Abstract
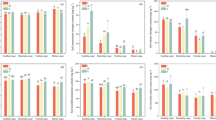
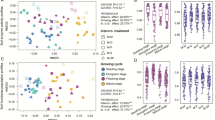
Soil microbes are essential links between above- and below-ground ecosystems and play an important role in regulating ecological functions in the soil. Dynamic interactions within the soil-microbial community in a cereal-legume intercrop** ecosystem influence the composition and structure of N-cycling microbial groups (e.g. nitrogen-fixing bacteria). However, these effects have not been extensively studied in some intercrop** patterns or in response to varying nitrogen fertilization levels. In the present study, we evaluated the effects of reduced and conventional nitrogen application in a sweet maize (Zea may L.)/soybean (Glycine max L.) strip intercrop** system under three crop** patterns over a 3-year time period. High-throughput sequencing and quantitative PCR techniques were used to investigate changes to both the microbial community structure and the expression of key nitrogen-cycling genes in the rhizosphere. Our results indicate that reduced nitrogen application affected the microbial community structure in the rhizosphere, but microbial diversity in the sweet maize rhizosphere was relatively stable. Both the abundance and activity of functional marker genes for microbial nitrogen fixation (nifH), nitrification (amoA), denitrification (nirS, nirK, nosZ), and decomposition (chiA) increased significantly from 2013 to 2016. Taken together, these data demonstrate that the quantified shifts in the soil microbial community and the observed increases in the expression of key functional genes involved in N-cycling were the result of reduced nitrogen application in this strip intercrop** system. This study, therefore, provides essential insight into the potential relationships between functional nitrogen-cycling genes and mitigation of nitrogen-loss and N2O emissions in a cereal-legume strip intercrop** system.



Similar content being viewed by others
References
Bainard LD, Koch AM, Gordon AM, Klironomos JN (2013) Growth response of crops to soil microbial communities from conventional monocrop** and tree-based intercrop** systems. Plant Soil 363:345–356
Bao SD (2013) Soil agro-chemistrical analysis, 3rd edn. China Agriculture Press, Bei**g
Berthrong ST, Yeager CM, Gallegos-Graves L, Steven B, Eichorst SA, Jackson RB, Kuske CR (2014) Nitrogen fertilization has a stronger effect on soil nitrogen-fixing bacterial communities than elevated atmospheric CO2. Appl Environ Microb 80:3103–3112
Brooker RW, Bennett AE, Cong W, Daniell TJ, George TS, Hallett PD, Hawes C, Iannetta PPM, Jones HG, Karley AJ, Li L, McKenzie BM, Pakeman RJ, Paterson E, Schöb C, Shen J, Squire G, Watson CA, Zhang C, Zhang F, Zhang J, White PJ (2015) Improving intercrop**: a synthesis of research in agronomy, plant physiology and ecology. New Phytol 206(1):107–117
Campbell BJ, Poison SW, Hanson TE, Mack MC, Schuur EAG (2010) The effect of nutrient deposition on bacterial communities in Arctic tundra soil. Environ Microb 12:1842–1854
Chu GX, Shen Q, Cao J (2004) Nitrogen fixation and N transfer from peanut to rice cultivated in aerobic soil in an intercrop** system and its effect on soil N fertility. Plant Soil 263:17–27
Cong WF, Hoffland E, Li L, Six J, Sun JH, Bao XG, Zhang FS, Van Der Werf W (2015) Intercrop** enhances soil carbon and nitrogen. Glob Change Biol 21:1715–1726
Correa-Galeote D, Bedmar EJ, Fernandezlopez M, Arone GJ (2016) Bacterial communities in the rhizosphere of amilaceous maize (Zea mays L.) as assessed by pyrosequencing. Front Plant Sci 7(e197):1016
Corre-Hellou G, Brisson N, Launay M, Fusteca J, Crozata Y (2007) Effect of root depth penetration on soil nitrogen competitive interactions and dry matter production in pea–barley intercrops given different soil nitrogen supplies. Field Crops Res 103:76–85
Coskun D, Britto DT, Weiming S, Kronzucker HJ (2017) Nitrogen transformations in modern agriculture and the role of biological nitrification inhibition. Nat Plants 3:17074
Duchene O, Vian J-F, Celette F (2017) Intercrop** with legume for agroecological crop** systems: complementarity and facilitation processes and the importance of soil microorganisms—a review. Agr Ecosyst Environ 240:148–161
Fadrosh DW, Ma B, Gajer P, Sengamalay N, Ott S, Brotman RM, Ravel J (2014) An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform. Microbiome 2:6
Fan FL, Li ZJ, Steven AW, Wakelin Y, Liang YC (2012) Mineral fertilizer alters cellulolytic community structure and suppresses soil cellobiohydrolase activity in a long-term fertilization experiment. Soil Biol Biochem 55:70–77
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6:1007–1017
Gaba S, Lescourret F, Boudsocq S, Enjalbert J, Hinsinger P, Journet E-P, Navas M-L, Wery J, Louarn G, Malézieux E, Pelzer E, Prudent M, Ozier-Lafontaine H (2015) Multiple crop** systems as drivers for providing multiple ecosystem services: from concepts to design. Agron Sustain Dev 35:607–623
Hai B, Diallo NH, Sall S, Haesler F, Schauss K, Bonzi M, Assigbetse K, Chotte J-L, Munch JC, Schloter M (2009) Quantification of key genes steering the microbial nitrogen cycle in the rhizosphere of sorghum cultivars in tropical agroecosystems. Appl Environ Microb 75:4993–5000
Hargreaves SK, Williams RJ, Hofmockel KS (2015) Environmental filtering of microbial communities in agricultural soil shifts with crop growth. PLoS ONE 10(7):e0134345
Hauggaard-Nielsen H, Jørnsgaard B, Kinane J, Jensen ES (2008) Grain legume–cereal intercrop**: the practical application of diversity, competition and facilitation in arable and organic crop** systems. Renew Agric Food Syst 23:3–12
Hink L, Gubry-Rangin C, Nicol GW, Prosser JI (2018) The consequences of niche and physiological differentiation of archaeal and bacterial ammonia oxidisers for nitrous oxide emissions. ISME J 12:1084–1093
Kong AYY, Hristova K, Scow KM, Six J (2010) Impacts of different N management regimes on nitrifier and denitrifier communities and N cycling in soil microenvironments. Soil Biol Biochem 42(9):1523–1533
Lauber CL, Strickland MS, Bradford MA, Fierer N (2008) The influence of soil properties on the structure of bacterial and fungal communities across land-use types. Soil Biol Biochem 40:2407–2415
Li L, Zhang FS, Li XL, Christie P, Sun JH, Yang SC, Tang CX (2003) Interspecific facilitation of nutrient uptake by intercropped maize and faba bean. Nutr Cycl Agroecosyst 65:61–71
Li L, Tilman D, Lambers H, Zhang FS (2014) Plant diversity and overyielding: insights from belowground facilitation of intercrop** in agriculture. New Phytol 203:63–69
Li B, Li YY, Wu HM, Zhang FF, Li CJ, Li XX, Lambers H, Li L (2016) Root exudates drive interspecific facilitation by enhancing nodulation and N2 fixation. Proc Natl Acad Sci 113(23):6496–6501
Lindsay EA, Colloff MJ, Gibb NL, Wakelin SA (2010) The abundance of microbial functional genes in grassy woodlands is influenced more by soil nutrient enrichment than by recent weed invasion or livestock exclusion. Appl Environ Microb 76:5547–5555
Martin-Guay M-O, Paquette A, Dupras J, Rivest D (2018) The new green revolution: sustainable intensification of agriculture by intercrop**. Sci Total Environ 615:767–772
Philippot L, Hallin S, Schloterm M (2007) Ecology of denitrifying prokaryotes in agricultural soil. Adv Agron 96:135–190
Philippot L, Spor A, Henault C, Bru D, Bizouard F, Jones CM (2013) Loss in microbial diversity affects nitrogen cycling in soil. ISME J 7:1609–1619
Prosser JI, Nicol GW (2012) Archaeal and bacterial ammonia-oxidisers in soil: the quest for niche specialisation and differentiation. Trends Microbiol 20(11):523–531
Ribbons RR, Levy-Booth DJ, Masse J, Grayston SJ, McDonald MA, Vesterdal L, Prescott CE (2016) Linking microbial communities, functional genes and nitrogen-cycling processes in forest floors under four tree species. Soil Biol Biochem 103:181–191
Shen JP, Zhang LM, Guo JF, Jessica LR, He Z (2010) Impact of long-term fertilization practices on the abundance and composition of soil bacterial communities in Northeast China. Appl Soil Ecol 46:119–124
Shi P, Wang SP, Jia S, Gao Q (2015) Effect of 25-year fertilization on soil microbial biomass and community structure in a continuous maize crop** system. Arch Agron Soil Sci 61:1303–1317
Tang YL, Yu LL, Guan AM, Zhou XY, Wang ZG, Guo YG, Wang JW (2017) Soil mineral nitrogen residual and yield-scaled soil N2O emissions lowered by reducing nitrogen application and intercrop** with soybean for sweet maize production in southern China. J Integr Agr 16(11):2586–2596
Tatti E, Goyer C, Chantigny M, Wertz S, Zebarth BJ, Burton DL, Filion M (2014) Influences of over winter conditions on denitrification and nitrous oxide-producing microorganism abundance and structure in an agricultural soil amended with different nitrogen sources. Agr Ecosyst Environ 183:47–59
Wallenstein MD, Myrold DD, Firestone M, Voytek M (2006) Environmental controls on denitrifying communities and denitrification rates: insights from molecular methods. Ecol Appl 16(6):2143–2152
Williamson N, Brian P, Wellington EMH (2000) Molecular detection of bacterial and streptomycete chitinases in the environment. Anton Leeuw Int J 78(3):315–321
Xu M, Zhang Q, **a C, Zhong Y, Sun G, Guo J, Yuan T, Zhou J, He Z (2014) Elevated nitrate enriches microbial functional genes for potential bioremediation of complexly contaminated sediments. ISME J 8:1932–1944
Xue K, Wu L, Deng Y, He Z, Van Nostrand J, Robertson PG, Schmidt TM, Zhou J (2013) Functional gene differences in soil microbial communities from conventional, low-input, and organic farmlands. Appl Environ Microb 79:1284–1292
Zhou J, Guan D, Zhou B, Zhao B, Maa M, Qin J, Jiang X, Chen S, Cao F, Shen D, Li J (2015) Influences of 34-years of fertilization on bacterial communities in an intensively cultivated black soil in northeast China. Soil Biol Biochem 90:42–51
Zhu J, van der Werf W, Anten NPR, Vos J, Evers JB (2015) The contribution of phenotypic plasticity to complementary light capture in plant mixtures. New Phytol 207:1213–1222
Acknowledgements
A sincere thank you to Bernd Wollenweber for his diligent proofreading of this manuscript. And also we would like to thank Chen ** us to manage sweet maize cultivation in the field. We are grateful to the constructive comments from editor and two anonymous reviewers on this manuscript.
Funding
This work was supported by the National Natural Science Foundation of China [Grant Nos. 31770556, 31600348]; the Key Technologies R&D Program of China during the 12th five-year Plan period [Grant Nos. 2012BAD14B16-04]; and the Science and Technology Development Program of Guangdong [Grant Nos. 2015B090903077].
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Yu, L., Tang, Y., Wang, Z. et al. Nitrogen-cycling genes and rhizosphere microbial community with reduced nitrogen application in maize/soybean strip intercrop**. Nutr Cycl Agroecosyst 113, 35–49 (2019). https://doi.org/10.1007/s10705-018-9960-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10705-018-9960-4