Abstract
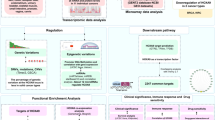
JAK/STAT pathway has been widely acknowledged in the development of human cancers. However, the role of different phosphorylated STAT proteins translocating into nucleus in transcription activation of target genes is not fully understood. In present research, ChIP-seq was carried on to investigate the genome-wide distribution of the activated STAT1, STAT2, STAT3, STAT5 and STAT6 in colorectal cancer HCT-116 cells. Our observations indicated that the homodimers rather than heterodimers of STAT protein predominantly occupied on genomic DNA. STAT3 accounted for the largest proportion among all STAT proteins HCT-116 cells. Furthermore, the biased binding motif targeted by different STAT homodimers suggested the distinct biological functions. Here, we noticed that NR5A2 was a specific co-activator of STAT3 by DNA motif analysis. Co-IP assay determined that NR5A2 indeed interacted with STAT3 homodimer rather than other homodimers or heterodimers. NR5A2 knockdown resulted in a reduced binding affinity of STAT3 homodimer in the original regions. Taken together, we characterize the genome-wide landscape of activated STAT proteins, and reveal the differences of binding patterns as well as the target genes and associated functions between homodimer and heterodimer of STAT proteins in HCT-116 cells. We also present some new findings and possible mechanisms regarding the role of NR5A2 on STAT3 in CRC. Our findings may provide new insights into the design of STAT inhibitors to treat CRC and other diseases.






Similar content being viewed by others
References
Vainchenker W, Constantinescu SN. JAK/STAT signaling in hematological malignancies. Oncogene. 2013;32:2601–13.
Groner B, von Manstein V. Jak Stat signaling and cancer: Opportunities, benefits and side effects of targeted inhibition. Mol Cell Endocrinol. 2017;451:1–14.
Darnell JE Jr. STATs and gene regulation. Science. 1997;277:1630–5.
Otero-Muras I, Yordanov P, Stelling J. Chemical reaction network theory elucidates sources of multi-stability in interferon signaling. PLoS Comput Biol. 2017;13: e1005454.
Zhang Y, Zhang Y, Yun H, Lai R, Su M. Correlation of STAT1 with apoptosis and cell-cycle markers in esophageal squamous cell carcinoma. PLoS ONE. 2014;9: e113928.
Sellier H, Rebillard A, Guette C, Barre B, Coqueret O. How should we define STAT3 as an oncogene and as a potential target for therapy? Jak-Stat. 2013;2: e24716.
Good SR, Thieu VT, Mathur AN, et al. Temporal induction pattern of STAT4 target genes defines potential for Th1 lineage-specific programming. J Immunol. 2009;183:3839–47.
Walford HH, Doherty TA. STAT6 and lung inflammation. Jak-Stat. 2013;2: e25301.
Heltemes-Harris LM, Farrar MA. The role of STAT5 in lymphocyte development and transformation. Curr Opin Immunol. 2012;24:146–52.
Murray PJ. The JAK-STAT signaling pathway: input and output integration. J Immunol. 2007;178:2623–9.
Tang S, Yuan X, Song J, Chen Y, Tan X, Li Q. Association analyses of the JAK/STAT signaling pathway with the progression and prognosis of colon cancer. Oncol Lett. 2019;17:159–64.
Wang SW, Sun YM. The IL-6/JAK/STAT3 pathway: potential therapeutic strategies in treating colorectal cancer (Review). Int J Oncol. 2014;44:1032–40.
Slattery ML, Lundgreen A, Kadlubar SA, Bondurant KL, Wolff RK. JAK/STAT/SOCS-signaling pathway and colon and rectal cancer. Mol Carcinog. 2013;52:155–66.
Dariya B, Muppala S, Srivani G, Momin S, Alam A, Saddala MS. Targeting STAT proteins via computational analysis in colorectal cancer. Mol Cell Biochem. 2021;476:165–74.
Zhou S, Shen Y, Zang S, Yin X, Li P. The epigenetic role of HTR1A antagonist in facilitaing GnRH expression for pubertal initiation control. Mol Ther Nucleic Acids. 2021;25:198–206.
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y. The I-TASSER Suite: protein structure and function prediction. Nat Methods. 2015;12:7–8.
Pierce BG, Wiehe K, Hwang H, Kim BH, Vreven T, Weng Z. ZDOCK server: interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics. 2014;30:1771–3.
Jiang L, Zhao XH, Mao YL, Wang JF, Zheng HJ, You QS. Long non-coding RNA RP11–468E2.5 curtails colorectal cancer cell proliferation and stimulates apoptosis via the JAK/STAT signaling pathway by targeting STAT5 and STAT6. J Exp Clin Cancer Res. 2019;38:465.
Murata T, Noguchi PD, Puri RK. IL-13 induces phosphorylation and activation of JAK2 Janus kinase in human colon carcinoma cell lines: similarities between IL-4 and IL-13 signaling. J Immunol. 1996;156:2972–8.
Luan J, Fu J, Wang D, et al. miR-150-Based RNA interference attenuates Tubulointerstitial fibrosis through the SOCS1/JAK/STAT pathway in vivo and in vitro. Mol Ther Nucleic Acids. 2020;22:871–84.
Pu Z, Xu M, Yuan X, **e H, Zhao J. Circular RNA circCUL3 accelerates the Warburg effect progression of gastric cancer through regulating the STAT3/HK2 Axis. Mol Ther Nucleic Acids. 2020;22:310–8.
Lim CP, Cao X. Structure, function, and regulation of STAT proteins. Mol BioSyst. 2006;2:536–50.
Decker T, Kovarik P. Transcription factor activity of STAT proteins: structural requirements and regulation by phosphorylation and interacting proteins. Cell Mol Life Sci. 1999;55:1535–46.
Fernandez-Marcos PJ, Auwerx J, Schoonjans K. Emerging actions of the nuclear receptor LRH-1 in the gut. Biochem Biophys Acta. 2011;1812:947–55.
Shen Y, Zhao H, Zhang L, et al. The roles of DNA methylation and hydroxymethylation at short interspersed nuclear elements in the hypothalamic arcuate nucleus during puberty. Mol Ther Nucleic Acids. 2021;26:242–52.
La Sala G, Michiels C, Kukenshoner T, et al. Selective inhibition of STAT3 signaling using monobodies targeting the coiled-coil and N-terminal domains. Nat Commun. 2020;11:4115.
Zhang T, Kee WH, Seow KT, Fung W, Cao X. The coiled-coil domain of Stat3 is essential for its SH2 domain-mediated receptor binding and subsequent activation induced by epidermal growth factor and interleukin-6. Mol Cell Biol. 2000;20:7132–9.
Acknowledgements
This work was supported by Science and Technology Development Plan of Suzhou (Medical and Health Science and Technology Innovation) (SKYD2022023), Start-up Funding of Suzhou Ninth People’s Hospital, Youth Program of Develo** Public Health through Science and Education of Suzhou (KJXW2019069), Program of Develo** Public Health through Science and Education of Wujiang District (wwk201811), The People’s Livelihood Science and Technology of Suzhou (SYSD2020043), Suzhou science and technology planning project (SKJY2021016), Joint Co-construction Project of Henan Medical Science and Technology Research Plan (LHGJ20210199), Key R & D and promotion projects in Henan Province of 2020 (scientific and technical program) (222102310266). We are grateful for the assistance from Shanghai Genefund Biotech Co. Ltd. in the generation and analysis of high-throughput sequencing data. We are also grateful for Coweldgen Scientific Co. Ltd. for STR authentication of cell lines.
Author information
Authors and Affiliations
Contributions
HY and SYF performed the cellular and molecular experiments and analyzed data. HY and ZY provided the clinical samples. TY and GY helped cell culture and molecular experiments. LC helped bioinformatic analysis. YT provided the major financial support for the project. SYH designed the research route, and was responsible for quality control of all raw data, and drafted and revised the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declared that they have no conflicts of interest to this work. The datasets and supporting materials generated during and/or analysis during the current study are available from the corresponding author on reasonable request.
Ethical approval
The research protocol was approved by Ethics Committee of Henan Cancer Hospital (2022-KY-0073-001). The experiments were conducted in accordance with the Declaration of Helsinki (2000 version).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13577_2022_815_MOESM1_ESM.xlsx
Supplementary file1 Table S1-5. The genomic annotation of STAT1 (S1), STAT2 (S2), STAT3 (S3), STAT5 (S4) and STAT6 (S5) in HCT116 cells. Column E lists the nearby gene element if exists. Column I-K list the information of probable typical enhancer (TE) and super-enhancer (SE) of HCT116 cells based from SEdbV1.05 Database. (XLSX 5445 KB)
13577_2022_815_MOESM6_ESM.xlsx
Supplementary file6 Table S6. The genomic annotation of binding areas with probable dimers with different STAT combinations. In Column D-H, the positive (“1”) and negative (“0”) abundance of the given protein are listed. In Column I, number means how many kind of STAT proteins bind on this area. In Column J to AD, difference of binding abundance between two STAT proteins are compared and shown by fold change and FDR. (XLSX 5909 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Hu, Y., Shen, Y., Zhao, Y. et al. Genomic distribution of signal transducer and activator of transcription (STAT) family in colorectal cancer. Human Cell 36, 286–295 (2023). https://doi.org/10.1007/s13577-022-00815-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13577-022-00815-0