Cancer. 2018;18:1179 https://doi.org/10.1186/s12885-018-5016-z.
Kuroda K, Fukuda T, Krstic-Demonacos M, Demonacos C, Okumura K, Isogai H, et al. miR-663a regulates growth of colon cancer cells, after administration of antimicrobial peptides, by targeting CXCR4-p21 pathway. BMC Cancer. 2017;17:33.
GTEx Consortium. The Genotype-Tissue Expression (GTEx) project. Nat Genet. 2013;45:580–5.
Weinstein JN, Collisson EA, Mills GB, Shaw KR, Ozenberger BA, Ellrott K, et al. The cancer genome atlas Pan-Cancer analysis project. Nat Genet 2013;45:1113–20.
Tang Z, Kang B, Li CW, Chen TX, Zhang ZM. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 2019; https://doi.org/10.1093/nar/gkz430.
You A, Tian W, Yuan H, Gu L, Zhou J, Deng D. TTC22 promotes m6A-mediated WTAP expression and colon cancer metastasis in an RPL4 binding-dependent pattern. Oncogene 2022;41:3925–38.
Zhuang M, Zhao S, Jiang Z, Wang S, Sun P, Quan J, et al. MALAT1 sponges miR-106b-5p to promote the invasion and metastasis of colorectal cancer via SLAIN2 enhanced microtubules mobility. EBioMedicine 2019;41:286–98.
Schmittgen TD, Jiang J, Liu Q, Yang L. A high-throughput method to monitor the expression of microRNA precursors. Nucleic Acids Res. 2004;32:e43.
Zhong Q, Fang Y, Lai Q, Wang S, He C, Li A, et al. CPEB3 inhibits epithelial-mesenchymal transition by disrupting the crosstalk between colorectal cancer cells and tumor-associated macrophages via IL-6R/STAT3 signaling. J Exp Clin Cancer Res. 2020;39:132.
Pirouzfar M, Amiri F, Dianatpour M, Takhshid MA. CRISPR/Cas9-mediated knockout of MLL5 enhances apoptotic effect of cisplatin in HeLa cells in vitro. EXCLI J. 2020;19:170–82.
Ran FA, Hsu PD, Wright J, Agarwala V, Scott DA, Zhang F. Genome engineering using the CRISPR-Cas9 system. Nat Protoc. 2013;8:2281–308.
Robles J, Doers M. pGEM®-T Vector Systems troubleshooting guide. Promega Notes. 1994;45:19–20.
Acknowledgements
This work was supported by grants from the National Natural Science Foundation of China (81372592), the Bei**g Hospitals Authority’s Mission Plan (SML20191101) to DD and the Peking University Medicine Fund of Fostering Young Scholars’ Scientific & Technological Innovation and the Fundamental Research Funds for the Central Universities (BMU2020PY005) to WT.
Author information
Authors and Affiliations
Contributions
DJD was responsible for designing the study, analyzing various RNA-seq datasets, interpreting the results, obtaining the main financial support, and writing the manuscript. WT was responsible for performing part of the experiments, interpreting the results, obtaining partial financial support, and writing the manuscript. HFY was responsible for performing most experiments and interpreting the results. QWR, YTD, YWM, LKG, and JZ performed some experiments. All authors approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yuan, H., Ren, Q., Du, Y. et al. LncRNA miR663AHG represses the development of colon cancer in a miR663a-dependent manner. Cell Death Discov. 9, 220 (2023). https://doi.org/10.1038/s41420-023-01510-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-023-01510-1
- Springer Nature Limited