Abstract
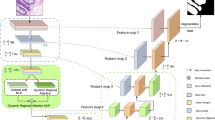
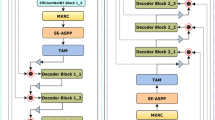
Deep learning has made important contributions to the development of medical image segmentation. Convolutional neural networks, as a crucial branch, have attracted strong attention from researchers. Through the tireless efforts of numerous researchers, convolutional neural networks have yielded numerous outstanding algorithms for processing medical images. The ideas and architectures of these algorithms have also provided important inspiration for the development of later technologies.Through extensive experimentation, we have found that currently mainstream deep learning algorithms are not always able to achieve ideal results when processing complex datasets and different types of datasets. These networks still have room for improvement in lesion localization and feature extraction. Therefore, we have created the dense multiscale attention and depth-supervised network (DmADs-Net).We use ResNet for feature extraction at different depths and create a Multi-scale Convolutional Feature Attention Block to improve the network’s attention to weak feature information. The Local Feature Attention Block is created to enable enhanced local feature attention for high-level semantic information. In addition, in the feature fusion phase, a Feature Refinement and Fusion Block is created to enhance the fusion of different semantic information.We validated the performance of the network using five datasets of varying sizes and types. Results from comparative experiments show that DmADs-Net outperformed mainstream networks. Ablation experiments further demonstrated the effectiveness of the created modules and the rationality of the network architecture.






























Similar content being viewed by others
Data availability
Data available on request from the authors.
References
Al-Dhabyani W, Gomaa M, Khaled H, Fahmy A (2019) Dataset of breast ultrasound images. Data Brief 28:104863
Alom MZ, Hasan M, Yakopcic C, Taha TM, Asari VK (2018) Recurrent residual convolutional neural network based on U-Net (R2U-Net) for medical image segmentation. ar**v preprint ar**v:1802.06955
Badrinarayanan V, Kendall A, Cipolla R (2017) SEGNet: a deep convolutional encoder–decoder architecture for image segmentation. IEEE Trans Pattern Anal Mach Intell 39(12):2481–2495
Chaitanya K, Erdil E, Karani N, Konukoglu E (2020) Contrastive learning of global and local features for medical image segmentation with limited annotations. Adv Neural Inf Process Syst 33:12546–12558
Chen H, Gu J, Zhang Z (2021) Attention in attention network for image super-resolution. ar**v preprint ar**v:2104.09497
Chen J, Lu Y, Yu Q, Luo X, Adeli E, Wang Y, Lu L, Yuille AL, Zhou Y (2021) TransUNet: Transformers make strong encoders for medical image segmentation. ar**v preprint ar**v:2102.04306
Chen LC, Papandreou G, Kokkinos I, Murphy K, Yuille AL (2017) DeepLab: semantic image segmentation with deep convolutional nets, atrous convolution, and fully connected CRFs. IEEE Trans Pattern Anal Mach Intell 40(4):834–848
Chen Y, Dai X, Liu M, Chen D, Yuan L, Liu Z (2020) Dynamic convolution: attention over convolution kernels. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 11030–11039
Darshik A, Dev A, Bharath M, Nair BA, Gopakumar G (2020) Semantic segmentation of spectral images: a comparative study using FCN8s and U-Net on RIT-18 dataset. In: 2020 11th international conference on computing, communication and networking technologies (ICCCNT). IEEE, pp 1–6
Ding Q, Shao Z, Huang X, Altan O (2021) DSA-Net: a novel deeply supervised attention-guided network for building change detection in high-resolution remote sensing images. Int J Appl Earth Obs Geoinf 105:102591
Dong Z, Li J, Hua Z (2024) Transformer-based multi-attention hybrid networks for skin lesion segmentation. Expert Syst Appl 244:123016
Fu B, Peng Y, He J, Tian C, Sun X, Wang R (2024) HMSU-Net: a hybrid multi-scale u-net based on a CNN and transformer for medical image segmentation. Comput Biol Med 170:108013
Gutman D, Codella NC, Celebi E, Helba B, Marchetti M, Mishra N, Halpern A (2016) Skin lesion analysis toward melanoma detection: a challenge at the international symposium on biomedical imaging (ISBI) 2016, hosted by the international skin imaging collaboration (ISIC). ar**s in deep residual networks. In: Computer Vision–ECCV 2016: 14th European conference, Amsterdam, The Netherlands, October 11–14, 2016, Proceedings, Part IV 14. Springer, pp 630–645
Hu J, Shen L, Sun G (2018) Squeeze-and-excitation networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 7132–7141
Hu X, Zeng D, Xu X, Shi Y (2021) Semi-supervised contrastive learning for label-efficient medical image segmentation. In: International conference on medical image computing and computer-assisted intervention. Springer, pp 481–490
Huang G, Liu Z, Van Der Maaten L, Weinberger KQ (2017) Densely connected convolutional networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 4700–4708
Ibtehaz N, Rahman MS (2020) MultiResUNet: rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural Netw 121:74–87
Jha D, Smedsrud PH, Riegler MA, Johansen D, De Lange T, Halvorsen P, Johansen HD (2019) ResUNet++: an advanced architecture for medical image segmentation. In: 2019 IEEE international symposium on multimedia (ISM). IEEE, pp 225–2255
Krizhevsky A, Sutskever I, Hinton GE (2017) ImageNet classification with deep convolutional neural networks. Commun ACM 60(6):84–90
Lee CY, **e S, Gallagher P, Zhang Z, Tu Z (2015) Deeply-supervised nets. In: Artificial intelligence and statistics. PMLR, pp 562–570
Li R, Zheng S, Zhang C, Duan C, Su J, Wang L, Atkinson PM (2021) Multiattention network for semantic segmentation of fine-resolution remote sensing images. IEEE Trans Geosci Remote Sens 60:1–13
Li S, Liu J, Song Z (2022) Brain tumor segmentation based on region of interest-aided localization and segmentation U-Net. Int J Mach Learn Cybern 13(9):2435–2445
Li X, Chen H, Qi X, Dou Q, Fu CW, Heng PA (2018) H-DenseUNet: hybrid densely connected UNet for liver and tumor segmentation from CT volumes. IEEE Trans Med Imaging 37(12):2663–2674
Li X, Wang W, Hu X, Yang J (2019) Selective kernel networks. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 510–519
Li Y, Yang H, Wang J, Zhang C, Liu Z, Chen H (2022) An image fusion method based on special residual network and efficient channel attention. Electronics 11(19):3140
Liu J, Zhang W, Tang Y, Tang J, Wu G (2020) Residual feature aggregation network for image super-resolution. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition, pp 2359–2368
Liu L, Huang Z, Liò P, Schönlieb CB, Aviles-Rivero AI (2022) PC-SwinMorph: patch representation for unsupervised medical image registration and segmentation. ar**v preprint ar**v:2203.05684
Liu X, Wang Z, Wan J, Zhang J, ** Y, Liu R, Miao Q (2023) RoadFormer: road extraction using a swin transformer combined with a spatial and channel separable convolution. Remote Sens 15(4):1049
Liu Z, Lin Y, Cao Y, Hu H, Wei Y, Zhang Z, Lin S, Guo B (2021) Swin transformer: hierarchical vision transformer using shifted windows. In: Proceedings of the IEEE/CVF international conference on computer vision, pp 10012–10022
Long J, Shelhamer E, Darrell T (2015) Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 3431–3440
Makandar A, Halalli B (2016) Threshold based segmentation technique for mass detection in mammography. J Comput 11(6):472–478
Oktay O, Schlemper J, Folgoc LL, Lee M, Heinrich M, Misawa K, Mori K, McDonagh S, Hammerla NY, Kainz B et al (2018) Attention U-Net: learning where to look for the pancreas. ar**v preprint ar**v:1804.03999
Qu H, Riedlinger G, Wu P, Huang Q, Yi J, De S, Metaxas D (2019) Joint segmentation and fine-grained classification of nuclei in histopathology images. In: 2019 IEEE 16th international symposium on biomedical imaging (ISBI 2019). IEEE, pp 900–904
Rajalakshmi NR, Vidhyapriya R, Elango N, Ramesh N (2021) Deeply supervised U-Net for mass segmentation in digital mammograms. Int J Imaging Syst Technol 31(1):59–71
Ronneberger O, Fischer P, Brox T (2015) U-Net: convolutional networks for biomedical image segmentation. In: International conference on medical image computing and computer-assisted intervention. Springer, pp 234–241
Shiraishi J, Katsuragawa S, Ikezoe J, Matsumoto T, Kobayashi T, Ki Komatsu, Matsui M, Fujita H, Kodera Y, Doi K (2000) Development of a digital image database for chest radiographs with and without a lung nodule: receiver operating characteristic analysis of radiologists’ detection of pulmonary nodules. Am J Roentgenol 174(1):71–74
Simonyan K, Zisserman A (2014) Very deep convolutional networks for large-scale image recognition. ar**v preprint ar**v:1409.1556
Sirinukunwattana K, Pluim JP, Chen H, Qi X, Heng PA, Guo YB, Wang LY, Matuszewski BJ, Bruni E, Sanchez U et al (2017) Gland segmentation in colon histology images: the GLaS challenge contest. Med Image Anal 35:489–502
Sohl-Dickstein J, Weiss E, Maheswaranathan N, Ganguli S (2015) Deep unsupervised learning using nonequilibrium thermodynamics. In: International conference on machine learning. PMLR, pp 2256–2265
Sridevi M, Mala C (2012) A survey on monochrome image segmentation methods. Procedia Technol 6:548–555
Vahadane A, Atheeth B, Majumdar S (2021) Dual encoder attention u-net for nuclei segmentation. In: 2021 43rd annual international conference of the IEEE Engineering in Medicine & Biology Society (EMBC). IEEE, pp 3205–3208
Valanarasu JMJ, Oza P, Hacihaliloglu I, Patel VM (2021) Medical transformer: gated axial-attention for medical image segmentation. In: International conference on medical image computing and computer-assisted intervention. Springer, pp 36–46
Valanarasu JMJ, Sindagi VA, Hacihaliloglu I, Patel VM (2021) KIU-Net: overcomplete convolutional architectures for biomedical image and volumetric segmentation. IEEE Trans Med Imaging 41(4):965–976
Vaswani A, Shazeer N, Parmar N, Uszkoreit J, Jones L, Gomez AN, Kaiser Ł, Polosukhin I (2017) Attention is all you need. Adv Neural Inf Process Syst 30
Wang Q, Wu B, Zhu P, Li P, Zuo W, Hu Q (2020) ECA-Net: efficient channel attention for deep convolutional neural networks. In: Proceedings of the IEEE/CVF conference on computer vision and pattern recognition (CVPR)
Wang X, Girshick R, Gupta A, He K (2018) Non-local neural networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition, pp 7794–7803
Wu K, Zhang D (2015) Robust tongue segmentation by fusing region-based and edge-based approaches. Expert Syst Appl 42(21):8027–8038
Xu Q, Duan W, He N (2022) DCSAU-Net: a deeper and more compact split-attention U-Net for medical image segmentation. ar**v preprint ar**v:2202.00972
Yang Y, Wan W, Huang S, Zhong X, Kong X (2023) RADCU-Net: residual attention and dual-supervision cascaded U-Net for retinal blood vessel segmentation. Int J Mach Learn Cybern 14:1605–1620
Zagoruyko S, Komodakis N (2016) Wide residual networks. ar**v preprint ar**v:1605.07146
Zhan B, Song E, Liu H, Gong Z, Ma G, Hung CC (2023) CFNet: a medical image segmentation method using the multi-view attention mechanism and adaptive fusion strategy. Biomed Signal Process Control 79:104112
Zhao X, Zhang L, Lu H (2021) Automatic polyp segmentation via multi-scale subtraction network. In: International conference on medical image computing and computer-assisted intervention. Springer, pp 120–130
Zhou Z, Rahman Siddiquee MM, Tajbakhsh N, Liang J (2018) UNet++: a nested U-Net architecture for medical image segmentation. In: Deep learning in medical image analysis and multimodal learning for clinical decision support. Springer, pp 3–11
Acknowledgements
The authors acknowledge the National Natural Science Foundation of China (Grant nos. 61772319, 62002200, 62202268, 62272281), Shandong Provincial Science and Technology Support Program of Youth Innovation Team in Colleges (under Grant 2021KJ069, 2019KJN042), Yantai science and technology innovation development plan (2022JCYJ031).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no potential Conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Fu, Z., Li, J., Chen, Z. et al. DmADs-Net: dense multiscale attention and depth-supervised network for medical image segmentation. Int. J. Mach. Learn. & Cyber. (2024). https://doi.org/10.1007/s13042-024-02248-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13042-024-02248-7