Abstract
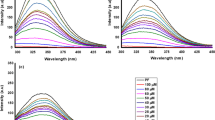
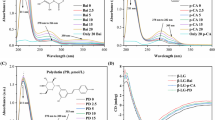
β-Lactoglobulin (β-LG), the most important whey protein, has high capacity for binding many small hydrophobic molecules. The binding properties of β-LG in the presence of four different synthetic curcumin-based derivatives as potent inhibitors of intestinal α-glucosidase (α-Gls) were investigated. To study the binding properties both UV–vis and florescence spectroscopy were used accompanied by molecular modeling and simulation. The formed complex between curcumin and β-LG was investigated using spectroscopic techniques. The spectroscopic analyses results propose acceptable binding affinities between the curcumin derivatives and β-LG. Also, it was proved that the hydrophobic interactions play a noteworthy role in the protein–ligand binding. All of the curcumin inhibitors showed high affinity to β-LG based on the structural and thermodynamic analyses. Binding constant of complex formation (K) and thermodynamic parameters (ΔG°, ΔH° and ΔS°) of complex formation were calculated. The results suggested that the process was endothermic and entropy driven. Moreover, our results indicated that protein–ligand binding was spontaneous signifying the presence of static quenching between the two molecules. Our finding suggests the major role of hydrophobic effect in the interaction of the synthetic compounds and β-LG. Moreover, molecular docking analyses exposed that curcumin-based ligands bind with high affinity to the protein in a hydrophobic pocket which located on the protein surface. The stability of ligand–protein interaction was further confirmed by molecular dynamics simulation analysis. This study suggests β-LG as a good carrier for these potentially curcumin-based antidiabetic compounds to deliver safely through the stomach to small intestine which is the crucial site of their activity.











Similar content being viewed by others

Abbreviations
- β-LG:
-
β-Lactoglobulin
- ANS:
-
1-Anilinonaphthalene-8-Sulfonic acid
- MVD:
-
Molegro Virtual Docker
- PDB:
-
Protein data bank
- CGenFF:
-
CHARMM general force field
- GROMACS:
-
Groningen machine for chemical simulation
- PME:
-
Particle mesh ewald
- RMSD:
-
Rout mean square deviation
- RMSF:
-
Root mean square fluctuation
References
G. Kontopidis, C. Holt, L. Sawyer, The ligand-binding site of bovine β-lactoglobulin: evidence for a function. J. Mol. Biol. 318(4), 1043–1055 (2002)
S. Brownlow, J.H. Morais Cabral, R. Cooper, D.R. Flower, S.J. Yewdall, I. Polikarpov, A.C.T. North, L. Sawyer, Bovine β-lactoglobulin at 18 Å resolution—still an enigmatic lipocalin. Structure 5(4), 481–495 (1997)
K. Takagi, R. Teshima, H. Okunuki, J.I. Sawada, Comparative study of in vitro digestibility of food proteins and effect of preheating on the digestion. Biol. Pharm. Bull. 26(7), 969–973 (2003)
L. Liang, H. Tajmir-Riahi, M. Subirade, Interaction of β-lactoglobulin with resveratrol and its biological implications. Biomacromol 9(1), 50–56 (2007)
L. Sawyer, G. Kontopidis, The core lipocalin, bovine β-lactoglobulin. Biochim. Biophys. Acta (BBA) Protein Struct. Mol. Enzymol. 1482(1–2), 136–148 (2000)
T. Lefèvre, M. Subirade, Conformational rearrangement of β-lactoglobulin upon interaction with an anionic membrane. Biochim. Biophys. (BBA) Protein Struct. Mol. Enzymol. 1549(1), 37–50 (2001)
H.L. Monaco, G. Zanotti, P. Spadon, Crystal structure of the trigonal form of bovine beta-lactoglobulin and of its complex with retinol at 2.5 Å resolution. J. Mol. Biol. 197(4), 695–706 (1987)
G. Kontopidis, C. Holt, L. Sawyer, Invited review: β-lactoglobulin: binding properties, structure, and function. J. Dairy Sci. 87(4), 785–796 (2004)
M.H. Mehraban, R. Yousefi, A. Taheri-Kafrani, F. Panahi, A. Khalafi-Nezhad, Binding study of novel anti-diabetic pyrimidine fused heterocycles to β-lactoglobulin as a carrier protein. Colloids Surf. B 112, 374–379 (2013)
P. Busti, C.A. Gatti, N.J. Delorenzi, Some aspects of β-lactoglobulin structural properties in solution studied by fluorescence quenching. Int. J. Biol. Macromol. 23(2), 143–148 (1998)
A. Divsalar, A.A. Saboury, H. Mansoori-Torshizi, M. Islami Moghaddam, F. Ahmad, G.H. Hakimelahi, Comparative studies on the interaction between bovine β-lacto-globulin type a and B and a new designed Pd (II) complex with anti-tumor activity at different temperatures. J. Biomol. Struct. Dyn. 26(5), 587–597 (2009)
A.K. Bordbar, N. Sohrabi, H. Gharibi, Binding set analysis for interaction of human serum albumin with cethyl trimethylammonium bromide. Bull. Korean Chem. Soc. 25(6), 791–795 (2004)
B.R. Chandrika, J. Subramanian, S.D. Sharma, Managing protein flexibility in docking and its applications. Drug Discov. Today 14(7–8), 394–400 (2009)
L.J. Shai, P. Masoko, M.P. Mokgotho, S.R. Magano, A.M. Magale, N. Boaduo, J.N. Eloff, Yeast alpha glucosidase inhibitory and antioxidant activities of six medicinal plants collected in Phalaborwa, South Africa. South Afr. J. Bot. 76(3), 465–470 (2010)
V. Ghadyale, S.H. Takalikar, V. Haldavnekar, A. Arvindekar, Effective control of postprandial glucose level through inhibition of intestinal alpha glucosidase By Cymbopogon martinii (Roxb). Evid. Based Complement. Altern. Med. 2012, e372909 (2012)
S.D. Kim, α-Gls inhibitor from Buthus martensi Karsch. Food Chem. 136(2), 297–300 (2013)
Y. Liu, L. Ma, W. Chen, H. Park, Z. Ke, B. Wang, Binding mechanism and synergetic effects of xanthone derivatives as noncompetitive α-Gls inhibitors: a theoretical and experimental study. J. Phys. Chem. B 117(43), 13464–13471 (2013)
F. Panahi, R. Yousefi, M.H. Mehraban, A. Khalafi-Nezhad, Synthesis of new pyrimidine-fused derivatives as potent and selective antidiabetic α-Gls inhibitors. Carbohydr. Res. 380, 81–91 (2013)
V.P. Menon, A.R. Sudheer, Antioxidant and anti-inflammatory properties of curcumin. Mol. Targets Ther. Uses Curcumin Health Dis. 595, 105–125 (2007)
H. Hatcher, R. Planalp, J. Cho, F.M. Torti, S.V. Torti, Curcumin: from ancient medicine to current clinical trials. Cell. Mol. Life Sci. 65(11), 1631–1652 (2008)
R.A. Kowluru, M. Kanwar, Effects of curcumin on retinal oxidative stress and inflammation in diabetes. Nutr. Metab. 4(1), 8 (2007)
A. Yousefi, R. Yousefi, F. Panahi, S. Sarikhani, A.R. Zolghadr, A. Bahaoddini, A. Khalafi-Nezhad, Novel curcumin-based pyrano [2, 3-d] pyrimidine anti-oxidant inhibitors for α-amylase and α-Gls: Implications for their pleiotropic effects against diabetes complications. Int. J. Biol. Macromol. 78, 46–55 (2015)
A. Kumar, G. Das, B. Bose, Recombinant receptor-binding domain of diphtheria toxin increases the potency of curcumin by enhancing cellular uptake. Mol. Pharm. 11(1), 208–217 (2013)
S.C. Gupta, S. Prasad, J.H. Kim, S. Patchva, L.J. Webb, I.K. Priyadarsini, B.B. Aggarwal, Multitargeting by curcumin as revealed by molecular interaction studies. Natl. Prod. Rep. 28(12), 1937–1955 (2011)
A.H. Sneharani, J.V. Karakkat, S.A. Singh, A.G. Appu Rao, Interaction of curcumin with β-lactoglobulin- stability, spectroscopic analysis, and molecular modeling of the complex. J. Agric. Food Chem. 58(20), 11130–11139 (2010)
P. Mailliart, B. Ribadeau-Dumas, Preparation of β-lactoglobulin and p-lactoglobulin-free proteins from whey retentate by NaCI salting out at low pH. J. Food Sci. 53(3), 743–745 (1988)
P. Bourassa, S. Dubeau, G.M. Maharvi, A.H. Fauq, T.J. Thomas, H.A. Tajmir-Riahi, Binding of antitumor tamoxifen and its metabolites 4-hydroxytamoxifen and endoxifen to human serum albumin. Biochimie 93(7), 1089–1101 (2011)
F. Mohammadi, A.K. Bordbar, A. Divsalar, K. Mohammadi, A.A. Saboury, Interaction of curcumin and diacetylcurcumin with the lipocalin member β-lactoglobulin. Protein J. 28(3–4), 117–123 (2009)
G. Zhang, X. Chen, J. Guo, J. Wang, Spectroscopic investigation of the interaction between chrysin and bovine serum albumin. J. Mol. Struct. 921(1–3), 346–351 (2009)
Y.J. Hu, Y. Liu, J. Wang, X. **ao, S. Qu, Study of the interaction between monoammonium glycyrrhizinate and bovine serum albumin. J. Pharm. Biomed. Anal. 36(4), 915–919 (2004)
F. Ding, W. Liu, J.X. Diao, B. Yin, L. Zhang, Y. Sun, Complexation of insecticide chlorantraniliprole with human serum albumin: biophysical aspects. J. Lumin. 131(7), 1327–1335 (2011)
K. Chen, T. Li, T. Cao, Tribe-PSO: a novel global optimization algorithm and its application in molecular docking. Chemom. Intell. Lab. Syst. 82(1–2), 248–259 (2006)
M. Xu, M.A. Lill, Utilizing experimental data for reducing ensemble size in flexible-protein docking. J. Chem. Inf. Model. 52(1), 187–198 (2011)
R. Dias, W. Filgueira, Jr. de Azevedo, Molecular docking algorithms. Curr. Drug Targets 9(12), 1040–1047 (2008)
R. Thomsen, M.H. Christensen, MolDock: a new technique for high-accuracy molecular docking. J. Med. Chem. 49(11), 3315–3321 (2006)
M.D. Hanwell, D.E. Curtis, D.C. Lonie, T. Vandermeersch, E. Zurek, G.R. Hutchison, Avogadro: an advanced semantic chemical editor, visualization, and analysis platform. J. Cheminf. 4(1), 17 (2012)
F. Keshavarz, D. Mohammad-Aghaie, Dual-target anticancer drug candidates: Rational design and simulation studies. Phys. Chem. Res. 3(2), 125–143 (2015)
E. Dufour, C. Genot, T. Haertlé, β-Lactoglobulin binding properties during its folding changes studied by fluorescence spectroscopy. Biochim. Biophys. Acta (BBA) Protein Struct. Mol. Enzymol. 1205(1), 105–112 (1994)
K. Vanommeslaeghe, E.P. Raman, A.D. MacKerell Jr., Automation of the CHARMM General Force Field (CGenFF) II: assignment of bonded parameters and partial atomic charges. J. Chem. Inf. Model. 52(12), 3155–3168 (2012)
K. Vanommeslaeghe, A.D. MacKerell Jr., Automation of the CHARMM General Force Field (CGenFF) I: bond perception and atom ty**. J. Chem. Inf. Model. 52(12), 3144–3154 (2012)
M.J. Abraham, T. Murtola, R. Schulz, S. Pall, J.C. Smith, B. Hess, E. Lindahl, GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015)
J. Huang, A.D. MacKerell, CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 34(25), 2135–2145 (2013)
H.J. Berendsen, J.P.M. Postma, W.F. van Gunsteren, A. DiNola, J.R. Haak, Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81(8), 3684–3690 (1984)
M. Parrinello, A. Rahman, Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52(12), 7182–7190 (1981)
T. Darden, D. York, L. Pedersen, Particle mesh ewald: an N⋅ log (N) method for Ewald sums in large systems. J. Chem. Phys. 98(12), 10089–10092 (1993)
B. Hess, H. Bekker, H.J.C. Berendsen, J.G.E.M. Fraaije, LINCS: a linear constraint solver for molecular simulations. J. Comput. Chem. 18(12), 1463–1472 (1997)
E.F. Pettersen, T.D. Goddard, C.C. Huang, G.S. Couch, D.M. Greenblatt, E.C. Meng, T.E. Ferrin, UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25(13), 1605–1612 (2004)
R.A. Laskowski, M.B. Swindells, LigPlot+: multiple ligand–protein interaction diagrams for drug discovery. J. Chem. Inf. Model. 51(10), 2778–2786 (2011)
X. Daura, K. Gademann, B. Jaun, D. Seebach, W.F. Gunsteren, A.L. Mark, Peptide folding: when simulation meets experiment. Angew. Chem. Int. Ed. 38(1–2), 236–240 (1999)
J.M. Bland, D.G. Altman, Applying the right statistics: analyses of measurement studies. Ultrasound Obstet. Gynecol. 22(1), 85–93 (2003)
M.Z. Papiz, L. Sawyer, E.E. Eliopoulos, A.C.T. North, J.B.C. Findlay, R. Sivaprasadarao, T.A. Jones, M.E. Newcomer, P.J. Kraulis, The structure of β-lactoglobulin and its similarity to plasma retinol-binding protein. Nature 324(6095), 383–385 (1986)
S. Le Maux, S. Bouhallab, L. Giblin, A. Brodkorb, T. Croguennec, Bovine β-lactoglobulin/fatty acid complexes: binding, structural, and biological properties. Dairy Sci. Technol. 94(5), 409–426 (2014)
M.C. Yang, H.H. Guan, M.Y. Liu, Y.H. Lin, J.M. Yang, W.L. Chen, C.J. Chen, S.J.T. Mao, Crystal structure of a secondary vitamin D3 binding site of milk β-lactoglobulin. Proteins Struct. Funct. Bioinf. 71(3), 1197–1210 (2008)
S.-Y. Wu, M.D. Perez, P. Puyol, L. Sawyer, β-Lactoglobulin binds palmitate within its central cavity. J. Biol. Chem. 274(1), 170–174 (1999)
I. Hasni, P. Bourassa, H.A. Tajmir-Riahi, Binding of cationic lipids to milk β-lactoglobulin. J. Phys. Chem. B 115(20), 6683–6690 (2011)
Acknowledgements
The authors appreciatively acknowledge the financial support of Shiraz University Research Council, Iran National Science foundation (INSF).
Funding
This work was supported by INSF (Grant number 960084).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yousefi, A., Ahrari, S., Panahi, F. et al. Binding analysis of the curcumin-based synthetic alpha-glucosidase inhibitors to beta-lactoglobulin as potential vehicle carrier for antidiabetic drugs. J IRAN CHEM SOC 19, 489–503 (2022). https://doi.org/10.1007/s13738-021-02323-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13738-021-02323-8