Abstract
The Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) is related with the COVID-19 pandemic. Recent spike protein variations have had an effect on the transmission of the virus. In addition to ACE-2, spike proteins can employ DC-SIGN and its analogous receptor, DC-SIGNR, for host evasion. Spike variations in the DC-SIGN interaction region and role of DC-SIGN in immune evasion have not been well defined. To understand the spike protein variations and their binding mode, phylogenetic analysis of the complete GISAID (Global Initiative for Sharing Avian Influenza Data) data of the SARS-CoV-2 spike protein was considered. In addition, an in silico knockout network evaluation of the SARS-CoV-2 single-cell transcriptome was conducted to determine the key role of DC-SIGN/R in immunological dysregulation. Within the DC-SIGN-interacting region of the SARS-CoV spike protein, the spike protein of SARS-CoV-2 displayed remarkable similarity to the SARS-CoV spike protein. Surprisingly, the phylogenetic analysis revealed that the SARS-CoV-2’s spike exhibited significantly diverse variants in the DC-SIGN interaction domain, which altered the frequency of these variants. The variation within the DC-SIGN-interacting domain of spike proteins affected the binding of a limited number of variants with DC-SIGN and DC-SIGNR and affected their evolution. MMGBSA binding free energies evaluation differed for variants from those of the wild type, suggesting the influence of substitution mutations on the interaction pattern. In silico knockout network analysis of the single-cell transcriptome of Bronchoalveolar Lavage and peripheral blood mononuclear cells revealed that SARS-CoV-2 altered DC-SIGN/R signaling. Early surveillance of diverse SARS-CoV-2 strains could preclude a worsening of the pandemic and facilitate the development of an optimum vaccine against variations. The spike Receptor Binding Domain genetic variants are thought to boost SARS CoV-2 immune evasion, resulting in its higher longevity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
A novel contiguous virus was found in Wuhan city of China, in late December 2019. It belongs to the coronavirus and is classified as SARS-CoV-2 [1]. It has infected billions of people and caused millions of deaths across the globe and poses a risk to human life. a pandemic situation was declared on March 11, 2020 (WHO) [2, 3]. A pandemic infection on March 11th, 2020 [2, 4]. SARS-CoV-2 is the seventh virus known to infect humans date [3, 5], and it is a member of the beta coronavirus family [3], which possesses single-stranded RNA such as SARS CoV and MERS. It reduces lung function [6] by causing vascular thrombosis, endothelial damage, and microangiopathy. The fusing of cellular membranes is a crucial stage in viral infection [6]. The fusing of cellular membranes is a crucial stage in viral infection. The spike (S) protein, which consists of 1273 amino acids and is a transmembrane fusion glycoprotein [7], is essential for membrane fusion. It has two domains named S1 and S2 [4, 6, 8], and a region between amino acids 682 and 685 where furin can disassemble it. Angiotensin-converting enzyme 2 (ACE2) is the major receptor for SARS-CoV spike [9] and SARS-CoV-2 [10], which regulates how the virus travels from one cell to the next [11]. In addition, the receptor binding domain (RBD) amino acid alteration improved spike protein stability and transmission antigenicity [12]. Nevertheless, the low expression of ACE-2 on a subset of lung epithelial and endothelial cells demonstrates that additional receptors are involved in infection and entrance [11, 13,14,15,16]. Additionally, alterations in the RBD of spike protein impact the propagation of the virus [17]. Spike N501Y (U.K. variant B.1.1.7) [18] and D614G have recently found, with the potential to increase ACE2 recognition and effect the considerable dissemination of these variants across different populations [19]. The D614G variant had a high transmission rate, indicating significant viremia in COVID patients but it had no effect on disease severity [19].
DC-SIGN and DC-SIGNR are member of c-type lectins protein family [20] which recognizes carbohydrates. They function as adhesion and pathogen pattern-recognition receptors (PRR) that identify high mannose and complex glycan substrates and influence viral spread [20]. These interactions disrupt the signaling of other PRRs, such as TLRs, and induce immunological regulation. The infection of SARS CoV-2 begins by recognizing ACE2 [8, 15]. Utilizing auxiliary receptors restricts virus transmission, though. CD209L (DC-SIGNR) and CD209 (DC-SIGN) are known as alternative receptors for SARS CoV, SARS CoV-2, and HIV-1 to thrive on the host [13]. DC-SIGN/CD209 is found on PBMCs [19] such as dendritic cells and tissue-specific alveolar and dermal macrophages [20]. Similarly, DC-SIGNR is strongly expressed on type II alveolar lung epithelium and on the endothelium surface of the liver, lung, and lymph node [20]. Despite 77% similarities, their glycan recognition profiles are distinct. [21]. Moreover, DC-SIGN/R participation in infection of SARS-CoV-2 was previously demonstrated [13]. The correlation between DC-SIGN expression and disease severity suggests a potential target for SARS-CoV-2. Therefore, analyzing spike variations in regions where DC-SIGN/DC-SIGNR interact, may be essential for understanding how infections and evasions arise.
Our study describes the region of the SARS CoV-2 spike RBD where DC-SIGN and DC-SIGNR interacted and their variants. Based on the amino acid sequence, the spike RBD domain interacts variably with DC-SIGN/DC-SIGNR in GISAID sequences from 2020 to 2021. Also, the link between amino acid substitution in spike RBD, evolution, and binding was examined. Spike protein variations had unique molecular interactions with DC-SIGN/R, but not ACE2. In addition, we found variation in their binding free energy and per residue contribution utilizing MMGBSA of HawkDock [22]. By analyzing the single-cell transcriptomes of BALF and PBMC from SARS-CoV-2 patients, we further showed that DC-SIGN/R influenced the propagation of network signals. The diverse variations may boost fitness by making infections in several organs more likely to cause their failure. The development of new treatments to limit the spread of covid infection may depend on the identification of spike RBD variants and the manner in which they evolve.
Methods
Retrieval of DC-SIGN structure and active site identification
The 3D structures of DC-SIGN, DC-SIGNR (CD209/L) and ACE2 (PDB ID: 2XR5 1XPH and 6M17) were retrieved from the Protein Data Bank. The models were refined and minimized using the protein preparation wizard tool of Maestro in Schrodinger’s software suite. The interacting residues of DC-SIGN, DC-SIGNR and ACE2 were obtained from literature [23]. Specifically, residues N365, D366, N367, K368, F313, F374, D320, L321, Q323, G325, T326, and W327 comprise the active site of DC-SIGN and DC-SIGNR, and amino-acid stretches 30–41, 82–84 and 353–357 form the ACE2 active site.
Sequence alignment and modeling of spike protein
Primary amino acid sequence of the spike glycoprotein of SARS CoV (Accession no. AAR86788.1) and SARS CoV-2 (YP_009724390.1) was retrieved from the NCBI database. A model of SARS-CoV-1 and SARS-CoV-2 was generated using the SWISS-MODEL server. The SARS-CoV (PDB ID: 2GHV) was used as a template because it shares 99.89% sequence similarity with SARS-CoV-2. The reliable models were recognized based on GMQE and QMEAN [24]. GMQE represents the alignment and coverage of a template to query sequence, and QMEAN indicates the similarity of a model to the experimentally solved protein structures. The stereochemical property of models were evaluated by the Ramachandran plot generated using PROCHECK [25]. Moreover, quality of the generated models was evaluated using the Mol Probity scores. Mol Probity score provides an estimate of the computed sum of clashes, unfavored residues in the Ramachandran plot (%), and bad rotamers (%). A low numerical score signifies considerable quality of a particular model. It exhibits the resolution of structure at which the corresponding values might be applicable. Prior to studying protein–protein interaction, 3D models were further refined and energy minimized using Schrodinger’s Maestro protein preparation wizard [26]. The sequence of spike glycoprotein of SARS CoV and SARS CoV-2 was aligned by using Clustal W [27] and the percentage identity between the two was determined. Next, Multiple sequence alignment (MSA) was employed to evaluate the potential DC-SIGN binding residues in SARS-CoV-2 Spike RBD using SARS CoV and DC-SIGN interacting residues as a template.
Mutational analysis of DC-SIGN/R binding region of spike protein
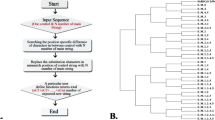
Mutations in SARS CoV-2 DC-SIGN/R binding region (337–399 amino acid) were analyzed for sequences available in GISAID from Oct 2020 to dec 2021. Mutations were analyzed using the GISAID CoV Server (www.gisaid.org/epiflu-applications/covsurver-mutations-app) with ‘hCoV-19/Wuhan/WIV04/2019’ (Accession number NC-045512.2). The information about the collected dataset is presented as Table S1. Noisy sequences were excluded from the analysis. The frequency of identified mutations was followed for a year (from oct 2020 to Oct 2021).
The 3D modeling of the mutated spike protein
FASTA file with altered amino acid sequences was submitted to the SWISS MODEL server to build mutated protein variants of the wild type spike protein (Wuhan SARS CoV-2). The stereochemical quality of the modeled mutants was verified using Procheck module of PDBsum tool [12, 28]. The original and mutant protein structures were viewed and studied using PyMOL visualization program [16].
Molecular docking of spike protein with DC-SIGN/R
Solving the crystal structures of interacting proteins is an inconvenient and laborious task. Several methods have been devised to study protein–protein modeling. The High Ambiguity Driven protein–protein Docking (HADDOCK) modeling approach is employed because it uses biochemical and biophysical data and provides the lowest energy structure [15, 16]. The protein–protein modeling was employed to study the intermolecular interaction of spike glycoprotein with DC-SIGN using HADDOCK 2.4 server [29]. For initiating docking, actively participating residues were provided to the HADDOCK 2.4 server [30]. The active residues of DC-SIGN were retrieved from literature [23] which encompasses calcium coordinated primary binding sites and secondary binding sites. The active site residues from 324 to 386 of spike SARS CoV were utilized. For SARS CoV-2 spike protein, the active site residues were predicted via sequence alignment of spike proteins of SARS CoV and SARS COV-2, that displays high sequence similarity [31] and the corresponding region was considered as the active site for the intermolecular interaction. The passive residues were generated automatically to a distance of 6.5A° around the protein.
The structural models were ranked based on their Van-der wall and electrostatic interactions, desolvation, Ambiguous interaction restraints (AIRs), and energies of the buried surface energies. The most favored model as indicated by the lowest HADDOCK score was further considered for analyzing details of the intermolecular interactions between SARS CoV-2 and DC-SIGN/R. Structural models with lowest energy score were analyzed and visualized using [32] PyMOL and Mapiya [33]. The intermolecular interacting residues across the proteins involved in interaction were recognized by PDB sum (http://www.ebi.ac.uk/pdbsum) and visualized as a pictorial interaction map [34]Mapiya visualizes the interaction using a contact map (CM) and a distance map (DM). Mapiya showed a unique feature of coloring and filtering contacts based on the physiochemical properties of interactions Mapiya uses the cutoff distance in angstroms between two heavy molecules of different residues to make the CM interaction map. The pair of residues is colored when the distance between the residues is shorter than a cutoff value. We used a filter for electrostatic and hydrophobic interactions to analyze their respective interactions. Complex 3D structures were displaced using Mapiya's built-in Molstar viewer [33].
Estimation of binding free energy of spike variant-DC-SIGN/R using MMGBSA/MMPBSA method
The free binding energy of intermolecular interaction of Spike protein and DC-SIGN/R was evaluated by utilizing MMGBSA (Molecular Mechanics Generalized Born Surface Area)/ /MMPBSA (Molecular mechanics Poisson Boltzmann surface area) which exhibits more crucial than scoring functions, through Hawk Dock web server [22]. Moreover, it provides the contribution of each residue in binding free energy of PP interaction and depicts the key residues which indicate the near native interaction and their binding affinity. MM/GBSA was employed to determine the interface residues of Spike protein and DC-SIGN/R using Hawk dock scoring used in Hawk Dock webserver.
Influence of point mutation on dynamic of wild type and Spike variants
The effect of mutation on the flexibility, structural conformation and molecular stability of the SARS-CoV-2 protein was investigated using the DynaMut software [35].DynaMut is a software server that uses well established normal mode techniques to analyze and visualize protein dynamics. By uploading the modelled spike structure to the DynaMut software, the influence of point mutations on several protein structure stability characteristics such as deformation energies, atomic fluctuations and vibrational entropy was studied. DynaMut calculates the vibrational entropy changes to determine the impact of mutations on protein stability and dynamics.
DC-SIGN regulatory network construction
The DC-SIGN regulatory network was constructed using the STRING v.10.5 database [36] and Cytoscape [36, 37]. In the constructed network, proteins with highest degree of binding in the network explains a key regulatory role as a hub. The topological properties of the network were calculated using Network Analyzer [38], a plugin of Cytoscape v3.7.2. The network was functionally annotated using Metascape.
Topological properties of PPI network
Topological analysis facilitates the comprehension of a network's structure, hence facilitating the comprehension of its hidden mechanisms. Betweenness (CB) and closeness characterize the centrality assessment of the PPI network of differentially regulated proteins (DEGs) (Cc). Using Network Analyzer, measurements of centrality were determined [38]. These topological characteristics were utilized to track the topological alterations brought about by network disruption. To explore the network's fundamental characteristics, the following network properties were considered.
Degree
In PPI network analysis, the degree k indicates the total number of links established by a network node. This is also used to determine the local significance of a node in the process of network control. G = (N, E) represents the graph, where N represents the nodes and E represents the edges. Betweenness and proximity centrality are the fundamental centrality metrics and the parameters for estimating a node's global functional importance in a network regulation [39].
Betweenness centrality
Betweenness centrality quantifies a node's participation in the quickest communication between other pairs of nodes in the network. This nodal metric, c B (u), represents the number of shortest routes that pass through a node u. (Eq. 1).
Betweenness centrality is the degree to which a node shares the shortest-path traffic from all viable routes through other nodes. Consequently, it is the parameter of a node's ability to benefit from the flow of information throughout the network [40] and to exert control over the signal processing of other nodes in the network [41].
Closeness centrality
Closeness centrality is a measure of a node's proximity (via short pathways) to other nodes in a network. This nodal measure CC (u) is the inverse of the sum of the distances between the node of interest u and every other node v \(\in\) V\u (Eq. 2).
The closeness centrality of a network indicates how quickly information spreads from one node to another related node [38]. The function of a node is reliant on the centralities of its neighbors and changes across networks of highly connected nodes [42]. It is regarded as a significant measure of a node's information transmission capability inside a network since the likelihood of node isolation is reduced within a zone of tightly connected nodes.
Integration of single cell sequence data from COVID-19 patients
The expression analysis of DC-SIGN and associated proteins were validated through a database portal (http://covid19.cancer-pku.cn/). This database is repository of single cell transcriptome dataset of BALF and PBMC from mild and severely infected patients. We retrieved the gene expression data of different cell type from http://covid19.cancer-pku.cn/. We visualized the expression of DC-SIGN and associated proteins as violin plot and heat map by using ggplot2.
Results
DC-SIGN/R interacts with a unique area of spike protein's receptor binding domain (RBD)
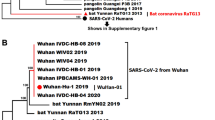
Analysis of the spike proteins of SARS-CoV and SARS-CoV-2 revealed that their amino acid sequences were 98.89% similar [18] and that eight conserved and semi-conserved areas interacted [3, Data is with the authors and will be provided on request through corresponding author. Angiotensin converting enzyme Bronchoalveolar lavage fluid Chemokine (C-X-C motif) ligand 1 Chemokine (C–C motif) ligand Dendritic cell specific integrin grabbing non-integrin Interleukin Kinase suppressor of ras Leucocyte specific protein Nuclear factor kappa light chain enhancer of activated B cells Receptor binding domain Lempp FA, et al. Lectins enhance SARS-CoV-2 infection and influence neutralizing antibodies. Nature. 2021;598:342–7. Sirohi PR, Gupta J, Somvanshi P, Prajapati VK, Grover A. Multiple epitope-based vaccine prediction against SARS-CoV-2 spike glycoprotein. J Biomol Struct Dyn. 2020;40:1. Ibrahim IM, Abdelmalek DH, Elshahat ME, Elfiky AA. COVID-19 spike-host cell receptor GRP78 binding site prediction. J Infect. 2020;80:554. Cucinotta D, Vanelli M. WHO declares COVID-19 a pandemic. Acta Biomed. 2020;91:157–60. Andersen KG, Rambaut A, Lipkin WI, Holmes EC, Garry RF. The proximal origin of SARS-CoV-2. Nat Med. 2020;26:450–2. Islam AB, Khan M. Lung transcriptome of a COVID-19 patient and systems biology predictions suggest impaired surfactant production which may be druggable by surfactant therapy. Sci Rep. 2020;10(1):1–6. Mcgonagle D. Since January 2020 Elsevier has created a COVID-19 resource centre with free information in English and Mandarin on the novel coronavirus COVID-19. The COVID-19 resource centre is hosted on Elsevier Connect, the company‘s public news and information. 2020. https://cdn.who.int/media/docs/default-source/whhd-2021/scientific-publications/2.jhi_5may2021.pdf?sfvrsn=6526a2a5_5. Lan J, et al. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature. 2020;581:215–20. Dimitrov DS. The secret life of ACE2 as a receptor for the SARS virus. Cell. 2003;115:652–3. Hoffmann M, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and Is blocked by a clinically proven protease inhibitor. Cell. 2020;181:271-280.e8. Stoffolano JG. Free information in english and mandarin on the novel coronavirus COVID- water chemistry and microbiology. Adv In Insect Phys. 2020;57:72–80. Hajazadeh F, et al. SARS-COV-2 RBD (receptor binding domain) mutations and variants (A sectional-analytical study). Microb Pathog. 2022;168:105595. Amraei R, et al. CD209L/L-SIGN and CD209/DC-SIGN act as receptors for SARS-CoV-2. ACS Cent Sci. 2021;7:1156–65. Soh WT et al. The N terminal domain of spike glycoprotein mediates SARS-CoV-2 infection by associating with L-SIGN and DC-SIGN. BioRxiv. 2020. https://doi.org/10.1101/2020.11.05.369264. Elfiky AA, Ibrahim IM. Host-cell recognition through Cs-GRP78 is enhanced in the new Omicron variant of SARS-CoV-2, in silico structural point of view. J Infect. 2022;84:722. Elfiky AA, Ibrahim IM. Host-cell recognition through GRP78 is enhanced in the new UK variant of SARS-CoV-2, in silico. J Infect. 2021;82:186. Dinnon KH, et al. A mouse-adapted model of SARS-CoV-2 to test COVID-19 countermeasures. Nature. 2020;586:560–6. Fratev F. The SARS-CoV-2 S1 spike protein mutation N501Y alters the protein interactions with both hACE2 and human derived antibody: A Free energy of perturbation study. BioRxiv. 2020. https://doi.org/10.1101/2020.11.05.369264. Luan B, Wang H, Huynh T. Enhanced binding of the N501Y-mutated SARS-CoV-2 spike protein to the human ACE2 receptor: insights from molecular dynamics simulations. FEBS Lett. 2021;595:1454–61. Jeffers SA, Tusell SM, Gillim-Ross L, Hemmila EM, Achenbach JE, Babcock GJ, Thomas Jr WD, et al. CD209L (L-SIGN) is a receptor for severe acute respiratory syndrome coronavirus. Proc Nat Acad Sci. 2004;101(44):15748–53. https://doi.org/10.1073/pnas.040381210. Amraei R, et al. CD209L/L-SIGN and CD209/DC-SIGN act as receptors for SARS-CoV-2. ACS Central Sci. 2021;7(7):1156–65. https://doi.org/10.1021/acscentsci.0c01537. Weng G, et al. HawkDock: a web server to predict and analyze the protein–protein complex based on computational docking and MM/GBSA. Nucleic Acids Res. 2019;47:W322. Lenza MP, et al. Structural characterization of N-Linked glycans in the receptor binding domain of the SARS-CoV-2 spike protein and their interactions with human lectins. Angewandte Chemie Int Ed. 2020;59:23763–71. Benkert P, Biasini M, Schwede T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics. 2011;27:343–50. Laskowski RA, MacArthur MW, Moss DS, Thornton JM. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26(2):283–91. https://doi.org/10.1107/S0021889892009944. Chen VB, et al. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Crystallogr D Biol Crystallogr. 2010;66:12–21. Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucl Acids Res. 1994;22(22):4673–80. https://doi.org/10.1093/nar/22.22.4673. Laskowski RA, et al. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Crystallogr. 1993;26:283–91. van Zundert GCP et al. The HADDOCK2.2 web server: user-friendly integrative modeling of biomolecular complexes. J Mol Biol. (2016); 428: 720–725 Dominguez C, Boelens R, Bonvin AMJJ. HADDOCK: a protein-protein docking approach based on biochemical or biophysical information. J Am Chem Soc. 2003;125:1731–7. Grifoni A, et al. A sequence homology and bioinformatic approach can predict candidate targets for immune responses to SARS-CoV-2. Cell Host Microbe. 2020;27:671-680.e2. Seeliger D, de Groot BL. Ligand docking and binding site analysis with PyMOL and autodock/Vina. J Comput Aided Mol Des. 2010;24:417–22. Badaczewska-Dawid AE, Nithin C, Wroblewski K, Kurcinski M, Kmiecik S. Mapiya contact map server for identification and visualization of molecular interactions in proteins and biological complexes. Nucleic Acids Res. 2022;50:W474–82. Laskowski RA, Jabłońska J, Pravda L, Vařeková RS, Thornton JM. PDBsum: Structural summaries of PDB entries. Protein Sci. 2018;27:129–34. Rodrigues CHM, Pires DEV, Ascher DB. DynaMut: predicting the impact of mutations on protein conformation, flexibility and stability. Nucleic Acids Res. 2018;46:W350–5. von Mering C, et al. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 2003;31:258–61. https://doi.org/10.1093/nar/gkg034. Shannon P, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504. Assenov Y, Ramírez F, Schelhorn SESE, Lengauer T, Albrecht M. Computing topological parameters of biological networks. Bioinformatics. 2008;24:282–4. Newman ME, Girvan M. Finding and evaluating community structure in networks. Phys Rev E. 2004;69(2):026113. Brandes U. A faster algorithm for betweenness centrality. J Math Sociol. 2001;25:163–77. Chirom K, Malik MZ, Mangangcha IR, Somvanshi P, Singh RB. Network medicine in ovarian cancer: topological properties to drug discovery. Brief Bioinfo. 2022;23(3):bbac085. Canright GS, Engø-Monsen K. Spreading on networks: a topographic view. Complexus. 2006;3:131–46. Ren X, et al. COVID-19 immune features revealed by a large-scale single-cell transcriptome atlas. Cell. 2021;184:1895-1913.e19. Amraei R, Yin W, Napoleon MA, Suder EL, Berrigan J, Zhao Q, Olejnik J, Chandler KB, **a C, Feldman J, Hauser BM. CD209L/L-SIGN and CD209/DC-SIGN act as receptors for SARS-CoV-2 and are differentially expressed in lung and kidney epithelial and endothelial cells. bioRxiv. 2021. https://doi.org/10.1101/2020.06.22.165803. Xue LC, Rodrigues JP, Kastritis PL, Bonvin AM, Vangone A. PRODIGY: a web server for predicting the binding affinity of protein–protein complexes Li. Bioinformatics. 2016;32:3676–8. Thépaut M, Luczkowiak J, Vivès C, Labiod N, Bally I, Lasala F, Grimoire Y, Fenel D, Sattin S, Thielens N, Schoehn G. DC/L-SIGN recognition of spike glycoprotein promotes SARS-CoV-2 trans-infection and can be inhibited by a glycomimetic antagonist. PLoS Pathogens. 2021;17(5):e1009576. Geijtenbeek TB, et al. DC-SIGN, a dendritic cell–specific HIV-1-binding protein that enhances trans-infection of T cells. Cell. 2000;100(5):587–97. Zhang L, Jackson CB, Mou H, Ojha A, Peng H, Quinlan BD, Rangarajan ES, Pan A, Vanderheiden A, Suthar MS, Li W. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nature Commun. 2020;11(1):6013. Jarvis CM, et al. Antigen structure affects cellular routing through DC-SIGN. Proc Natl Acad Sci U S A. 2019;116:14862–7. Wan S, et al. Characteristics of lymphocyte subsets and cytokines in peripheral blood of 123 hospitalized patients with 2019 novel coronavirus pneumonia (NCP). MedRxiv. 2020. https://doi.org/10.1101/2020.02.10.20021832. Pan Y, et al. SARS-CoV-2-specific immune response in COVID-19 convalescent individuals. Signal Trans Target Ther. 2021;6(1):256. Sharma S, Singh I, Haider S, Malik MZ, Ponnusamy K, Rai E. ACE2 homo-dimerization, human genomic variants and interaction of host proteins explain high population specific differences in outcomes of COVID19. BioRxiv. 2020. https://doi.org/10.1101/2020.04.24.050534. Gringhuis SI, Den Dunnen J, Litjens M, Van Der Vlist M, Geijtenbeek TB. Carbohydrate-specific signaling through the DC-SIGN signalosome tailors immunity to Mycobacterium tuberculosis, HIV-1 and Helicobacter pylori. Nat Immunol. 2009;10(10), 1081–8. J.G is thankful to the Department of Biotechnology (DBT), Government of India, for Fellowship (DBT/2016/JNU/480). All the authors are also grateful for the Department of Biotechnology, Jawaharlal Nehru University, New Delhi, India for providing research platform to conduct the research. AKR is thankful to IOE, University of Delhi for the research grant (FRP 2022). RC is thankful to SERB, Department of Science, Goverment of India for research grant (ECR/2015/000429). Authors declare there is no conflict of interest. All the authors have read and approved the manuscript in all respect for publication. Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations. Below is the link to the electronic supplementary material. Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law. Gupta, J., Malik, M.Z., Chaturvedi, M. et al. SARS CoV-2 spike protein variants exploit DC-SIGN/DC-SIGNR receptor for evolution and severity: an in-silico insight.
VirusDis. 34, 278–296 (2023). https://doi.org/10.1007/s13337-023-00820-3 Received: Accepted: Published: Issue Date: DOI: https://doi.org/10.1007/s13337-023-00820-3Data availability
Abbreviations
References
Acknowledgements
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Consent of publication
Additional information
Publisher's Note
Supplementary Information
Rights and permissions
About this article
Cite this article
Keywords