Abstract
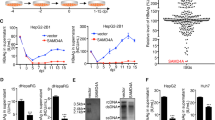
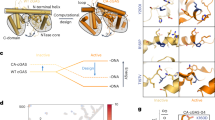
Cyclic dinucleotides (CDNs) are known to activate stimulator of interferon genes (STING) and induce type I interferon responses, therefor possess great potentials to be of immunotherapeutic value for cancers and infectious diseases. However, the existence of different single nucleotide polymorphism (SNP) of human STING (hSTING) gene poses an obstacle to achieve broad-spectrum activation by CDNs. We reported here the design and synthesis of a total of 36 CDNs, representing all structural variations, that contain four bases (A, G, C, U) and two linkage directions (2′-5′-linked and 3′-5′-linked phosphodiester). Through systematic evaluation of IFN-β induction with a dual-luciferase reporter assay, we discovered that wild type hSTING and two isoforms (HAQ and AQ) showed strong response while hSTING-R232H and R293Q exhibited the relatively weak response to CDNs stimulation. For the first time, we found that the c[G(2′,5′)U(2′,5′)] showed excellent activity against all five hSTING variants even equivalent to the endogenous ligand c[G(2′,5′)A(3′,5′)]. Furthermore, we have also demonstrated that 3′-3′ CDNs with two 3′-5′ phosphodiesters showed higher serum and hydrolase stability than 2′-2′ CDNs with two 2′-5′ phosphodiesters and 2′-3′ CDNs with one 2′-5′ and one 3′-5′ phosphodiester. It is very interesting to note that 2′-2′ CDNs has been found for the first time to show strong activity. These findings will stimulate our exploration for the new functional role of CDNs, and provide guidelines to design CDNs based hSTING targeted drugs.
Similar content being viewed by others
References
Krasteva PV, Sondermann H. Nat Chem Biol, 2017, 13: 350–359
Schaap P. IUBMB Life, 2013, 65: 897–903
Sun L, Wu J, Du F, Chen X, Chen ZJ. Science, 2013, 339: 786–791
Wu J, Sun L, Chen X, Du F, Shi H, Chen C, Chen ZJ. Science, 2013, 339: 826–830
Gao P, Ascano M, Zillinger T, Wang W, Dai P, Serganov AA, Gaffney BL, Shuman S, Jones RA, Deng L, Hartmann G, Barchet W, Tuschl T, Patel DJ. Cell, 2013, 154: 748–762
Zhang X, Shi H, Wu J, Zhang X, Sun L, Chen C, Chen ZJ. Mol Cell, 2013, 51: 226–235
Ablasser A, Goldeck M, Cavlar T, Deimling T, Witte G, Röhl I, Hopfner KP, Ludwig J, Hornung V. Nature, 2013, 498: 380–384
Lioux T, Mauny MA, Lamoureux A, Bascoul N, Hays M, Vernejoul F, Baudru AS, Boularan C, Lopes-Vicente J, Qushair G, Tiraby G. J Med Chem, 2016, 59: 10253–10267
Burdette DL, Monroe KM, Sotelo-Troha K, Iwig JS, Eckert B, Hyodo M, Hayakawa Y, Vance RE. Nature, 2011, 478: 515–518
Shi H, Wu J, Chen ZJ, Chen C. Proc Natl Acad Sci USA, 2015, 112: 8947–8952
Wang C, Sinn M, Stifel J, Heiler AC, Sommershof A, Hartig JS. J Am Chem Soc, 2017, 139: 16154–16160
Fu J, Kanne DB, Leong M, Glickman LH, McWhirter SM, Lemmens E, Mechette K, Leong JJ, Lauer P, Liu W, Sivick KE, Zeng Q, Soares KC, Zheng L, Portnoy DA, Woodward JJ, Pardoll DM, Dubensky Jr. TW, Kim Y. Sci Transl Med, 2015, 7: 283ra52
Ghaffari A, Peterson N, Khalaj K, Vitkin N, Robinson A, Francis JA, Koti M. Br J Cancer, 2018, 119: 440–449
Wang H, Hu S, Chen X, Shi H, Chen C, Sun L, Chen ZJ. Proc Natl Acad Sci USA, 2017, 114: 1637–1642
Smith TT, Moffett HF, Stephan SB, Opel CF, Dumigan AG, Jiang X, Pillarisetty VG, Pillai SPS, Wittrup KD, Stephan MT. J Clin Investigat, 2017, 127: 2176–2191
Deng L, Liang H, Xu M, Yang X, Burnette B, Arina A, Li XD, Mauceri H, Beckett M, Darga T, Huang X, Gajewski TF, Chen ZJ, Fu YX, Weichselbaum RR. Immunity, 2014, 41: 843–852
Li T, Cheng H, Yuan H, Xu Q, Shu C, Zhang Y, Xu P, Tan J, Rui Y, Li P, Tan X. Sci Rep, 2016, 6: 19049
Yi G, Brendel VP, Shu C, Li P, Palanathan S, Cheng Kao C. PLoS ONE, 2013, 8: e77846
Whiteley AT, Eaglesham JB, de Oliveira Mann CC, Morehouse BR, Lowey B, Nieminen EA, Danilchanka O, King DS, Lee ASY, Mekalanos JJ, Kranzusch PJ. Nature, 2019, 567: 194–199
Gentili M, Kowal J, Tkach M, Satoh T, Lahaye X, Conrad C, Boyron M, Lombard B, Durand S, Kroemer G, Loew D, Dalod M, Théry C, Manel N. Science, 2015, 349: 1232–1236
Girardin SE, Boneca IG, Carneiro LAM, Antignac A, Jéhanno M, Viala J, Tedin K, Taha MK, Labigne A, Zähringer U, Coyle AJ, DiStefano PS, Bertin J, Sansonetti PJ, Philpott DJ. Science, 2003, 300: 1584–1587
Case DA. Case D, Berryman J, Betz RM, Cerutti DS, Cheatham T, Darden T, Duke RE, Giese TJ, Gohlke H, Götz A, Homeyer N, Izadi S, Janowski P, Kaus J, Kovalenko A, Lee TS, LeGrand S, Li P, Luchko T. Kollman PA. Amber 2015, University of California, San Francisco, 2015
Salomon-Ferrer R, Götz AW, Poole D, Le Grand S, Walker RC. J Chem Theor Comput, 2013, 9: 3878–3888
Maier JA, Martinez C, Kasavajhala K, Wickstrom L, Hauser KE, Simmerling C. J Chem Theor Comput, 2015, 11: 3696–3713
Wang B, Wang Z, Javornik U, ** Z, Plavec J. Sci Rep, 2017, 7: 16550
Darden T, York D, Pedersen L. J Chem Phys, 1993, 98: 10089–10092
Ryckaert JP, Ciccotti G, Berendsen HJC. J Comput Phys, 1977, 23: 327–341
Zhang C, Shang G, Gui X, Zhang X, Bai XC, Chen ZJ. Nature, 2019, 567: 394–398
Zhao B, Du F, Xu P, Shu C, Sankaran B, Bell SL, Liu M, Lei Y, Gao X, Fu X, Zhu F, Liu Y, Laganowsky A, Zheng X, Ji JY, West AP, Watson RO, Li P. Nature, 2019, 569: 718–722
Mukai K, Konno H, Akiba T, Uemura T, Waguri S, Kobayashi T, Barber GN, Arai H, Taguchi T. Nat Commun, 2016, 7: 11932
Corrales L, Glickman LH, McWhirter SM, Kanne DB, Sivick KE, Katibah GE, Woo SR, Lemmens E, Banda T, Leong JJ, Metchette K, Dubensky Jr. TW, Gajewski TF. Cell Rep, 2015, 11: 1018–1030
Yin Q, Tian Y, Kabaleeswaran V, Jiang X, Tu D, Eck MJ, Chen ZJ, Wu H. Mol Cell, 2012, 46: 735–745
Che X, Zhang J, Quan H, Yang L, Gao YQ. J Phys Chem B, 2018, 122: 1862–1868
Volpi S, Insalaco A, Caorsi R, Santori E, Messia V, Sacco O, Terheggen-Lagro S, Cardinale F, Scarselli A, Pastorino C, Moneta G, Cangemi G, Passarelli C, Ricci M, Girosi D, Derchi M, Bocca P, Diociaiuti A, El Hachem M, Cancrini C, Tomà P, Granata C, Ravelli A, Candotti F, Picco P, DeBenedetti F, Gattorno M. J Clin Immunol, 2019, 39: 476–485
Motwani M, Pawaria S, Bernier J, Moses S, Henry K, Fang T, Burkly L, Marshak-Rothstein A, Fitzgerald KA. Proc Natl Acad Sci USA, 2019, 116: 7941–7950
Konno H, Chinn IK, Hong D, Orange JS, Lupski JR, Mendoza A, Pedroza LA, Barber GN. Cell Rep, 2018, 23: 1112–1123
Melki I, Rose Y, Uggenti C, Van Eyck L, Frémond ML, Kitabayashi N, Rice GI, Jenkinson EM, Boulai A, Jeremiah N, Gattorno M, Volpi S, Sacco O, Terheggen-Lagro SWJ, Tiddens HAWM, Meyts I, Morren MA, De Haes P, Wouters C, Legius E, Corveleyn A, Rieux-Laucat F, Bodemer C, Callebaut I, Rodero MP, Crow YJ. J Allergy Clin Immunol, 2017, 140: 543–552.e5
Huang YH, Liu XY, Du XX, Jiang ZF, Su XD. Nat Struct Mol Biol, 2012, 19: 728–730
Shang G, Zhu D, Li N, Zhang J, Zhu C, Lu D, Liu C, Yu Q, Zhao Y, Xu S, Gu L. Nat Struct Mol Biol, 2012, 19: 725–727
Shu C, Yi G, Watts T, Kao CC, Li P. Nat Struct Mol Biol, 2012, 19: 722–724
Shang G, Zhang C, Chen ZJ, Bai XC, Zhang X. Nature, 2019, 567: 389–393
Ouyang S, Song X, Wang Y, Ru H, Shaw N, Jiang Y, Niu F, Zhu Y, Qiu W, Parvatiyar K, Li Y, Zhang R, Cheng G, Liu ZJ. Immunity, 2012, 36: 1073–1086
Ergun SL, Fernandez D, Weiss TM, Li L. Cell, 2019, 178: 290–301.e10
Kato K, Nishimasu H, Oikawa D, Hirano S, Hirano H, Kasuya G, Ishitani R, Tokunaga F, Nureki O. Nat Commun, 2018, 9: 4424
Li L, Yin Q, Kuss P, Maliga Z, Millán JL, Wu H, Mitchison TJ. Nat Chem Biol, 2014, 10: 1043–1048
Che X, Zhang J, Zhu Y, Yang L, Quan H, Gao YQ. J Phys Chem B, 2016, 120: 2670–2680
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2017YFD0200500) and the National Natural Science Foundation of China (21740002, 21837001). We also thanks Prof. Junmin Quan for the support of THP-1-Lucia cells and Prof. Dongsheng Guo for the help of ITC assay.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest The authors declare that they have no conflict of interest.
Supporting information
11426_2019_9662_MOESM1_ESM.pdf
Design, synthesis and systematic evaluation of all possible cyclic dinucleotides (CDNs) that activate human stimulator of interferon genes (STING) variants
Rights and permissions
About this article
Cite this article
Wang, ZH., Zhao, CC., Zhang, QZ. et al. Design, synthesis and systematic evaluation of all possible cyclic dinucleotides (CDNs) that activate human stimulator of interferon genes (STING) variants. Sci. China Chem. 63, 534–545 (2020). https://doi.org/10.1007/s11426-019-9662-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11426-019-9662-5