Abstract
Leaves are the key organs of plants that produce photosynthates. Leaf senescence aids nutrient remobilization from source to sink tissues, which dramatically affects crop quality and yield. Although several senescence-associated genes (SAGs) have been recognized in cotton, the function of lipoic acid synthase in leaf senescence remains unclear. Therefore, we isolated a lipoic acid synthase gene named GhLIP1 from cotton (Gossypium hirsutum L.). Protein domain analysis revealed that GhLIP1 had a LIAS-N domain and an Elp3 domain, and yeast mutant complementation experiments demonstrated that GhLIP1 was a lipoic acid synthase. GhLIP1 expression pattern analysis showed that GhLIP1 was vastly expressed in fibres, and the phytohormone IAA (indole-3-acetic acid) induced its transcription. When GhLIP1 was transformed into Arabidopsis, the leaves of overexpression Arabidopsis exhibited a late leaf senescence phenotype compared to wild-type Arabidopsis, Col-0. Moreover, downregulation of GhLIP1 expression using virus-induced gene silencing (VIGS) technology resulted in early leaf senescence in cotton. qRT-PCR analyses revealed that two SAGs, GhWRKY53 and GhNAP, were upregulated in cotton plants with GhLIP1 knock-down. Cell ultrastructure observations found that chloroplasts were degraded in an orderly manner in cotton leaf cells with GhLIP1 knock-down. These results suggest that GhLIP1 plays a significant role in leaf senescence in cotton.







Similar content being viewed by others
References
Babiychuk E, Vandepoele K, Wissing J, Garcia-Diaz M, De Rycke R, Akbari H, Joubes J, Beeckman T, Jansch L, Frentzen M, Van Montagu MCE, Kushnir S (2011) Plastid gene expression and plant development require a plastidic protein of the mitochondrial transcription termination factor family. Proc Natl Acad Sci USA 108:6674–6679
Balazadeh S, Siddiqui H, Ad A, Lp MR, Caldana C, Mehrnia M, Mi Z, Khler B, Muellerroeber B (2010) A gene regulatory network controlled by the NAC transcription factor ANAC092/AtNAC2/ORE1 during salt-promoted senescence. Plant J 62:250–264
Balazadeh S, Schildhauer J, Araújo WL, Munné-Bosch S, Fernie AR, Proost S, Humbeck K, Mueller-Roeber B (2014) Reversal of senescence by N resupply to N-starved Arabidopsis thaliana: transcriptomic and metabolomic consequences. J Exp Bot 65:3975–3992
Barth C, Moeder W, Klessig DF, Conklin PL (2004) The timing of senescence and response to pathogens is altered in the ascorbate-deficient Arabidopsis mutant vitamin c-1. Plant Physiol 134:1784–1792
Breeze E, Harrison E, McHattie S, Hughes L, Hickman R, Hill C, Kiddle S, Kim YS, Penfold CA, Jenkins D, Zhang CJ, Morris K, Jenner C, Jackson S, Thomas B, Tabrett A, Legaie R, Moore JD, Wild DL, Ott S, Rand D, Beynon J, Denby K, Mead A, Buchanan-Wollaston V (2011) High-resolution temporal profiling of transcripts during Arabidopsis leaf senescence reveals a distinct chronology of processes and regulation. Plant Cell 23:873–894
Brouwer B, Ziolkowska A, Bagard M, Keech O, Gardestrom P (2012) The impact of light intensity on shade-induced leaf senescence. Plant Cell Environ 35:1084–1098
Buchanan-Wollaston V, Page T, Harrison E, Breeze E, Lim PO, Nam HG, Lin JF, Wu SH, Swidzinski J, Ishizaki K, Leaver CJ (2005) Comparative transcriptome analysis reveals significant differences in gene expression and signalling pathways between developmental and dark/starvation-induced senescence in Arabidopsis. Plant J 42:567–585
Chen ER, Wang XQ, Gong Q, Butt HI, Chen YL, Zhang CJ, Yang ZR, Wu ZX, Ge XY, Zhang XL, Li FG, Zhang XY (2017) A novel GhBEE1-Like gene of cotton causes anther indehiscence in transgenic Arabidopsis under uncontrolled transcription level. Gene 627:49–56
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Fujiwara K, Okamura-Ikeda K, Motokawa Y (1990) cDNA sequence, in vitro synthesis, and intramitochondrial lipoylation of H-protein of the glycine cleavage system. J Biol Chem 265:17463–17467
Gan S, Amasino RM (1997) Making sense of senescence (Molecular genetic regulation and manipulation of leaf senescence). Plant Physiol 113:313–319
Gepstein S, Sabehi G, Carp MJ, Hajouj T, Nesher MFO, Yariv I, Dor C, Bassani M (2004) Large-scale identification of leaf senescence-associated genes. Plant J 36:629–642
Guinn G (1985) Abscisic acid and cutout in cotton. Plant Physiol 77:16–20
Guo P, Li Z, Huang P, Li B, Fang S, Chu J, Guo H (2017) A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence. Plant Cell 29:2854–2870
Huang G, Wu Z, Percy RG, Bai M, Li Y, Frelichowski JE, Hu J, Wang K, Yu JZ, Zhu Y (2020) Genome sequence of Gossypium herbaceum and genome updates of Gossypium arboreum and Gossypium hirsutum provide insights into cotton A-genome evolution. Nat Genet 52:516–524
Jiang YJ, Liang G, Yang SZ, Yu DQ (2014) Arabidopsis WRKY57 functions as a node of convergence for jasmonic acid- and auxin-mediated signaling in jasmonic acid-induced leaf senescence. Plant Cell 26:230–245
Jibran R, Hunter DA, Dijkwel PP (2013) Hormonal regulation of leaf senescence through integration of developmental and stress signals. Plant Mol Biol 82:547–561
**g HC, Schippers JH, Hille J, Dijkwel PP (2005) Ethylene-induced leaf senescence depends on age-related changes and OLD genes in Arabidopsis. J Exp Bot 56:2915–2923
Kim Y, Oliver DJ (1990) Molecular cloning, transcriptional characterization, and sequencing of cDNA encoding the H-protein of the mitochondrial glycine decarboxylase complex in peas. J Biol Chem 265:848–853
Kim HJ, Ryu H, Hong SH, Woo HR, Lim PO, Lee IC, Sheen J, Nam HG, Hwang I (2006) Cytokinin-mediated control of leaf longevity by AHK3 through phosphorylation of ARR2 in Arabidopsis. Proc Natl Acad Sci USA 103:814–819
Kusaba M, Tanaka A, Tanaka R (2013) Stay-green plants: what do they tell us about the molecular mechanism of leaf senescence. Photosynth Res 117:221–234
Lee S, Seo PJ, Lee HJ, Park CM (2012) A NAC transcription factor NTL4 promotes reactive oxygen species production during drought-induced leaf senescence in Arabidopsis. Plant J 70:831–844
Leopold AC (1961) Senescence in plant development: the death of plants or plant parts may be of positive ecological or physiological value. Science 134:1727–1732
Li Z, Peng J, Wen X, Guo H (2012) Gene network analysis and functional studies of senescence-associated genes reveal novel regulators of Arabidopsis leaf senescence. J Integr Plant Biol 54:526–539
Liang CZ, Wang YQ, Zhu YN, Tang JY, Hu B, Liu LC, Ou SJ, Wu HK, Sun XH, Chu JF, Chu CC (2014) OsNAP connects abscisic acid and leaf senescence by fine-tuning abscisic acid biosynthesis and directly targeting senescence-associated genes in rice. Proc Natl Acad Sci USA 111:10013–10018
Lim PO, Woo HR, Nam HG (2003) Molecular genetics of leaf senescence in Arabidopsis. Trends Plant Sci 8:272–278
Lim PO, Kim HJ, Nam HG (2007) Leaf senescence. Annu Rev Plant Biol 58:115–136
Lin M, Pang CY, Fan SL, Song MZ, Wei HL, Yu SX (2015) Global analysis of the Gossypium hirsutum L. transcriptome during leaf senescence by RNA-SEq. BMC Plant Biol 15:43
Liu J, Pang CY, Wei HL, Song MZ, Meng YY, Ma JH, Fan SL, Yu SX (2015) iTRAQ-facilitated proteomic profiling of anthers from a photosensitive male sterile mutant and wild-type cotton (Gossypium hirsutum L.). J Proteomics 126:68–81
Macherel D, Lebrun M, Gagnon J, Neuburger M, Douce R (1990) cDNA cloning, primary structure and gene expression for H-protein, a component of the glycine-cleavage system (glycine decarboxylase) of pea (Pisum sativum) leaf mitochondria. Biochem J 268:783–789
Mattevi A, de Kok A, Perham RN (1992) The pyruvate dehydrogenase multienzyme complex. Curr Opin Struc Biol 2:877–887
Miller JR, Busby RW, Jordan SW, Cheek J, Henshaw TF, Ashley GW, Broderick JB, Cronan JE, Marletta MA (2000) Escherichia coli LipA is a lipoyl synthase: In vitro biosynthesis of lipoylated pyruvate dehydrogenase complex from octanoyl-acyl carrier protein. Biochemistry 39:15166–15178
Nam HG (1997) The molecular genetic analysis of leaf senescence. Curr Opin Biotechnol 8:200–207
Noodén LD (1988) The Phenomena of Senescence and Aging. In: Noodén LD, Leopold AC (eds) Senescence and Aging in Plants. Academic Press, pp 1–50
Oh SA, Sang YL, Chung IK, Lee CH, Hong GN (1996) A senescence-associated gene of Arabidopsis thaliana is distinctively regulated during natural and artificially induced leaf senescence. Plant Mol Biol 30:739–754
Ouellet F, Overvoorde PJ, Theologis A (2001) IAA17/AXR3: biochemical insight into an auxin mutant phenotype. Plant Cell 13:829–841
Pang J, Zhu Y, Li Q, Liu J, Tian Y, Liu Y, Wu J (2013) Development of Agrobacterium-mediated virus-induced gene silencing and performance evaluation of four marker genes in Gossypium barbadense. PLoS One 8:e73211
Perham RN (1991) Domains, motifs, and linkers in 2-oxo acid dehydrogenase multienzyme complexes: a paradigm in the design of a multifunctional protein. Biochemistry 30:8501–8512
Quirino BF, Noh YS, Himelblau E, Amasino RM (2000) Molecular aspects of leaf senescence. Trends Plant Sci 5:278–282
Reed KE, Cronan JE Jr (1993) Lipoic acid metabolism in Escherichia coli: sequencing and functional characterization of the lipA and lipB genes. J Bacteriol 175:1325–1336
Reed LJ, Hackert ML (1990) Structure-function relationships in dihydrolipoamide acyltransferases. J Bio Chem 16:8971–8974
Ren MZ, Venglat P, Qiu SQ, Feng L, Cao YG, Wang E, **ang DQ, Wang JH, Alexander D, Chalivendra S, Logan D, Mattoo A, Selvaraj G, Datla R (2012) Target of rapamycin signaling regulates metabolism, growth, and life span in Arabidopsis. Plant Cell 24:4850–4874
Richard-Molard C, Krapp A, Brun F, Ney B, Daniel-Vedele F, Chaillou S (2008) Plant response to nitrate starvation is determined by N storage capacity matched by nitrate uptake capacity in two Arabidopsis genotypes. J Exp Bot 59:779–791
Rouse D, Mackay P, Stirnberg P, Estelle M, Leyser O (1998) Changes in auxin response from mutations in an AUX/IAA gene. Science 279:1371–1373
Senchina DS, Alvarez I, Cronn RC, Liu B, Rong JK, Noyes RD, Paterson AH, Wing RA, Wilkins TA, Wendel JF (2003) Rate variation among nuclear genes and the age of polyploidy in Gossypium. Mol Biol Evol 20:633–643
Shi HT, Chen L, Ye TT, Liu XD, Ding KJ, Chan ZL (2014) Modulation of auxin content in Arabidopsis confers improved drought stress resistance. Plant Physiol Bioch 82:209–217
Shoji K, Addicott FT, Swets WA (1951) Auxin in relation to leaf blade abscission. Plant Physiol 26:189–191
Singh VP, Jegadheesan A (2003) Effect of alpha-lipoic acid on senescence in gladiolus flowers. Indian J Plant Physiol Special Issue No 1:72–79
Sulo P, Martin NC (1993) Isolation and characterization of LIP5. A lipoate biosynthetic locus of Saccharomyces cerevisiae. J Biol Chem 268:17634–17639
van der Graaff E, Schwacke R, Schneider A, Desimone M, Flugge UI, Kunze R (2006) Transcription analysis of Arabidopsis membrane transporters and hormone pathways during developmental and induced leaf senescence. Plant Physiol 141:776–792
Wang X, Chen E, Ge X, Gong Q, Butt H, Zhang C, Yang Z, Li F, Zhang X (2018) Overexpressed BRH1, a RING finger gene, alters rosette leaf shape in Arabidopsis thaliana. Sci China Life Sci 61:79–87
Weaver LM, Gan S, Quirino B, Amasino RM (1998) A comparison of the expression patterns of several senescence-associated genes in response to stress and hormone treatment. Plant Mol Biol 37:455–469
Webber JM (1936) Cytogenetic notes on cotton and cotton relatives. II Science 84:378
Wendel JF (1989) New World tetraploid cottons contain Old World cytoplasm. Proc Natl Acad Sci USA 86:4132–4136
Woo HR, Chung KM, Park JH, Oh SA, Ahn T, Hong SH, Jang SK, Nam HG (2001) ORE9, an F-Box protein that regulates leaf senescence in Arabidopsis. Plant Cell 13:1779–1790
Woo HR, Koo HJ, Kim J, Jeong H, Yang JO, Lee IH, Jun JH, Choi SH, Park SJ, Kang B, Kim YW, Phee BK, Kim JH, Seo C, Park C, Kim SC, Park S, Lee B, Lee S, Hwang D, Nam HG, Lim PO (2016) Programming of plant leaf senescence with temporal and inter-organellar coordination of transcriptome in Arabidopsis. Plant Physiol 171:452–467
Wright BP (1998) Research into early senescence syndrome in cotton. Better Crops International 12:14–16
Yang SD, Seo PJ, Yoon HK, Park CM (2011) The Arabidopsis NAC transcription factor VNI2 integrates abscisic acid signals into leaf senescence via the COR/RD genes. Plant Cell 23:2155–2168
Yasuno R, Wada H (1998) Biosynthesis of lipoic acid in Arabidopsis: cloning and characterization of the cDNA for lipoic acid synthase. Plant Physiol 118:935–943
Yasuno R, Wada H (2002) The biosynthetic pathway for lipoic acid is present in plastids and mitochondria in Arabidopsis thaliana. Febs Lett 517:110–114
Zhao J, Jiang T, Liu Z, Zhang W, Jian G, Qi F (2012) Dominant gene cplsr1 corresponding to premature leaf senescence resistance in cotton (Gossypium hirsutum L.). J Integr Plant Biol 54:577–583
Zhou J, Wang J, Cheng Y, Chi YJ, Fan B, Yu JQ, Chen Z, Gassmann W (2013) NBR1-mediated selective autophagy targets insoluble ubiquitinated protein aggregates in plant stress responses. PloS Genet 10:e1004477
Acknowledgements
We thank Dr. Shijun Chen of the Experimental Center at Henan Institute of Science and Technology for his help with the cell ultrastructure observations.
Funding
This work was supported by the National Natural Science Foundation of China (Grant Number 31872129) and Henan Science and Technology Project (Grant Number 202102110146).
Author information
Authors and Affiliations
Contributions
EY Chen and CW Li conceived and designed the research; EY Chen, HY Hu, XB Yang, DX Li and QC Wei performed the experiments; F Zhou, YY Guan, YA Yu and PW Song analyzed the data; EY Chen and CW Li wrote, reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Longbiao Guo.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material
Fig. S1
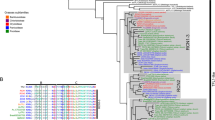
Multiple sequence alignment of the GhLIP1 protein and its homologs in other species. (PDF 16 kb)
Fig. S2
Phenotypes of cotton plants with down-regulated expression of GhLIP1 and GhPDS using VIGS technology. The TRV2::GhPDS cotton line was used as a positive control in the VIGS procedure. (TIF 7179 kb)
Table S1
Primers used in this study. Table S2 The qRT-PCR primers specifically used for the analysis of GhLIP1 and senescence-associated gene expression patterns. (DOCX 15 kb)
Rights and permissions
About this article
Cite this article
Chen, E., Hu, H., Yang, X. et al. GhLIP1, a lipoic acid synthase gene, negatively regulates leaf senescence in cotton. Plant Growth Regul 94, 73–85 (2021). https://doi.org/10.1007/s10725-021-00697-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-021-00697-6