Abstract
Epithelial-mesenchymal transition (EMT) is a continuum that includes epithelial, partial EMT, and mesenchymal states, each of which is associated with cancer progression, invasive capabilities, and ultimately, metastasis. We used a lineage-traced sporadic model of pancreatic cancer to generate a murine organoid biobank from primary and secondary tumors, including sublines that underwent partial EMT and complete EMT. Using an unbiased proteomics approach, we found that organoid morphology predicts the EMT state, and the solid organoids are associated with a partial EMT signature. We also observed that exogenous TGFβ1 induces solid organoid morphology that is associated with changes in the S100 family, complete EMT, and the formation of high-grade tumors. S100A4 may be a useful biomarker for predicting EMT state, disease progression, and outcome in patients with pancreatic cancer.

Similar content being viewed by others
Introduction
Pancreatic ductal adenocarcinoma (PDAC) is among the most lethal malignancies, with a 5-year survival rate of less than 12% [1]. Many patients succumb to the disease within the first 6 months, reflecting late diagnosis, metastasis, and therapy resistance [2, 3]. A distinctive feature of PDAC is the progression of early pancreatic intraepithelial neoplasia to an advanced unresectable disease [2, 3]. During this process, the histopathological phenotype of the tumor changes from low-grade, characterized by well-formed glandular epithelial structures, to high-grade, whereby epithelial cells are diffuse throughout the desmoplastic stroma [2, 3]. This process of dissemination has been linked to the activation of epithelial-mesenchymal transition (EMT), a cell-biological program that is essential for the early stages of embryogenesis and organ formation and is exploited by cancer cells [4]. EMT supports pathological events associated with the loss of epithelial behavior and the acquisition of mesenchymal features that enable invasion and metastasis [4].
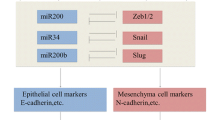
In PDAC, patient analysis, in vitro assays, xenograft models, and genetically engineered mouse models (GEMMs) have suggested that a set of transcription factors (TF) enables EMT [5,6,7,8,9,10]. We now know that PDAC progression is not solely driven by the classic EMT-TFs Snail or Twist [6]; however, the loss of EMT-TF Zeb1 can result in the growth of low-grade tumors [8]. These EMT-TFs were originally considered part of a binary process, whereby cells transition into two distinct epithelial or mesenchymal states, the latter defined by the loss of expression of the epithelial protein E-cadherin and gain of expression of the mesenchymal protein vimentin [4]. This concept has now shifted to reflect a continuum of partial EMT states with overlap** transcriptional features found in multiple cancers, including PDAC [4, 11]. Cancer cells can also revert from partial EMT to epithelial states through mesenchymal-epithelial transition (MET) [9, 10]. This spectrum of heterogeneous EMT states may be associated with differences in epithelial plasticity and cell migration, thereby influencing tumor progression, metastasis, and response to treatment [11, 12]. The intrinsic mechanisms that drive the partial EMT or complete EMT remain unclear.
PDAC tumor cells reside within a dense desmoplastic stroma comprising cancer-associated fibroblasts (CAFs) and associated extracellular matrix (ECM), which constitutes more than 80% of the tumor mass [13]. The mechanism by which the stroma modulates the behavior of tumor cells, particularly the progression of EMT, remains unclear. A primary focus has been placed on the cytokine TGFβ1, which is potently secreted by CAFs [14], resulting in autocrine signaling and differentiation into myofibroblastic CAFs that promote the deposition of ECM proteins [15, 16]. Paracrine TGFβ1 signaling can also act on benign neoplastic epithelial cells, leading to cell cycle arrest or on neoplastic cells, to induce proliferation, motility, and EMT [11]. Similar to previous reports on patient-derived PDAC organoids [17, 44,45,46,47,48], we showed that solid murine organoids are often derived from high-grade tumors. Our proteomic analysis revealed that the morphology of PDAC organoids correlates with the continuum of EMT states, which has been previously reported for mammary organoids [49, 50]. We found that within organoids isolated from both primary and secondary tumors, glandular organoids were often in an epithelial state, whereas solid organoids often underwent partial EMT. Although solid morphology was a feature of acinar organoids in previously published work [51, 52], no enrichment for acinar signatures were detected in our solid organoid proteomics dataset, ruling out an acinar origin for these samples (Fig. S4A). We also found that partial EMT organoids give rise to different EMT spectra along a previously described trajectory of the epithelial, EMT-hybrid, and mesenchymal states [22]. Interestingly, following serial passaging and allograft generation, a subset of solid organoids with a partial EMT phenotype underwent MET, characterized by reversion to a glandular phenotype and re-expression of EpCAM. This phenomenon is not uncommon, and has previously been reported ex vivo, following the culture of partial EMT cell lines [11] and in vivo following the migration of cells from primary tumors to distant organs [9, 10].
Among the proteins identified within the partial EMT state in solid organoids is the S100 protein family, which consists of 25 members with a variety of intracellular and extracellular cellular functions, including calcium homeostasis, proliferation, and apoptosis [53]. S100 proteins have also been shown to interact with cytoskeletal proteins, thereby affecting cellular morphology and migration [43, 53]. We were intrigued by the S100 proteins, as they have previously been shown to be secreted by epithelial cells in response to tissue damage or inflammatory responses [28, 54]. We found that S100a14 was increased in the organoids derived from PDAC tumors compared to the normal organoids, which is consistent with the increased expression observed in murine cancer cells and patient cancer cell lines and was linked to poor overall survival [22, 38, 55]. S100a14 has been described as a mesenchymal marker because upregulation of the transcription factor Gli1 promotes EMT and increases s100a14 expression [56]. However, using murine models [55] and human cell lines [38], we and others have shown that s100a14 is associated with epithelial and not EMT states, as its expression is lower in glandular MOs, murine tumors, and patients with low-grade tumors.
We were also interested in the increased expression of s100a4, which is associated with partial EMT in solid organoids, because s100a4 has also been associated with poor survival in PDAC [57]. S100a4 has also been shown to play a role in EMT and metastasis and is highly expressed in murine PDAC mesenchymal tumor cells, patient cell lines, and PDAC patients with high-grade tumors, suggesting a clear association with EMT [22, 38, 56]. As s100a14 is also increased in tumor cells and is associated with poor prognosis, S100A4 may be a more useful biomarker for treatment of advanced disease since high S100A4 expression correlates with resistance to gemcitabine, the mainstay chemotherapy provided to patients with PDAC [58, 59], and EMT is known to alter the response to chemotherapy [60].
Secreted factors within the tumor microenvironment are known to influence EMT in PDAC [21]. We explored the contribution of TGFβ1 and found that recombinant TGFβ1 could transition a glandular organoid to a solid phenotype, which is associated with the induction of a classic EMT gene signature. As previously shown in PDAC patient-derived organoids [17], we observed that a branching phenotype could also be induced by recombinant TGFβ1 suggesting a transition to a complete EMT state. We were particularly intrigued by the observation that TGFβ1 could upregulate S100a4 consistent with other studies [7, 8]. It is thought that TGFβ1 may regulate S100a4 through the EMT-TF, Zeb1, as Zeb1 overexpression also resulted in an increase in S100A4 expression in cell lines, while S100A14 decreased, consistent with our observations [38].
Despite being an obvious therapeutic target to prevent the transition from early stage EMT to complete EMT, there is likely limited clinical benefit following targeting of TGFβ1 in cancer due to the high probability of on-target toxicities due to the complexities of TGF signaling and function within the immune system. For this reason, we explored the role of IL-6 family cytokines in EMT, which are readily targeted therapeutically, and which we and others have shown to be induced downstream of TGFβ1 signaling [18, 19]. However, we failed to detect changes in organoid morphology following stimulation with recombinant IL-6 family cytokines nor did we detect an increase in classic EMT markers, suggesting that this cytokine family alone does not promote EMT. Moreover, despite previous reports showing that IL-11 upregulates S100A4 and S100A14 in human PDAC cell lines [38], IL-11Rα1 expression was not detected in the organoid biobank. Other studies have also shown that the treatment of lung cancer cell lines with IL-6 alone did not induce EMT, although the presence of receptor components was not confirmed [61]. We also did not observe the induction of S100a4 following stimulation with IL-6 or LIF. However, we observed a significant induction of S100a14 suggesting that although IL-6 and LIF may contribute to tumor features, they do not directly contribute to EMT. Thus, the therapeutic inhibition of the IL-6 cytokine family alone is unlikely to prevent EMT. However, the activation of signal transducer and activator of transcription (STAT)−3, a pro-tumorigenic transcription factor downstream of the IL-6 family of cytokines, is significantly correlated with S100A4 expression in PDAC patient tissues [38] suggesting that cooperative induction of S100A4 is possible. Previous reports have also suggested that synergy between ZEB1 EMT-TF and IL-6 family cytokines results in the augmentation of S100A4 expression, which was not explored in this study [38].
Taken together, our observations suggest that both extracellular and cell-intrinsic complete EMT programs converge on S100A4 and may represent useful biomarkers for predicting disease progression and prognosis.
Materials and methods
Mouse strains
PdxCre; RosaYFP (CY) and PdxCre; KrasG12V; p53R172H; RosaYFP (CKPY) mice [5] were maintained on a C57BL/6 background and bred and maintained in a specific pathogen-free animal facility at WEHI. All experiments involving mice were approved by the WEHI Animal Ethics Committee (AEC approval #2019.015 and #2020.032). CKPY mice were aged between 12 and 27 weeks and were collected together with aged and sex-matched CY mice.
Generation of organoids
FACs sorted cells from murine tissue were resuspended in >90% v/v Matrigel (Corning) and grown in murine pancreatic organoid medium (MPOM) containing advanced DMEM/F12 (Gibco) containing 10 mmol/L HEPES (Gibco), 1X GlutaMAX (Gibco), 1x penicillin/streptomycin (Gibco), 5% v/v Rspo2-Fc conditioned medium (harvested from transiently transfected FreestyleTM-293F cells, Thermo Fisher), 5% v/v Noggin-conditioned medium (harvested from Noggin-expressing 239 cells obtained from Foundation Hubrecht Organoid Technology (HUB), Hubrecht Institute, Uterecht, The Netherlands), 10 mmol/L nicotinamide, 1% v/v B-27 supplement without vitamin A, 1 mmol/L N-acetyl-L-cysteine, 100 ng/mL rh FGF-10, 50 ng/mL rh EGF, 10 nmol/L rh [Leu15]-gastrin I, and 3 µmol/L prostaglandin E2. Following passaging, the organoids were cultured in MPOM with 10 µM Y-27632 (Sigma) and 5 µM GSK-3 inhibitor (Sigma) for the first three days, followed by culturing in normal MPOM for regular maintenance, as described previously [62].
Generation of allografts
Wild-type C57BL/6 mice were used to establish murine organoid-derived allografts. Wild-type C57BL/6 mice were bred and maintained in a specific pathogen-free animal facility at WEHI. All experiments involving mice were approved and monitored by the WEHI Animal Ethics Committee (AEC Approval #2017.033 and ##2020.032).
To generate allografts, one confluent well of organoids (approximately 50,000 cells) was resuspended in 100 µL of 50% v/v PBS/ 50% v/v Matrigel and subcutaneously injected into each flank of C57BL/6 mice (gender matched to the organoid). If no visible tumor was present 3 months after engraftment, engraftment was deemed unsuccessful and the mouse was euthanized.
Mass spectrometry-based proteomics
Organoids were lysed in RIPA buffer containing 100 mM NaCl, 10 mM Tris-HCl, 1% (v/v) glycerol, 50 mM NaF, 2 mM EDTA, 1% (v/v) Triton X-100, 1 mM Na3VO4, complete mini protease inhibitor cocktail, and complete mini phosphatase inhibitor, and 20 µg per replicate was prepared for proteomic analysis using the USP3 protocol previously described [63] with some minor modifications. Lysates were heated at 95 °C for 10 min in buffer containing 1% (v/v) SDS, 100 mM Tris (pH 8), 10 mM Tris (2-carboxyethyl) phosphine (TCEP), and 40 mM 2-chloracetamide. Magnetic PureCube Carboxy agarose beads (Cube Biotech) were added to all the samples along with acetonitrile (70% v/v final concentration) and incubated at room temperature for 20 min. Samples were placed on a 96-well magnetic rack, supernatants were discarded, and beads were washed twice with 70% ethanol and once with neat ACN. ACN was completely evaporated from the tubes using a CentriVap (Labconco) before the addition of digestion buffer (50 mm Tris) containing 0.8 µg Lys-C (Wako, 129–02541) and 0.8 µg Trypsin-gold (Promega, V5280). Enzymatic digestion was performed with agitation (400 rpm) for 1 h at 37 °C. Following digestion, the samples were transferred to pre-equilibrated C18 StageTips for sample clean-up. The eluates were lyophilized to dryness using CentriVap (Labconco) before being reconstituted in 60 µL of 0.1% FA/2% ACN ready for mass spectrometry analysis.
For DDA analysis, peptides (2 µL) were separated by reverse-phase chromatography on a C18 fused silica column packed into an emitter tip (IonOpticks) using a nano-flow HPLC (M-class, Waters). HPLC was coupled to a timsTOF Pro (Bruker) equipped with a CaptiveSpray source. Peptides were loaded directly onto the column at a constant flow rate of 400 nL/min with buffer A (99.9% v/v Milli-Q water, 0.1% v/v FA) and eluted with a 90-min linear gradient from 2 to 34% buffer B (99.9% v/v ACN, 0.1% v/v FA). The timsTOF Pro was operated in PASEF mode using Compass Hystar 5.1. Settings were as follows: Mass Range 100 to 1700 m/z, 1/K0 Start 0.6 V·s/cm2 End 1.6 V·s/cm2, Ramp time 110.1 ms, Lock Duty Cycle to 100%, Capillary Voltage 1600V, Dry Gas 3 l/min, Dry Temp 180 °C, PASEF settings: 10 MS/MS scans (total cycle time 1.27 sec), charge range 0-5, active exclusion for 0.4 min, Scheduling Target intensity 10000, Intensity threshold 2500, CID collision energy 42 eV.
Data availability
Mass spectrometry proteomics data were deposited in the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD030992.
Change history
22 March 2023
A Correction to this paper has been published: https://doi.org/10.1038/s41418-023-01151-y
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA: A Cancer J Clin. 2020;70:7–30.
Low RRJ, Lim WW, Nguyen PM, Lee B, Christie M, Burgess AW, et al. The diverse applications of pancreatic ductal adenocarcinoma organoids. Cancers. 2021;13:4979.
Feldmann G, Beaty R, Hruban RH, Maitra A. Molecular genetics of pancreatic intraepithelial neoplasia. J Hepatobiliary Pancreat Surg. 2007;14:224–32.
Yang J, Antin P, Berx G, Blanpain C, Brabletz T, Bronner M, et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat Rev Mol Cell Biol. 2020;21:341–52.
Rhim AD, Mirek ET, Aiello NM, Maitra A, Bailey JM, McAllister F, et al. EMT and dissemination precede pancreatic tumor formation. Cell. 2012;148:349–61.
Zheng X, Carstens JL, Kim J, Scheible M, Kaye J, Sugimoto H, et al. Epithelial-to-mesenchymal transition is dispensable for metastasis but induces chemoresistance in pancreatic cancer. Nature. 2015;527:525.
Chen Y, LeBleu VS, Carstens JL, Sugimoto H, Zheng X, Malasi S, et al. Dual reporter genetic mouse models of pancreatic cancer identify an epithelial‐to‐mesenchymal transition‐independent metastasis program. EMBO Mol Med. 2018;10:e9085.
Krebs AM, Mitschke J, Lasierra Losada M, Schmalhofer O, Boerries M, Busch H, et al. The EMT-activator Zeb1 is a key factor for cell plasticity and promotes metastasis in pancreatic cancer. Nat Cell Biol. 2017;19:518–29.
Beerling E, Seinstra D, de Wit E, Kester L, van der Velden D, Maynard C, et al. Plasticity between epithelial and mesenchymal states unlinks EMT from metastasis-enhancing stem cell capacity. Cell Rep. 2016;14:2281–8.
Reichert M, Bakir B, Moreira L, Pitarresi JR, Feldmann K, Simon L, et al. Regulation of epithelial plasticity determines metastatic organotropism in pancreatic cancer. Dev cell. 2018;45:696–711.e698.
Aiello NM, Maddipati R, Norgard RJ, Balli D, Li J, Yuan S, et al. EMT subtype influences epithelial plasticity and mode of cell migration. Dev Cell. 2018;45:681–95.e684.
Lüönd F, Sugiyama N, Bill R, Bornes L, Hager C, Tang F, et al. Distinct contributions of partial and full EMT to breast cancer malignancy. Dev Cell. 2021;56:3203–21.e3211.
Dougan SK. The pancreatic cancer microenvironment. Cancer J (Sudbury, Mass). 2017;23:321–5.
Marzoq AJ, Mustafa SA, Heidrich L, Hoheisel JD, Alhamdani MSS. Impact of the secretome of activated pancreatic stellate cells on growth and differentiation of pancreatic tumour cells. Sci Rep. 2019;9:5303.
Biffi G, Oni TE, Spielman B, Hao Y, Elyada E, Park Y, et al. IL1-Induced JAK/STAT signaling is antagonized by TGFβ to shape CAF heterogeneity in pancreatic ductal adenocarcinoma. Cancer Disco. 2019;9:282–301.
Choi JW, Lee SK, Kim MJ, Kim DG, Shin JY, Zhou Z, et al. Piperine ameliorates the severity of fibrosis via inhibition of TGF‑β/SMAD signaling in a mouse model of chronic pancreatitis. Mol Med Rep. 2019;20:3709–18.
Huang W, Navarro-Serer B, Jeong YJ, Chianchiano P, **a L, Luchini C, et al. Pattern of invasion in human pancreatic cancer organoids is associated with loss of SMAD4 and clinical outcome. Cancer Res. 2020;80:2804.
Su J, Morgani SM, David CJ, Wang Q, Er EE, Huang YH, et al. TGF-β orchestrates fibrogenic and developmental EMTs via the RAS effector RREB1. Nature. 2020;577:566–71.
David CJ, Huang YH, Chen M, Su J, Zou Y, Bardeesy N, et al. TGF-β tumor suppression through a Lethal EMT. Cell. 2016;164:1015–30.
Gabitova-Cornell L, Surumbayeva A, Peri S, Franco-Barraza J, Restifo D, Weitz N, et al. Cholesterol pathway inhibition induces TGF-β signaling to promote basal differentiation in pancreatic cancer. Cancer Cell. 2020;38:567–83.e511.
Ligorio M, Sil S, Malagon-Lopez J, Nieman LT, Misale S, Di Pilato M, et al. Stromal microenvironment shapes the intratumoral architecture of pancreatic cancer. Cell. 2019;178:160–75.e127.
Simeonov KP, Byrns CN, Clark ML, Norgard RJ, Martin B, Stanger BZ, et al. Single-cell lineage tracing of metastatic cancer reveals selection of hybrid EMT states. Cancer Cell. 2021;39:1150–62.e1159.
Dumartin L, Whiteman HJ, Weeks ME, Hariharan D, Dmitrovic B, Iacobuzio-Donahue CA, et al. AGR2 is a novel surface antigen that promotes the dissemination of pancreatic cancer cells through regulation of cathepsins B and D. Cancer Res. 2011;71:7091–102.
Deng Y, Ma H, Hao J, **e Q, Zhao RMCM2. and NUSAP1 are potential biomarkers for the diagnosis and prognosis of pancreatic cancer. BioMed Res Int. 2020;2020:8604340.
Hu D, Ansari D, Zhou Q, Sasor A, Said Hilmersson K, Andersson R. Galectin 4 is a biomarker for early recurrence and death after surgical resection for pancreatic ductal adenocarcinoma. Scand J Gastroenterol. 2019;54:95–100.
Alikanoglu AS, Gunduz S, Demirpence O, Suren D, Gunduz UR, Sezer C, et al. Expression pattern and prognostic significance of claudin 1, 4 and 7 in pancreatic cancer. Asian Pac J Cancer Prev. 2015;16:4387–92.
Holczbauer Á, Gyöngyösi B, Lotz G, Szijártó A, Kupcsulik P, Schaff Z, et al. Distinct claudin expression profiles of hepatocellular carcinoma and metastatic colorectal and pancreatic carcinomas. J Histochem Cytochem. 2013;61:294–305.
Tian C, Clauser KR, Öhlund D, Rickelt S, Huang Y, Gupta M, et al. Proteomic analyses of ECM during pancreatic ductal adenocarcinoma progression reveal different contributions by tumor and stromal cells. Proc Natl Acad Sci USA. 2019;116:19609–18.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–W102.
Raphael BJ, Hruban RH, Aguirre AJ, Moffitt RA, Yeh JJ, Stewart C, et al. Integrated genomic characterization of pancreatic ductal adenocarcinoma. Cancer Cell. 2017;32:185–203.e113.
Elyada E, Bolisetty M, Laise P, Flynn WF, Courtois ET, Burkhart RA, et al. Cross-species single-cell analysis of pancreatic ductal adenocarcinoma reveals antigen-presenting cancer-associated fibroblasts. Cancer Disco. 2019;9:1102–23.
Bernard V, Semaan A, Huang J, San Lucas FA, Mulu FC, Stephens BM, et al. Single-cell transcriptomics of pancreatic cancer precursors demonstrates epithelial and microenvironmental heterogeneity as an early event in neoplastic progression. Clin Cancer Res. 2019;25:2194.
David CJ, Huang YH, Chen M, Su J, Zou Y, Bardeesy N, et al. TGF-beta tumor suppression through a lethal EMT. Cell. 2016;164:1015–30.
Alvarez MA, Freitas JP, Mazher Hussain S, Glazer ES. TGF-β inhibitors in metastatic pancreatic ductal adenocarcinoma. J Gastrointest Cancer. 2019;50:207–13.
Lee JW, Stone ML, Porrett PM, Thomas SK, Komar CA, Li JH, et al. Hepatocytes direct the formation of a pro-metastatic niche in the liver. Nature. 2019;567:249–52.
Shi Y, Gao W, Lytle NK, Huang P, Yuan X, Dann AM, et al. Targeting LIF-mediated paracrine interaction for pancreatic cancer therapy and monitoring. Nature. 2019;569:131–5.
Ohlund D, Handly-Santana A, Biffi G, Elyada E, Almeida AS, Ponz-Sarvise M, et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J Exp Med. 2017;214:579–96.
Al-Ismaeel Q, Neal CP, Al-Mahmoodi H, Almutairi Z, Al-Shamarti I, Straatman K, et al. ZEB1 and IL-6/11-STAT3 signalling cooperate to define invasive potential of pancreatic cancer cells via differential regulation of the expression of S100 proteins. Br J Cancer. 2019;121:65–75.
Nieto MA, Huang Ruby Y-J, Jackson Rebecca A, Thiery Jean P. EMT: 2016. Cell. 2016;166:21–45.
Boj SF, Hwang CI, Baker LA, Chio II, Engle DD, Corbo V, et al. Organoid models of human and mouse ductal pancreatic cancer. Cell. 2015;160:324–38.
Messal HA, Alt S, Ferreira RMM, Gribben C, Wang VM-Y, Cotoi CG, et al. Tissue curvature and apicobasal mechanical tension imbalance instruct cancer morphogenesis. Nature. 2019;566:126–30.
Roe JS, Hwang CI, Somerville TDD, Milazzo JP, Lee EJ, Da Silva B, et al. Enhancer reprogramming promotes pancreatic cancer metastasis. Cell. 2017;170:875–88.e820.
Gires O, Pan M, Schinke H, Canis M, Baeuerle PA. Expression and function of epithelial cell adhesion molecule EpCAM: where are we after 40 years? Cancer Metastasis Rev. 2020;39:969–87.
Tiriac H, Belleau P, Engle DD, Plenker D, Deschenes A, Somerville T, et al. Organoid profiling identifies common responders to chemotherapy in pancreatic cancer. Cancer Disco. 2018;8:1112–29.
Seino T, Kawasaki S, Shimokawa M, Tamagawa H, Toshimitsu K, Fujii M, et al. Human pancreatic tumor organoids reveal loss of stem cell niche factor dependence during disease progression. Cell Stem Cell. 2018;22:454–67.e456.
Romero-Calvo I, Weber C, Ray M, Brown M, Kirby K, Nandi RK, et al. Human organoids share structural and genetic features with primary pancreatic adenocarcinoma tumors. Mol Cancer Res. 2019;17:70–83.
Driehuis E, van Hoeck A, Moore K, Kolders S, Francies HE, Gulersonmez MC, et al. Pancreatic cancer organoids recapitulate disease and allow personalized drug screening. Proc Natl Acad Sci USA. 2019;116:26580–90.
Sharick JT, Walsh CM, Sprackling CM, Pasch CA, Pham DL, Esbona K, et al. Metabolic heterogeneity in patient tumor-derived organoids by primary site and drug treatment. Front Oncol. 2020;10:553.
Jung H-Y, Fattet L, Tsai JH, Kajimoto T, Chang Q, Newton AC, et al. Apical–basal polarity inhibits epithelial–mesenchymal transition and tumour metastasis by PAR-complex-mediated SNAI1 degradation. Nat Cell Biol. 2019;21:359–71.
Shamir ER, Pappalardo E, Jorgens DM, Coutinho K, Tsai WT, Aziz K, et al. Twist1-induced dissemination preserves epithelial identity and requires E-cadherin. J Cell Biol. 2014;204:839–56.
Ferreira RMM, Sancho R, Messal HA, Nye E, Spencer-Dene B, Stone RK, et al. Duct- and acinar-derived pancreatic ductal adenocarcinomas show distinct tumor progression and marker expression. Cell Rep. 2017;21:966–78.
Huang L, Desai R, Conrad DN, Leite NC, Akshinthala D, Lim CM, et al. Commitment and oncogene-induced plasticity of human stem cell-derived pancreatic acinar and ductal organoids. Cell Stem Cell. 2021;28:1090–104.e1096.
Wang T, Huo X, Chong Z, Khan H, Liu R, Wang T. A review of S100 protein family in lung cancer. Clin Chim Acta. 2018;476:54–59.
**a C, Braunstein Z, Toomey AC, Zhong J, Rao X. S100 proteins as an important regulator of macrophage inflammation. Front Immunol. 2018;8:1908.
Hosein AN, Huang H, Wang Z, Parmar K, Du W, Huang J, et al. Cellular heterogeneity during mouse pancreatic ductal adenocarcinoma progression at single-cell resolution. JCI Insight. 2019;5:e129212.
Xu X, Su B, **e C, Wei S, Zhou Y, Liu H, et al. Sonic hedgehog-Gli1 signaling pathway regulates the epithelial mesenchymal transition (EMT) by mediating a new target gene, S100A4, in pancreatic cancer cells. PLoS One. 2014;9:e96441–e96441.
Dreyer SB, Pinese M, Jamieson NB, Scarlett CJ, Colvin EK, Pajic M, et al. Precision oncology in surgery: patient selection for operable pancreatic cancer. Ann Surg. 2020;272:366–76.
Mahon PC, Baril P, Bhakta V, Chelala C, Caulee K, Harada T, et al. S100A4 contributes to the suppression of BNIP3 expression, chemoresistance, and inhibition of apoptosis in pancreatic cancer. Cancer Res. 2007;67:6786.
Ma G, Sun Y, Fu S. Evaluation of S100A4 mRNA in EUS-FNA specimens for the assessment of chemosensitivity to gemcitabine from patients with unresectable pancreatic cancer. Int J Clin Exp Pathol. 2015;8:13284–8.
Shibue T, Weinberg RA. EMT, CSCs, and drug resistance: the mechanistic link and clinical implications. Nat Rev Clin Oncol. 2017;14:611–29.
Liu RY, Zeng Y, Lei Z, Wang L, Yang H, Liu Z, et al. JAK/STAT3 signaling is required for TGF-β-induced epithelial-mesenchymal transition in lung cancer cells. Int J Oncol. 2014;44:1643–51.
Broutier L, Andersson-Rolf A, Hindley CJ, Boj SF, Clevers H, Koo BK, et al. Culture and establishment of self-renewing human and mouse adult liver and pancreas 3D organoids and their genetic manipulation. Nat Protoc. 2016;11:1724–43.
Louis C, Souza-Fonseca-Guimaraes F, Yang Y, D’Silva D, Kratina T, Dagley L, et al. NK cell-derived GM-CSF potentiates inflammatory arthritis and is negatively regulated by CIS. J Exp Med. 2020;217:e20191421.
Anaya J. OncoLnc: linking TCGA survival data to mRNAs, miRNAs, and lncRNAs. PeerJ Computer Sci. 2016;2:e67.
Acknowledgements
We wish to thank the WEHI bioservices and histological facilities for their excellent technical support.
Funding
This work was supported by generous donations from the Philip Hemstritch Pancreatic Cancer Research Program and Donald Cant Watts Corke (Australia). TLP is supported by the Sylvia and Charles Viertel Charitable Foundation Senior Medical Research Fellowship. Funding from the Victorian State Government Operational Infrastructure Support Scheme is acknowledged. Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
RRJL and TLP conceived of the study, designed the experiments, and wrote the manuscript. RRJL performed all MO experiments. KYF and AP contributed to in vivo experiments and KYF, AP, and PN contributed to TGFβ MO experiments. HG analyzed the public transcriptomic and proteomic datasets. JY, SJEC, LFD, and RHL performed proteomic analysis. BL and KYF performed the serum analyses, and PG provided human samples. NK, AWB, and MDWG contributed critical reagents and intellectual input. FH, MDWG, and SMG supervised experiments and provided intellectual support. All the authors have read and agreed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no conflict of interest. TLP has consulted for enterprises involved in biological drug development (Mestag Therapeutics, Enleofen Ltd). MDWG has consulted for enterprises involved in biological drug development (Mestag Therapeutics).
Ethics approval
All experiments involving mice were approved by the WEHI Animal Ethics Committee (AEC approval #2019.015 and #2020.032). Blood was collected from de-identified healthy or PDAC patients who had consented to a project governed by WEHI (G16/05).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: During the revision process, dr paul m. nguyen contributed to the design and assisted with the experiment presented in supplemental figure 4g/h. paul’s name was inadvertently left off the manuscript when it was re-submitted. this was an error, and his name should have been included to recognise his contribution. paul’s initials were included in the author contributions, it is only his name in the author list that is missing from the publication. Paul M. Nguyen should be added to position 9 of the author list on the manuscript. The affiliations SHOULD BE: 3. University of Melbourne Centre for Cancer Research, Victorian Comprehensive Cancer Centre, Parkville, VIC, 3000, Australia. 4. Department of Clinical Pathology, University of Melbourne, Parkville, VIC, 3000, Australia.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Low, R.R.J., Fung, K.Y., Gao, H. et al. S100 family proteins are linked to organoid morphology and EMT in pancreatic cancer. Cell Death Differ 30, 1155–1165 (2023). https://doi.org/10.1038/s41418-023-01126-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-023-01126-z
- Springer Nature Limited
This article is cited by
-
Unbiasedly decoding the tumor microenvironment with single-cell multiomics analysis in pancreatic cancer
Molecular Cancer (2024)
-
OrganoIDNet: a deep learning tool for identification of therapeutic effects in PDAC organoid-PBMC co-cultures from time-resolved imaging data
Cellular Oncology (2024)
-
A systematic review on the culture methods and applications of 3D tumoroids for cancer research and personalized medicine
Cellular Oncology (2024)