Abstract
Temozolomide (TMZ) resistance is a major clinical challenge for glioblastoma (GBM). O6-methylguanine-DNA methyltransferase (MGMT) mediated DNA damage repair is a key mechanism for TMZ resistance. However, MGMT-null GBM patients remain resistant to TMZ, and the process for resistance evolution is largely unknown. Here, we developed an acquired TMZ resistant xenograft model using serial implantation of MGMT-hypermethylated U87 cells, allowing the extraction of stable, TMZ resistant (TMZ-R) tumors and primary cells. The derived tumors and cells exhibited stable multidrug resistance both in vitro and in vivo. Functional experiments, as well as single-cell RNA sequencing (scRNA-seq), indicated that TMZ treatment induced cellular heterogeneity including quiescent cancer stem cells (CSCs) in TMZ-R tumors. A subset of these were labeled by NES+/SOX2+/CADM1+ and demonstrated significant advantages for drug resistance. Further study revealed that Epidermal Growth Factor Receptor (EGFR) deficiency and diminished downstream signaling may confer this triple positive CSCs subgroup’s quiescent phenotypes and chemoresistance. Continuous EGF treatment improved the chemosensitivity of TMZ-R cells both in vitro and in vivo, mechanically reversing cell cycle arrest and reduced drug uptake. Further, EGF treatment of TMZ-R tumors favorably normalized the response to TMZ in combination therapy. Here, we characterize a unique subgroup of CSCs in MGMT-null experimental glioblastoma, identifying EGF + TMZ therapy as a potential strategy to overcome cellular quiescence and TMZ resistance, likely endowed by deficient EGFR signaling.
Similar content being viewed by others
Introduction
Glioblastoma (GBM), the most malignant human brain tumor, is characterized by resistance to anti-cancer therapy [1]. Temozolomide (TMZ), an oral alkylating agent, is one of the first-line drugs commonly used in chemotherapy of GBM [2, 3]. The cytotoxicity of TMZ is commonly attributed to O6-methylguanine-induced DNA damage, which in turn induces DNA crosslinking, resulting in tumor cell death [4]. The lack of response to TMZ treatment arises from the intrinsic or acquired resistance of tumors to recurrent drug exposure. The DNA repair enzyme, O6-methylguanine methyltransferase (MGMT)- mediated demethylation at the O6 site is a key pathway that has been implicated in intrinsic TMZ resistance [5]. In gliomas, the methylation level of MGMT is also closely related to the efficacy of TMZ, where patients with hypomethylation of the MGMT promoter displaying an improved response to TMZ and survival outcomes [6]. The expression of MGMT is negatively correlated with promoter methylation status and nearly half of GBM patients experience MGMT silencing due to hypermethylation of the promoter [7, 8]. While most of them eventually develop tumor progression and acquired resistance to TMZ [9], the available evidence indicates that both MGMT promoter methylation status and MGMT protein remain relatively stable during GBM progress and TMZ treatment [6, 15], changing half of the medium every 2 days. The growth of the 3D-Spheroid cultured cells was monitored by a microscope with a real-time camera (EVOS® FL Auto Imaging System, Life Technologies, Carlsbad, CA, USA). For sphere growth assay, photographs of tumor spheres were taken at the indicated time points and sphere diameter was measured to reflect sphere growth.
Reverse-transcription semi-quantitative PCR (RT-PCR) and quantitative PCR (RT-qPCR)
Total RNA was isolated using Trizol reagent (Thermo) according to the manufacturer’s instructions. HiFiScript cDNA synthesis Kit (CoWin Bioscience, China) was used according to the manufacturer’s protocol. Quantitative PCR (qPCR) analyses were conducted to quantify mRNA relative expression using Real SYBR Mixture (CoWin Bioscience), with GAPDH as an internal control. For semi-quantitative PCR of MGMT expression, total RNA was isolated and first-strand cDNA was generated according to the manufacturer’s instructions. PCR was performed with PrimeSTAR Max DNA Polymerase (Takara). The PCR products were separated by agarose gel electrophoresis, and scanned with a Gel imager (Tanon, China). The primers are shown in Supplementary Tables 1 and 2.
Methylation-specific PCR
The MGMT promoter methylation status was determined by methylation-specific PCR (MSP). Briefly, 2 μg of DNA were subjected to bisulfite treatment using the EpiArt® DNA Methylation Bisulfite Kit (Vazyme; EM101). DNA was cleaned up following the manufacturer’s instructions and quantified. 30 ng of DNA per sample were PCR-amplified with the EpiArtTM HS Taq DNA Polymerase (Vazyme, EM201) and specific primers to detect methylated and unmethylated MGMT promoter (Supplementary Table 2). The PCR amplification protocol was as follows: 95 °C for 5 min, denature at 95 °C for 30 s, anneal at 60 °C for 30 s, extension at 70 °C for 30 s for 40 cycles, followed by a 5 min final extension.
Single-cell RNA-sequencing and data analysis
Single-cell RNA sequencing was performed using 10× Chromium single-cell platform (10X Genomics). Briefly, cells were washed with a phosphate buffer solution containing 0.04% weight/volume bovine serum albumin (BSA, Sangon). Cells were counted using Countess®II Automated Cell Counter and the concentration was adjusted to 1 × 106 /ml. The cDNA libraries were constructed using the 10× Chromium TM Single cell 3’ Library Kit according to the manufacturer’s original protocol available on the 10x Genomics website. Cell Ranger 1.3 (http://10xgenomics.com) was used to process Chromium single cell 3’ RNA-seq output. The R package ‘Seurat’ (Version 3.2.3) was used to the initial clustering and Loupe Cell Browser 3.1.0 was used to view the clustering. The R package “SingleR” (Version 1.0.1) was utilized to identify the predominant cell types. Single-cell pseudotime trajectories were constructed with the R package Monocle (Version 2.1.0) [16]. Gene set variation analysis (GSVA) was also performed by the R package. GSVA (Version 1.32.0) was used to estimate the activity for diverse signatures and pathways [17]. GSVA scores for indicated signatures were calculated using predefined gene sets obtained from the MSigDB (Molecular Signatures Database) (http://www.gsea-msigdb.org/gsea/downloads.jsp).
Multiplex immunohistochemistry/immunofluorescence (mIHC/IF)
mIHC/IF was performed using an Opal 7-color Manual IHC Kit (PerkinElmer, USA), as previously described in other studies [18, 19]. Tissue sections (4 µm thick) were labeled with primary antibodies against EGFR, Nestin, Sox2, and CADM1, followed by appropriate secondary antibodies. All antibodies used were listed in Supplementary Table 3. The slides were mounted with ProLong Gold Antifade Reagent containing DAPI, and scanned using Vectra® Polaris™ Imaging System (Akoya Biosciences). Images were analyzed by Image J software (National Institutes of Health, USA).
Immunofluorescence staining
Immunofluorescence (IF) staining was performed as previously described [14]. Cells on coverslips were fixed, permeabilized and followed with an overnight incubation of primary antibodies (Supplementary Table 3) at 4 °C. On the following day, cells were incubated with Alexa Flour 488 or Alexa Flour 594-conjugated donkey anti-mouse/rabbit secondary antibodies (Thermo) at room temperature (RT) for 30 min. Cell nuclei were counterstained with Hoechst 33342 (Thermo). The staining was visualized using an EVOS FL microscope (Life Technology, Gaithersburg, MD, USA) or a laser scanning confocal microscope (Leica Microsystems GmbH, Mannheim, Germany).
Flow cytometric assay
CD133 positive cells were measured by flow cytometer. Briefly, cells (1 × 106) were washed and incubated with fluorescence labeling antibodies (PE mouse anti-CD133, clone AC133, Miltenyi Biotec; FITC mouse anti-CD44, clone C26, BD) at RT for 30 min. Samples were washed, assayed via FACS Canto II (BD) immediately, and analyzed secondarily using Diva software (BD).
Western blot analysis
Protein was extracted using RIPA lysis buffer (Cell Signaling Technology, Danvers, MA, USA). The concentration was quantified by the BCA Protein Assay Kit (CWBIO, China). Equal amounts of proteins were loaded onto 10% polyacrylamide gel and separated by SDS-PAGE. Samples were then transferred to PVDF membranes. After blocking with 5% non-fat milk, the membranes were incubated with primary antibodies and secondary antibodies. Target bands were visualized using chemiluminescence (Millipore, Billerica, MA, USA). Antibodies used to determine the indicated proteins are specified in Supplementary Table 2.
Cellular uptake assay
Cellular uptake profiles were measured by flow cytometry. Briefly, cells were seeded into six-well plates for 24 h. After incubation with or without Dox (10 μM) for 3 h, cells were harvested and washed with cold PBS. Cells without Dox incubation were used as background controls. Dox uptake was detected by flow cytometer and analyzed with FlowJo VX software (RRID:SCR_008520, FlowJo, LCC, OR, USA). The mean fluorescence intensity (MFI) ratio of Dox (+) to their blank control was used as relative MFI.
Tumor sphere formation and growth assay
Tumor sphere formation was analyzed as described previously [20]. Briefly, cells were seeded in 6-well plate (5000 cells/well) and maintained in NSC culture medium (DMEM/F-12 + 20 ng/ml of bFGF (PeproTech, Rocky Hill, NJ) + 20 ng/ml of EGF (PeproTech) + N2 (Thermo) + B27 (Thermo) and allowed to form spheres. Photographs of tumor spheres were taken at the indicated time points and sphere diameter was measured. At least 10 random fields were analyzed per experiment.
Limiting dilution analysis (LDA)
Glioma spheres were dissociated into single-cell suspensions using Accutase cell dissociation Reagent (Millipore, Billerica, MA) and plated in 96-well low attachment plates (Corning, NY) by a limiting dilution fashion at 5, 20, 50, 100, 200 cells per well and held in NSC culture media. Fresh medium was added every 3–4 days by removing 50% of the old medium and replacing it with fresh medium. After 2-3 weeks, each well was examined for the formation of tumor spheres. Clonal frequency and significance were analyzed using the Extreme Limiting Dilution Analysis (http://bioinf.wehi.edu.au/software/elda/).
Statistical analysis
All experiments were independently repeated at least three times, and the data were expressed as mean ± standard deviation (SD). All data were analyzed with the SPSS 20.0 software package (IBM Corp.) and consistent with the normal distribution and the homogeneity variance test. A t-test was used to compare the significant difference between two groups. One-way ANOVA was used for the mean comparison between multiple sample groups, and the Tukey ad hoc test was used for intra-group multiple comparisons. P < 0.05 was considered statistically significant. Graphpad Prism 6.0 software was used for image analysis.
Results
Construction and confirmation of an acquired TMZ-resistant GBM xenograft model
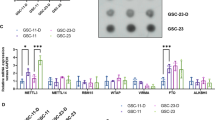
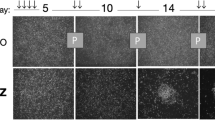
To obtain in vivo acquired TMZ-resistant GBM cells, we implanted U87 cells into nude mice and performed three cycles of TMZ treatment along with tumor passage in vivo using a mild concentration of TMZ (Fig. 1A). Temozolomide-resistant tumor tissues (TMZ-R) or cells were isolated from xenografts of serially treated mice. The parental tissues and cells isolated from saline-treated xenografts were named TMZ-S. These cells and tissues were frozen for subsequent in vitro and in vivo studies. Secondary xenograft models generated from the isolated frozen cells showed that tumors derived from TMZ-R cells grew more rapidly than those derived from TMZ-S cells (Fig. 1B). While in the presence of TMZ, TMZ-R tumors exhibited a loss response to chemotherapy, relative to the rate of DNA amplification of TMZ-S tumors (Fig. 1C). Moreover, chemotherapy-treated TMZ-R tumors showed comparable growth ability with saline-treated TMZ-S tumors (Fig. 1D). Compared with parental cells, TMZ-R generated cells showed impaired response to various anticancer drugs, as confirmed by an increased IC50 (Fig. 1E), indicating the acquisition of multidrug resistance. Two typical multidrug resistance-associated protein-coding genes, ABCB1 (MDR1) and ABCC3 [21, 22] were markedly upregulated in TMZ-R tumors and derived primary cells (Fig. 1F), revealing that drug efflux pumps may play an important role in mediating TMZ-resistance. DOX, a red fluorescent molecule, was used to determine whether drug efflux pump downregulation resulted in reduced drug accumulation in TMZ-R cells. The flow cytometry assay showed that the uptake of DOX by TMZ-R cells decreased significantly (Fig. 1G), confirming chemotherapy resistance is associated with impaired drug uptake [22, 23]. Consistent with the previous finding in U87 resistant cells [50], which is consistent with the proportion of triple positive CSCs observed in G0/G1 arrest herein.
The present study found a lowered expression of EGFR in drug-resistant, quiescent CSCs derived from TMZ-R tumors. Our findings corroborate EGFR expression and activity patterns observed in several TMZ-resistant cell lines and GBM patient tumor tissues [51]. For example, Areeb et al report that in addition to deficient EGFR expression in treatment-resistant cells, there is significant upregulation of the EGFR-targeting miR-221, which likely serves as a mechanism of reduced EGFR expression. The present findings also compliment a previous study wherein pharmacological inhibition of EGFR induced a population of quiescent GSCs expressing similar neural stem cell markers [52]. In contrast, one study found that the human EGFR inhibitor LRIG1 can regulate quiescent neural stem cells’ responsiveness to EGFR signaling (prompting NSC exit from quiescence) [53], while another reported that LRIG1 can reverse the course of multi-drug resistance by suppressing ABCB1 and ABCG2 [54] in glioblastoma cell lines. The latter studies support a role for EGFR/Akt/STAT3 signaling in conferring tumorgenicity [55] and sensitivity to temozolomide, which is at odds with our current findings. The discordance is likely rooted in the MGMT context of the different experimental models. For example, while TMZ resistance was associated with upregulation of STAT3 in the GBM cell line U87 [56, 57], this was dependent on concurrent upregulation of the DNA repair enzyme MGMT. In contrast, activation of AKT/STAT3 and upregulation of the EGFRvIII variant (present in 1/3 of GBMs) were associated with TMZ sensitivity but only in patients with highly methylated MGMT promoters (associated with loss of MGMT expression) [58]. Presented in the context of an MGMT-null GBM model, our findings of reduced EGFR expression and downstream AKT and STAT3 activation in quiescent and TMZ-resistant GSCs complement the findings of Struve et al and point to an important consideration across all GBM research.
Finally, we report that the decreased proliferation, cell cycle arrest, and sphere growth characteristic of the TMZ-R-derived CSC subpopulation were all reversible by EGF treatment. Moreover, the reduced drug response of these cells was reversed by EGF. In vivo, EGF-treated TMZ-R animals displayed a more normalized tumor volume/weight only when EGF was paired with TMZ treatment, but not by EGF alone, suggesting that while EGF can help quiescent CSCs revert to their differentiated state and the associated tumorigenic phenotypes, it cannot act alone as an effective glioblastoma therapeutic. The finding that deficient EGFR expression in TMZ-R-derived CSCs can be overridden by exogenous EGF supports the notion that the cellular phenotypes may be driven by reduced activation of EGFR signaling due to diminished expression of available receptors. In such a case, overwhelming the cells with EGFR ligand can increase the likelihood of binding/activation. Alternatively, one study has reported that exogenous EGF can stimulate the expression of NTN4, a regulator of GBM tumor progression/proliferation via ITGB4-Akt signal activation [59]. This is consistent with our observations that EGF treatment restored phosphorylated Akt in tandem with restored tumorigenic phenotypes. The Li et al study also lends support to the concept of redundancy along the EGFR/Akt/STAT3 pathways, which may incidentally explain the clinical ineffectiveness of various EGFR-targeting therapies to date [55]. Combinatorial, EGFR-targeted treatment has previously been shown to increase the efficacy of TMZ in GBM cell lines and animal-derived tumors [60]. While the aberrant EGFR expression in that study was not a downregulation of wild-type EGFR, rather expression of a common glioblastoma mutation (EGFRvIII), it supports the idea that specific cohorts of GBM patients may benefit from combined EGFR-targeting and TMZ therapy. It is worth noting that EGFR-targeted therapy is not the only treatment that aids in the re-sensitization of TMZ-R cells. Another study treated TMZ-R GBMs with the cytotoxic agent with aferin A to inhibit MGMT expression [61]. Combined withaferin A and TMZ treatment overcame the GBM’s chemoresistance and potentiated the efficacy of TMZ, similar to our findings. Collectively, our study and the existing literature support the idea that TMZ resistance of GBM tumors is multi-dimensional, likely rooted in a variety of signaling cascades and molecular events resulting in diverse cellular phenotypes which shape the drug response.
One of the limitations of this study was the subcutaneous tumor model, which may not accurately recapitulate the CNS microenvironment that shapes the course of glioblastomas clinically [62]. On the other hand, the drug-resistant cells obtained after acclimation continue to demonstrate drug-resistant properties in situ. Taken together, our model lends itself to repeated generation of stable, drug-resistant cells. Indeed, serial propagation in vivo is the defining characteristic of CSCs, as opposed to any particular molecular signature observed in vitro [63]. Toward that aim, another advantage of this model was the in vivo serial transplantation which resulted in a validated phenotype/genotype of CSCs. The nature of the model also circumvented many of the challenges observed in preclinical models which do not allow for the repopulation of tumors by surviving cancer stem cells nor the enrichment of repopulating clones due to long-term continuous chemotherapy or single-dose treatment [43]. In future studies, it will be important to thoughtfully investigate whether recurrent chemo and radio therapy truly confer the genotypes and phenotypes observed in drug-resistant quiescent CSCs or whether these subsets constitutively exist and drive the responsiveness of the tumor to treatment. Additionally, it will be pertinent to evaluate the patterns of these CSCs subsets across diverse glioblastoma patient types and characterize their plasticity throughout the course of therapeutic manipulation [63]. Finally, future efforts should continue to explore cross-talk between quiescent CSCs and adjacent non-CSCs, which have demonstrated negative feedback loops which favor the survival of oncogenic NSCs to promote tumorigenesis [40].
Despite the availability of GBM treatments, patient survival remains between 12-15 months due to tumor recurrence largely driven by treatment-resistant cells [2]. This study aimed to characterize a unique subset of quiescent, TMZ-R-derived CSCs using bioinformatics, genomics, and cellular assays. We describe the stable derivation of primary tumor cells which display the classical features of multi-drug resistant, quiescent CSCs, including reduced proliferation capacity, cell cycle arrest, diminished drug uptake, and lower amplification ability in 3D culture. Further, we identified a niche of CSCs which was particularly advantageous for evading responsivity to TMZ. The molecular signature of this subset included elevated expression of the canonical neural stem cell markers Nestin and Sox2, along with the elevated expression of CADM1. This triple-positive CSC subgroup additionally observed diminished expression of EGFR and downstream signaling which was overcome by exogenous EGF. We conclude that diminished EFGR activity in the context of MGMT-null organisms, may underlie the acquisition of TMZ resistance and quiescence of CSCs in recurrently-treated glioblastoma. In turn, restoration of EGFR signaling in patients with low MGMT expression or hypermethylated MGMT promoters may reprogram the chemotherapeutic sensitivity of TMZ-R tumors.
Data availability
The data that were analyzed during the current study are available from the corresponding author on reasonable request.
References
Alifieris C, Trafalis DT. Glioblastoma multiforme: pathogenesis and treatment. Pharmacol Ther. 2015;152:63–82.
Stupp R, Mason WP, van den Bent MJ, Weller M, Fisher B, Taphoorn MJ, et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N Engl J Med. 2005;352:987–96.
Baumann F, Bjeljac M, Kollias SS, Baumert BG, Brandner S, Rousson V, et al. Combined thalidomide and temozolomide treatment in patients with glioblastoma multiforme. J Neuro-Oncol. 2004;67:191–200.
Esteller M, Garcia-Foncillas J, Andion E, Goodman SN, Hidalgo OF, Vanaclocha V, et al. Inactivation of the DNA-repair gene MGMT and the clinical response of gliomas to alkylating agents. N Engl J Med. 2000;343:1350–4.
Hirose Y, Kreklau EL, Erickson LC, Berger MS, Pieper RO. Delayed repletion of O6-methylguanine-DNA methyltransferase resulting in failure to protect the human glioblastoma cell line SF767 from temozolomide-induced cytotoxicity. J Neurosurg. 2003;98:591–8.
Brandes AA, Franceschi E, Paccapelo A, Tallini G, De Biase D, Ghimenton C, et al. Role of MGMT methylation status at time of diagnosis and recurrence for patients with glioblastoma: clinical implications. Oncologist. 2017;22:432–7.
Hegi ME, Diserens AC, Gorlia T, Hamou MF, de Tribolet N, Weller M, et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N Engl J Med. 2005;352:997–1003.
Brennan CW, Verhaak RG, McKenna A, Campos B, Noushmehr H, Salama SR, et al. The somatic genomic landscape of glioblastoma. Cell 2013;155:462–77.
Happold C, Roth P, Wick W, Schmidt N, Florea AM, Silginer M, et al. Distinct molecular mechanisms of acquired resistance to temozolomide in glioblastoma cells. J Neurochem. 2012;122:444–55.
Yi GZ, Liu YW, **ang W, Wang H, Chen ZY, **e SD, et al. Akt and beta-catenin contribute to TMZ resistance and EMT of MGMT negative malignant glioma cell line. J Neurol Sci. 2016;367:101–6.
Gil Del Alcazar CR, Todorova PK, Habib AA, Mukherjee B, Burma S. Augmented HR repair mediates acquired temozolomide resistance in glioblastoma. Mol cancer Res: MCR. 2016;14:928–40.
An Z, Weiss WA. Cholesterol: an achilles’ heel for glioblastoma? Cancer Cell. 2016;30:653–4.
Kitange GJ, Mladek AC, Carlson BL, Schroeder MA, Pokorny JL, Cen L, et al. Inhibition of histone deacetylation potentiates the evolution of acquired temozolomide resistance linked to MGMT upregulation in glioblastoma xenografts. Clin Cancer Res. 2012;18:4070–9.
Jiao J, Zhang R, Li Z, Yin Y, Fang X, Ding X, et al. Nuclear Smad6 promotes gliomagenesis by negatively regulating PIAS3-mediated STAT3 inhibition. Nat Commun. 2018;9:2504.
Asp M, Giacomello S, Larsson L, Wu C, Furth D, Qian X, et al. A spatiotemporal organ-wide gene expression and cell atlas of the develo** human heart. Cell 2019;179:1647–60.e19.
Qiu X, Hill A, Packer J, Lin D, Ma YA, Trapnell C. Single-cell mRNA quantification and differential analysis with Census. Nat Methods. 2017;14:309–15.
Hanzelmann S, Castelo R, Guinney J. GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinform. 2013;14:7.
Yeong J, Suteja L, Simoni Y, Lau KW, Tan AC, Li HH, et al. Intratumoral CD39(+)CD8(+) T cells predict response to programmed cell death protein-1 or programmed death ligand-1 blockade in patients with NSCLC. J Thorac Oncol. 2021;16:1349–58.
Yeong J, Tan T, Chow ZL, Cheng Q, Lee B, Seet A, et al. Multiplex immunohistochemistry/immunofluorescence (mIHC/IF) for PD-L1 testing in triple-negative breast cancer: a translational assay compared with conventional IHC. J Clin Pathol. 2020;73:557–62.
Yin Y, Zhang X, Li Z, Deng L, Jiao G, Zhang B, et al. Glucocorticoid receptor beta regulates injury-mediated astrocyte activation and contributes to glioma pathogenesis via modulation of beta-catenin/TCF transcriptional activity. Neurobiol Dis. 2013;59:165–76.
Silva R, Vilas-Boas V, Carmo H, Dinis-Oliveira RJ, Carvalho F, de Lourdes Bastos M, et al. Modulation of P-glycoprotein efflux pump: induction and activation as a therapeutic strategy. Pharmacol therapeutics. 2015;149:1–123.
Robey RW, Pluchino KM, Hall MD, Fojo AT, Bates SE, Gottesman MM. Revisiting the role of ABC transporters in multidrug-resistant cancer. Nat Rev Cancer. 2018;18:452–64.
Gourbal B, Sonuc N, Bhattacharjee H, Legare D, Sundar S, Ouellette M, et al. Drug uptake and modulation of drug resistance in Leishmania by an aquaglyceroporin. J Biol Chem. 2004;279:31010–7.
Yi GZ, Huang G, Guo M, Zhang X, Wang H, Deng S, et al. Acquired temozolomide resistance in MGMT-deficient glioblastoma cells is associated with regulation of DNA repair by DHC2. Brain J Neurol. 2019;142:2352–66.
Colak S, Ten Dijke P. Targeting TGF-beta signaling in cancer. Trends Cancer. 2017;3:56–71.
Alimbetov D, Askarova S, Umbayev B, Davis T, Kipling D. Pharmacological targeting of cell cycle, apoptotic and cell adhesion signaling pathways implicated in chemoresistance of cancer cells. Int J Mol Sci. 2018;19:1690.
Guo F, Zhang H, Jia Z, Cui M, Tian J. Chemoresistance and targeting of growth factors/cytokines signalling pathways: towards the development of effective therapeutic strategy for endometrial cancer. Am J Cancer Res. 2018;8:1317–31.
Yang L, Shi P, Zhao G, Xu J, Peng W, Zhang J, et al. Targeting cancer stem cell pathways for cancer therapy. Signal Transduct Target Ther. 2020;5:8.
Mukherjee S. Quiescent stem cell marker genes in glioma gene networks are sufficient to distinguish between normal and glioblastoma (GBM) samples. Sci Rep. 2020;10:10937.
Banik A, Sharma R, Chauhan A, Singh S. Cutting the umbilical cord: cancer stem cell-targeted therapeutics. Life Sci. 2022;299:120502.
Sun X, Kaufman PD. Ki-67: more than a proliferation marker. Chromosoma. 2018;127:175–86.
Zheng GX, Terry JM, Belgrader P, Ryvkin P, Bent ZW, Wilson R, et al. Massively parallel digital transcriptional profiling of single cells. Nat Commun. 2017;8:14049.
Rutka JT, Ivanchuk S, Mondal S, Taylor M, Sakai K, Dirks P, et al. Co-expression of nestin and vimentin intermediate filaments in invasive human astrocytoma cells. Int J Dev Neurosci. 1999;17:503–15.
Sigismund S, Avanzato D, Lanzetti L. Emerging functions of the EGFR in cancer. Mol Oncol. 2018;12:3–20.
Cho DY, Lin SZ, Yang WK, Lee HC, Hsu DM, Lin HL, et al. Targeting cancer stem cells for treatment of glioblastoma multiforme. Cell Transpl. 2013;22:731–9.
Stepanenko AA, Chekhonin VP. On the critical issues in temozolomide research in glioblastoma: clinically relevant concentrations and MGMT-independent resistance. Biomedicines. 2019;7.
Rich JN, Bao S. Chemotherapy and cancer stem cells. cell stem cell. 2007;1:353–5.
Olmez I, Shen W, McDonald H, Ozpolat B. Dedifferentiation of patient-derived glioblastoma multiforme cell lines results in a cancer stem cell-like state with mitogen-independent growth. J Cell Mol Med. 2015;19:1262–72.
Tejero R, Huang Y, Katsyv I, Kluge M, Lin JY, Tome-Garcia J, et al. Gene signatures of quiescent glioblastoma cells reveal mesenchymal shift and interactions with niche microenvironment. EBioMedicine 2019;42:252–69.
Lawlor K, Marques-Torrejon MA, Dharmalingham G, El-Azhar Y, Schneider MD, Pollard SM, et al. Glioblastoma stem cells induce quiescence in surrounding neural stem cells via Notch signaling. Genes Dev. 2020;34:1599–604.
Sachdeva R, Wu M, Johnson K, Kim H, Celebre A, Shahzad U, et al. BMP signaling mediates glioma stem cell quiescence and confers treatment resistance in glioblastoma. Sci Rep. 2019;9:14569.
Garnier D, Meehan B, Kislinger T, Daniel P, Sinha A, Abdulkarim B, et al. Divergent evolution of temozolomide resistance in glioblastoma stem cells is reflected in extracellular vesicles and coupled with radiosensitization. Neuro Oncol. 2018;20:236–48.
Chan KS. Molecular pathways: targeting cancer stem cells awakened by chemotherapy to abrogate tumor repopulation. Clin Cancer Res. 2016;22:802–6.
Nunes T, Hamdan D, Leboeuf C, El Bouchtaoui M, Gapihan G, Nguyen TT, et al. Targeting cancer stem cells to overcome chemoresistance. Int J Mol Sci. 2018;19:4036.
**ushi M, Chiba S, Yoshiyama H, Masutomi K, Kinoshita I, Dosaka-Akita H, et al. Tumor-associated macrophages regulate tumorigenicity and anticancer drug responses of cancer stem/initiating cells. Proc Natl Acad Sci USA. 2011;108:12425–30.
Tang YA, Chen YF, Bao Y, Mahara S, Yatim S, Oguz G, et al. Hypoxic tumor microenvironment activates GLI2 via HIF-1alpha and TGF-beta2 to promote chemoresistance in colorectal cancer. Proc Natl Acad Sci USA. 2018;115:E5990–E9.
Singh N, Miner A, Hennis L, Mittal S. Mechanisms of temozolomide resistance in glioblastoma—a comprehensive review. Cancer Drug Resist. 2021;4:17–43.
Cai Q, Zhu A, Gong L. Exosomes of glioma cells deliver miR-148a to promote proliferation and metastasis of glioblastoma via targeting CADM1. Bull Cancer. 2018;105:643–51.
Li C, Wang Y, Wang H, Wang B, Li N, Qin Y. miR-486 promotes the invasion and cell cycle progression of ovarian cancer cells by targeting CADM1. Anal Cell Pathol. 2021;2021:7407086.
Zhang W, **e HY, Ding SM, **ng CY, Chen A, Lai MC, et al. CADM1 regulates the G1/S transition and represses tumorigenicity through the Rb-E2F pathway in hepatocellular carcinoma. Hepatobiliary Pancreat Dis Int. 2016;15:289–96.
Areeb Z, Stuart SF, West AJ, Gomez J, Nguyen HPT, Paradiso L, et al. Reduced EGFR and increased miR-221 is associated with increased resistance to temozolomide and radiotherapy in glioblastoma. Sci Rep. 2020;10:17768.
Jun HJ, Bronson RT, Charest A. Inhibition of EGFR induces a c-MET-driven stem cell population in glioblastoma. Stem Cells. 2014;32:338–48.
Marques-Torrejon MA, Williams CAC, Southgate B, Alfazema N, Clements MP, Garcia-Diaz C, et al. LRIG1 is a gatekeeper to exit from quiescence in adult neural stem cells. Nat Commun. 2021;12:2594.
Liu B, Guo Z, Dong H, Daofeng T, Cai Q, Ji B, et al. LRIG1, human EGFR inhibitor, reverses multidrug resistance through modulation of ABCB1 and ABCG2. Brain Res. 2015;1611:93–100.
Gao X, **a X, Li F, Zhang M, Zhou H, Wu X, et al. Circular RNA-encoded oncogenic E-cadherin variant promotes glioblastoma tumorigenicity through activation of EGFR-STAT3 signalling. Nat Cell Biol. 2021;23:278–91.
Kohsaka S, Wang L, Yachi K, Mahabir R, Narita T, Itoh T, et al. STAT3 inhibition overcomes temozolomide resistance in glioblastoma by downregulating MGMT expression. Mol Cancer Ther. 2012;11:1289–99.
Yang PL, Liu LX, Li EM, Xu LY. STAT3, the challenge for chemotherapeutic and radiotherapeutic efficacy. Cancers. 2020;12:2459.
Struve N, Binder ZA, Stead LF, Brend T, Bagley SJ, Faulkner C, et al. EGFRvIII upregulates DNA mismatch repair resulting in increased temozolomide sensitivity of MGMT promoter methylated glioblastoma. Oncogene. 2020;39:3041–55.
Li L, Huang Y, Gao Y, Shi T, Xu Y, Li H, et al. EGF/EGFR upregulates and cooperates with Netrin-4 to protect glioblastoma cells from DNA damage-induced senescence. BMC Cancer. 2018;18:1215.
Nitta Y, Shimizu S, Shishido-Hara Y, Suzuki K, Shiokawa Y, Nagane M. Nimotuzumab enhances temozolomide-induced growth suppression of glioma cells expressing mutant EGFR in vivo. Cancer Med. 2016;5:486–99.
Grogan PT, Sarkaria JN, Timmermann BN, Cohen MS. Oxidative cytotoxic agent withaferin A resensitizes temozolomide-resistant glioblastomas via MGMT depletion and induces apoptosis through Akt/mTOR pathway inhibitory modulation. Investig N Drugs. 2014;32:604–17.
Lenting K, Verhaak R, Ter Laan M, Wesseling P, Leenders W. Glioma: experimental models and reality. Acta Neuropathol. 2017;133:263–82.
Vidal SJ, Rodriguez-Bravo V, Galsky M, Cordon-Cardo C, Domingo-Domenech J. Targeting cancer stem cells to suppress acquired chemotherapy resistance. Oncogene. 2014;33:4451–63.
Acknowledgements
This work was supported by Natural Science Foundation of China (NFSC) grants (no. 81872056, 82172954, 81903391, 82003581, and 81802493), Natural Science Foundation of Jiangsu Province (BK20190148, 20191140), Wuxi key medical discipline construction project (CXTD2021006), Jiangsu Young Medical Talents (QNRC2016188), Wuxi Key Medical Talents (ZDRC001), Wuxi Science and Technology Development Fund (N20192048), Taihu Talent Plan (JZ), Reserve Talents of Double Hundred Talent Plan (HB2020017, HB2020018), General Program of Jiangsu Commission of Health (M2020012), Wuxi Translational Medicine Research Project (2020ZHZD04). We thank Clarity Manuscript Consultants for assistance with language editing (www.claritymanuscripts.com/about.html).
Author information
Authors and Affiliations
Contributions
All authors read the manuscript and provided feedback. JZ and YY designed the study and prepared the manuscript; LG and CC performed experiments and acquired data; DX and MW analyzed data; BZ, ZP, and QW prepared materials and raised animals.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics
The study was approved by the Institutional Ethics Committee of The Affiliated Wuxi People’s Hospital of Nan**g Medical University to the commencement of the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gong, L., Yin, Y., Chen, C. et al. Characterization of EGFR-reprogrammable temozolomide-resistant cells in a model of glioblastoma. Cell Death Discov. 8, 438 (2022). https://doi.org/10.1038/s41420-022-01230-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41420-022-01230-y
- Springer Nature Limited