Abstract
Background
Ferroptosis, as a unique form of cell death, plays crucial negative roles in tumorigenesis and progression. This study aimed to investigate the role and molecular mechanism of TEA domain transcription factor 1 (TEAD1) in HCC and its effect on sorafenib-induced ferroptosis.
Methods
TEAD1 expression was analyzed in HCC tissues using quantitative PCR, and western blot. The effects on cell proliferation, migration and invasion were determined by CCK-8, wound healing and Transwell assays. Intracellular iron, reactive oxygen species (ROS), malondialdehyde (MDA) and GSH measurement was used to assess ferroptosis. Chromatin immunoprecipitation and luciferase reporter gene assays were performed to verify the relationship between TEAD1 and solute carrier family 3 member 2 (SLC3A2). Expression of mTOR, ribosomal protein S6, glutathione peroxidase 4 (GPX4) and SLC3A2 was analyzed by western blot. Tumor xenografts were used assess the effect of TEAD1 on tumor growth in vivo.
Results
TEAD1 was more abundant in HCC compared with normal tissues. Overexpression of TEAD1 enhanced the proliferation, migration, and invasion of HCC cells, while knockdown of TEAD1 inhibited these cell behaviors. Further, TEAD1 inhibited ferroptosis, which was demonstrated by decreased intracellular Fe2+ content, ROS, and MDA levels, and increased GSH activity. Mechnistically, TEAD1 promotes the transcription of SLC3A2 and activates the mTOR signaling. Additionally, silenced TEAD1 restrained tumor growth and enhance sorafenib-induced antitumor activity in vivo.
Conclusions
TEAD1 confers resistance of HCC cells to ferroptosis, thereby promoting the progression of HCC, suggesting the potential value of TEAD1 in the diagnosis and treatment of HCC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hepatocellular carcinoma (HCC), as the most common type of liver cancer, accounts for about 80% of all liver cancers. Despite significant achievements in diagnosis and treatment in recent years, its clinical prognosis remains poor. Most HCC patients are diagnosed with late or even distant metastases, and the 5-year disease-free survival rate was about 18%, and the median survival time was about one year. Therefore, there is an urgent need to develop a less toxic and more effective treatment strategy.
Ferroptosis is a novel form of non-apoptotic cell death, which is triggered by iron-dependent lipid peroxides. It is generally believed that iron dependence, redox system dysfunction, and lipid peroxidation are the common markers of ferroptosis. A large amount of evidence shows that the mechanism of ferroptosis is closely related to the imbalance of redox balance in the cell. Ferroptosis is initiated when glutathione peroxidase GPX4, which is responsible for the clearance of lipid ROS, is directly or indirectly inhibited, resulting in cell death. Accumulating evidence has demonstrated that ferroptosis acts as a key negative regulator in the development and progression of human cancers, including HCC, and induction of ferroptosis is expected to be a promising anti-tumor therapy strategy [1,2,3]. Recently, several ferroptosis-related genes have been identified to be implicated in HCC progression and sorafenib sensitivity. Several molecules including quiescin sulfhydryl oxidase 1, GSTZ1 have been found to sensitize HCC to sorafenib-induced ferroptosis via inhibiting NRF2 activation. [4, 5]. Iron metabolism-related protein H-ferritin protects HCC from RSL3-induced ferroptosis [6]. Further, some proteins, such as TEAD1, ABCC5, are involved in ferroptosis via regulating SLC7A11 expression [7, 8]. Besides, CISD2 confers the resistance of HCC cell to sorafenib-induced ferroptosis through modulation of autophagy [9]. While regulating the expression of these key molecules can promote ferroptosis of HCC and enhance the efficacy of sorafenib. These findings indicate the significance of ferroptosis in HCC. However, there are still limited reports on regulatory proteins or signaling pathways of ferroptosis, and the exact regulatory network is still unknown.
TEA domain transcription factor 1 (TEAD1), as a member of the TEAD family, regulates cell viability, tissue regeneration, and stem cell maintenance [10]. The TEAD1 protein contains a TEA DNA binding domain, which can bind to the promoters of target genes, and a Yes-associated protein/transcriptional co-activator, which can bind to PDZ-binding motif (YAP/TAZ) binding domain of transcriptional cofactors [11]. TEAD1 coordinates with transcriptional cofactors including YAP, TAZ, and VGLL to modulate various biological processes in cancers [10]. Many studies reported that TEAD1 was up-regulated in malignant tumors including glioma, nasopharyngeal carcinoma, gastric cancer, colorectal cancer, renal cancer, prostate cancer, and HCC, which predicts poor survival of patients [10,11,12,13,14]. Targeting TEAD1 or blockade of YAP-TEAD association could inhibit tumor proliferation, migration, invasiveness, and enhanced apoptosis, which is a viable therapeutic strategy for certain cancers [11, 15]. In HCC, TEAD1 was found to elevate in HCC samples relative to nontumorous tissues and promoted tumor progression [16]. However, the role of TEAD1 in regulating the ferroptosis of HCC remains elusive.
Here, we explored the role and molecular mechanism of TEAD1 in HCC. We found that TEAD1 was overexpressed in HCC tissues, and upregulated TEAD1 activated mTORC1 signaling and increases GPX4 protein levels, thus conferring the resistance of HCC cells to ferroptosis, and eventually promoting cell survival. While depletion of TEAD1 triggers ferroptosis, thus inhibiting cancer cell growth and sensitizing HCC cells to sorafenib.
Materials and Methods
UALCAN
UALCAN (http://ualcan.path.uab.edu/index.html) was utilized to analyze TEAD1 mRNA expression in liver hepatocellular carcinoma (TIHC) tissues based on TCGA datasets [17].
HCC Patients
Thirty pairs of HCC tissues and adjacent normal tissues from HCC patients in the Cancer Center, Sichuan Provincial People's Hospital. The experimental scheme was approved by the Research Ethics Committee of the Sichuan Provincial People's Hospital (No. 2021363), and each enrolled participant signed informed consent.
Cell Lines and Cell Cuture
HCC cell lines were purchased from the National Collection of Authenticated Cell Cultures (Shanghai, China). These cells were cultured in RPMI 1640 medium with 10% FBS.
Plasmid Construction and Cell Transfection
The cDNA sequences of TEAD1 (NM_021961.6) were subcloned into pcDNA3.1 plasmid vector to construct TEAD1-overexpressing plasmids (TEAD1_OE). The small interfering RNAs targeting TEAD1 (si-TEAD1) or SLC3A2 (si-SLC3A2) was obtained from GenePharma (Suzhou, China). For cell tranfection, TEAD1-overexpressing plasmids or si-TEAD1 were transfected into HCC cells using Lipofectamine 3000 (Thermo Fisher Scientific, USA) according to manufacturer's instructions.
Quantitative Real-Time PCR (qRT-PCR)
Total RNA was isolated from tissue or cell samples with TRIzol reagent (ThermoFisher, Carlsbad, CA, USA). Then, PrimeScript RT reagent Kit (Takara, Bei**g, China) was used for reverse transcription. qRT-PCR was conducted using SYBR GreenER qPCR SuperMix Universal (Takara).
Western Blotting
Samples were lysed in radioimmunoprecipitation assay (RIPA) lysis buffer (Thermo Fisher). Protein was resolved in SDS-PAGE and transferred to polyvinylidene difluoride membranes and probed with primary and secondary antibodies. The antibodies used in this study as follows: the antibodies targeting RPS6, phospho-RPS6 (Ser235), mTOR, phospho-mTOR (Ser2448), GPX4 and Rabbit Anti-Mouse IgG H&L (HRP) were purchased from Abcam (Shanghai, China). TEAD1 and GAPDH antibodies were obtained from Proteintech Group (Wuhan, China).
Immunohistochemistry Staining (IHC)
Paraffin-embedded tissue sections were dewaxed, hydrated and antigen recovered successively. Next, the slides were quenched, and blocked, Then, TEAD1 antibody was added and incubated overnight at 4 °C, then reacted with secondary antibodies and detected with ABC-HRP kits. Representative images were obtained using a microscope.
Cell Counting Kit-8 (CCK8) Assay
Transfected HCC cells were seeded in 96-well plates at a concentration of 2 × 104 cells/well, and then treated with sorafenib for indicated times. Then, 10 μl of CCK8 reagent (Meilun, Dalian, China) was added and incubated at 37 °C for 2 h. The absorbance was detected at 450 nm using the microplate reader.
Wound Healing
The transfected cells were cultured in a 6-well plate to a 100% confluence monlayer. A 200-μL pipette tip was used to make a damage line in the petri dish. Images of the migrating cells were captured at 0 and 24 h for HCC cells. The average width of 8 random damage lines was measured and normalized to the 0 h control, which represents the relative migratory rate.
Transwell Assay
Transwell chambers were purchased from Corning Costar (USA). Transfected cells were serum-starved for 24 h, and 1 × 105 HCC cells in 200 μL medium without serum were placed onto the upper chambers pre-treated with Matrigel (BD Biosciences, CA, USA). Complete medium was added to the bottom chambers. After 24 h, the cells at the membrane were treated with 4% paraformaldehyde, and then stained with 0.5% crystal violet for 30 min. The invaded cells were photographed and counted in six random fields of view.
Intracellular Iron Assay
The intracellular iron was was detected with an Iron Assay Kit (Leagene, Bei**g, China). In brief, the cells were lysed and centrifuged, and the supernatant was taken for iron concentration determination. Reagents were added to 96-well plate according to kit’ instruction, and standard wells were set at the same time. After standing at room temperature for 15 min, the absorbance was measured at 562 nm with a microplate reader.
Lipid ROS Measurement
The cells were washed and incubated in BODIPY 581/591 C11 solution (Thermo Fisher, USA) for 30 min. Then the cells were collected and re-suspended for flow cytometry analysis.
Detection of Malondialdehyde (MDA)
After transfection or drug treatment, the cells were rinsed with PBS, and then lysated in shaking for 2 h. The MDA concentration was measured using Lipid Peroxidation (MDA) Assay Kit (Beyotime, China) following the manufacturer’s instructions. Absorbance was measured at 532 nm.
GSH Measurement
The content of GSH was measured using GSH Assay Kit (Beyotime, Shanghai,China) according to the instruction. The cells were treated with protein removal reagents, freezed-thawed twice, and then placed in ice bath for 5 min. Followed centrifugatng, the supernatant was used for the determination of GSH. The reagents and samples were added into 96-well plate in turn according to the instructions, mixed and incubated for 5 min at room temperature. After adding NADPH solution, the absorbance at 412 nm was detected by a microplate reader. The standard curve was drawn according to the different absorbance measured by different concentrations of standard materials. The total glutathione or GSSG content was calculated by comparing the sample to the standard curve. GSH = Total Glutathione-GSSG × 2.
Mitochondrial Membrane Potential Assay
Mitochondrial membrane potential assay kit with JC-1 was used for MMP measurement. After washing with PBS, the cells in the 6-well plate were added with 1 mL 1640 medium, 1 mL JC-1 staining solution, and incubated at 37 °C for 20 min. The supernatant was removed, cells were washed twice with JC-1 staining buffer, and added with 2 mL cell culture medium. JC-1 monomers and aggregates were observed at 490 nm and 525 nm under fluorescence microscope, respectively.
Chromatin Immunoprecipitation (ChIP) Assay
To detect TEAD1 and SLC3A2-binding sites, The ChIP kit (Abcam, USA) was used for ChIP assay. Briefly, BEL-7405 cells were collected and crosslinked with 1% formalin, lysed with SDS buffer, and DNA was fragmented by sonication. An anti-TEAD1 antibody (Proteintech, China) was used to precipitate protein/DNA complexes, and normal isotype-matched IgG was used as a negative control. The DNA was extracted and analyzed using real-time quantitative PCR.
Luciferase Reporter Gene Assay
Luciferase reporter vectors SLC3A2-wild-type (WT) and SLC3A2-mutant (MT) were constructed using PmirGLO (Promega, USA). Lipofectamine™ 2000 (Invitrogen, Carlsbad, California) was used to co-transfect SLC3A2-WT or SLC3A2-MT vectors with TEAD1 overexpressed plasmids. After 72 h of co-transfection, the Luciferase activity was detected using dual-Luciferase Reporter Assay System.
Tumor Xenografts
The BALB/c mice (four week-old) were obtained from West China Medical Center, Sichuan University (Chengdu, China). Hep-3B cells stably knocked down for TEAD1 and control cells were collected, and injected subcutaneously into the flank of nude mice with 2.5 × 106 cells. One week after inoculation, control or TEAD1-silenced mice were randomly divided into two groups to receive either vehicle or sorafenib treatment. The maximum diameter (a) and minimum width (b) of tumors were measured every 4 days and tumor volume was calculated as 0.5 × a × b2. After 30 days, the mice were sacrificed and the tumors were seperated and sliced for pathological and molecular analysis. The animal experiment protocol was approved by the Research Ethics Committee of the Sichuan Provincial People's Hospital (No. 2021363).
Statistical Analysis
In this study, GraphPad Prism (GraphPad Software, Inc.) was used for data statistics and analysis. Data are shown as mean ± SEM. At least three independent replicates were performed for each group. The student t-test and one-way analysis of variance were used for comparison between two or multiple groups. p < 0.05 was considered statistically significant.
Results
Expression of TEAD1 Is Upregulated in HCC
Analysis from UALCAN web resource revealed that the transcriptional levels of TEAD1 were upregulated in primary liver hepatocellular carcinoma (LIHC) tissues when compared to that of normal samples (Fig. 1A). Then, we determined the TEAD1 expression in 30 pairs of fresh HCC tissues and adjacent normal tissues. The qRT-PCR results of clinical tissue samples demonstrated that the levels of TEAD1 mRNA in HCC tissues were higher than that in adjacent non-cancerous tissues (Fig. 1B). And western blot showed the expression of TEAD1 protein was also upregulated in HCC tissues compared with adjacent non-cancerous samples (Fig. 1C). Besides, the expression of TEAD1 was examined in HCC cell lines and results showed that BEL-7405 had relatively lowest TEAD1 expression, while Hep-3B had highest TEAD1 expression (Fig. 1D, E).
TEAD1 expression in HCC tumor and matched adjacent normal tissues. A The levels of TEAD1 in HCC and normal tissues from UALCAN website. B RT-PCR analysis of TEAD1 mRNA in HCC tissues and matched normal tissues (n = 30). C Western blot results of TEAD1 protein in tissues and matched normal tissues. The representative images of Western blot (left). The plot of TEAD1 expression (n = 30). D The expression of TEAD1 mRNA in HCC cells, by RT-PCR. E The expression of TEAD1 protein in HCC cells, by Western blot. N normal tissues. T Tumor tissues
TEAD1 Promotes Cell Viability, Migration and Invasion in HCC Cells
To explore the role of TEAD1 in HCC cells, Hep-3B cells with highest TEAD1 expression were transfected with TEAD1 siRNAs (si-TEAD1#1, #2, #3) and BEL-7405 cells with lowest TEAD1 expression were transfected with TEAD1-expressing plasmids (TEAD1_OE). As shown in Figure S1A, B, si-TEAD1#1 showed the strongest inhibitory activity. Meantime, transfection of TEAD1_OE caused a significant increase of TEAD1 at the transcription and translation levels (Figures S1C, D). Next, the possible role of TEAD1 in cell viability, migration and invasion were assessed by CCK8, wound healing assay and Transwell invasion assays. TEAD1 depletion significantly repressed cell viability, motility and invasion in Hep-3B cells (Fig. 2A–C). On the contrary, overexpression TEAD1 markedly promoted cell viability, motility and invasion in BEL-7405 cells (Fig. 2D–F).
TEAD1 promotes proliferation, migration, and invasion in HCC Cells. After Hep-3B cells were transfected with si-TEAD1 or scrambled control (si-NC), A CCK8 assay was performed for cell viability, B wound healing was performed for cell migration, C Transwell invasion assay was conducted used for cell invasion. After BEL-7405 cells were transfected with TEAD1-overexpressing plasmids (TEAD1_OE) or empty vector (Vector), D CCK8 assay, E wound healing, and F Transwell invasion assays were conducted for cell viability, migration and invasion. **p < 0.01; ***p < 0.001
TEAD1 Regulates Sorafenib-Induced Ferroptosis
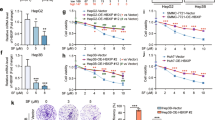
Next, we explored whether TEAD1 is related to sorafenib sensitivity in HCC cells. To determine this, HCC cells were treated with different concentrations of sorafenib for 48 h, and these cells included TEAD-overexpressed BEL-7405 cells or TEAD1-knockdown Hep-3B cells and the corresponding negative control groups. We observed that TEAD1 knockdown decreased IC50 from 12.73 to 8.56 μM in Hep-3B cells, indicating knockdown of TEAD1 enhanced the sensitivity of Hep-3B cells to sorafenib (Fig. 3A). Meanwhile, overexpressed TEAD1 increased IC50 from 10.64 μM to 15.53 μM in BEL-7405 cells, indicating TEAD1 decreased the sensitivity of BEL-7405 cells to sorafenib (Fig. 3B). Further, we found that sorafenib reduced the levels of total and nuclear TEAD1 in HCC cells, and increased the the levels of cytoplasma TEAD1, as evidenced by western blot (Fig. 3C). Moreover, immunofluorescence staining displayed that sorafenib treatment promoted the translocation of nuclear TEAD1 to the cytoplasm (Fig. 3D).
TEAD1 regulates ferroptosis. A The growth inhibition of Hep-3B/si-TEAD1 and Hep-3B/si-NC cells treated with various concentrations of Sorafenib was determined by CCK8 assay. B The growth inhibition of BEL-7405/TEAD1_OE and BEL-7405/vector cells treated with various concentrations of sorafenib was determined by CCK8 assay. C Western blot was performed to examine the expression of TEAD1 in nucleus and cytoplasm in HCC cells after treatment with 10 μM Sorafenib. D Immunofluorescent staining was used to observe the distribution of TEAD1 in HCC cells after treatment with 10 μM Sorafenib. E After BEL-7405/TEAD1_OE and BEL-7405/vector cells were treated with Sorafenib, ferrous iron, lipid ROS, MDA and GSH were analyzed. F After Hep-3B/si-TEAD1 and Hep-3B/si-NC cells were treated with Sorafenib, ferrous iron, lipid ROS, MDA and GSH were analyzed. G The cell viability of Hep-3B/si-TEAD1 and Hep-3B/si-NC cells treated with Fer-1 was determined by CCK8 assay. H After Hep-3B/si-TEAD1 and Hep-3B/si-NC cells were treated with Fer-1, ferrous iron, lipid ROS, MDA and GSH were analyzed. **p < 0.01; ***p < 0.001 verse the Vector or si-NC groups. ##p < 0.01; ###p < 0.001 verse the Vector or si-NC + SOR groups
Sorafenib has been identifed as an inducer of ferroptosis rather than apoptosis or necrosis. thus, we explored whether TEAD1 regulates Sorafenib-induced ferroptosis. As shown in Figs. 3D, Sorafenib remarkably increased intracellular redox-active iron accumulation, ROS generation, MDA content, and decreased GSH activity. However, TEAD1 overexpression not only reduced intracellular iron levels, ROS accumulation, MDA production, promoted GSH activity at basal level, but also significantly reversed the levels of these ferroptosis-related indicators induced by Sorafenib in BEL-7405 cells. Meanwhile, knockdown of TEAD1 resulted in increased levels of intracellular iron, ROS, and MDA, and a decreased levels of GSH upon basal culture or Sorafenib addition (Fig. 3E). In addition, we also examined the effect of TEAD1 on mitochondrial membrane potential (MMP). As shown in Figure S2, Sorafenib remarkably increased the formation of JC-1 monomer with green fluorescence, indicating MMP loss of HCC cells. However, TEAD1 overexpression reduced green fluorescence and increased the formation of JC-1 aggregates with red fluorescence, indicating that TEAD1 is essential for maintaining mitochondrial integrity.
To determine whether TEAD1 regulates ferroptosis in HCC cells, various specific inhibitors were applied to TEAD1-deficient HCC cells. The cell viability assay showed that ferroptosis inhibitors (Fer-1, 10 μM) alleviated impaired cell viability by si-TEAD1 transfection, while necrosis inhibitor Nec-1 (40 µM) displayed slight recovery on the cell viability, and apoptosis inhibitor Z-VAD-FMK (20 µM) displayed no obvious impact on the viability recovery in the TEAD1-deficient cells (Fig. 3G), suggesting that the cell death caused by TEAD1 knockdown is to a great extent due to ferroptosis. Meanwhile, compared to that of si-TEAD1 group, the addition of Fer-1 significantly decreased the levels of iron, ROS, and MDA, and increased GSH in TEAD1-deficient Hep-3B cells (Fig. 3H). Collectively, these data confirmed that TEAD1 is a crucial negative regulator of ferroptosis and represses Sorafenib-induced ferroptosis.
TEAD1 Regulates the Transcription of SLC3A2 in HCC Cells
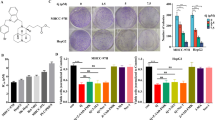
As a transcription factor, TEAD1 can specifically regulate the expression of downstream target genes. We used ChIPBase v2.0 (https://rna.sysu.edu.cn/chipbase/) database to screen potential target genes of TEAD1. Four ferroptosis-related genes, including SLC3A2, VDAC3, LPCAT3 and SLC39A8 were screened out (Fig. 4A). Analysis from UALCAN web resource showed the expression of SLC3A2 was markedly up-regulated in HCC (P = 1.63e-12, Fig. 4B), and was positively correlated with the expression of TEAD1 (P = 9.2e-05, R = 0.2, Fig. 4C). Furthermore, TEAD1 knockdown reduced SLC3A2 expression in Hep3B cells, while TEAD1 overexpression up-regulated SLC3A2 expression in BEL-2405 cells (Fig. 4D).
TEAD1 regulates the transcription of SLC3A2 in HCC cells. A Venn diagram reveals four ferroptosis-related genes, which were.potential target genes of TEAD1 predicted by ChIPBase database. B The mRNA expression levels of SLC3A2 in the LIHC samples was analyzed through UALCAN database. C The correlation between the mRNA expression levels of TEAD1 and SLC3A2 in the LIHC samples was analyzed through UALCAN database. D RT-PCR was used to measure the expression levels of TEAD1 protein in HCC cells after knockdowning or overexpressing TEAD1. E Western blot was used to measure the expression levels of TEAD1 protein in HCC cells after knockdowning or overexpressing TEAD1. F The schematic drawing shows the TEAD1 binding motifs on the SLC3A2 promoter. G The potential binding site between TEAD1 and SLC3A2 promoter was predicted by JASPAR bioinformatics website (http://jaspar.genereg.net/). H The dual-luciferase reporter assay verified the binding of TEAD1 in SLC3A2 promoter region in Hep-3B and BEL-7405 cells. I ChIP assay was performed to detect the enrichment of TEAD1 in SLC3A2 promoter region after overexpressing TEAD1 in BEL-7405 cells. J After BEL-7405 cells were co-treated with si-SLC3A2 and TEAD1_OE or alone, ferrous iron, lipid ROS, MDA and GSH were analyzed. *p < 0.05; **p < 0.01; ***p < 0.001 verse the control groups. #p < 0.05;##p < 0.01 verse the TEAD1_OE groups
To investigate whether SLC3A2 is a direct target gene of TEAD1, we analyzed the SLC3A2 promoter and identified two consensus binding motifs for TEAD1 through JASPAR online database (Fig. 4E), and the binding sites of TEAD1 and SLC3A2 promotor were shown in Fig. 4F. A 2 kb-long SLC3A2 promoter containing these two motifs was cloned into the pGL3 vector and the vector was then co-transfected into TEAD1-knockdown Hep-3B or TEAD1-overexpressed BEL-7405 cells. As shown in Fig. 4G–H, TEAD1 knockdown decreased the SLC3A2 promoter activity, while TEAD1 overexpression led to a reduced SLC3A2 promoter activity. However, when the TEAD1 binding sites were mutated, the SLC3A2 promoter activity was almost unresponsive to TEAD1. Moreover, the ChIP-PCR data derived from TEAD1-overexpressed BEL-7405 cells demonstrated that the abundance of SLC3A2 promotor Site 1 and Site 2 regions were significantly increased in TEAD1 Ab immunoprecipitation complex, especially when TEAD1 is overexpressed (Fig. 4I), indicating TEAD1 binding to the endogenous SLC3A2 promoter region. Collectively, SLC3A2 is the TEAD1 target gene.
Considering the role of SLC3A2 in ferroptosis, we hypothesized that TEAD1 might be involved in the regulation of ferroptosis through SLC3A2. To verify this, BEL-7405 cells were co-transfected with si-SLC3A2 or TEAD1-expressing plasmids or alone, and then treated with Sorafenib. It was found that SLC3A2 knockdown reversed TEAD1-mediated ferroptosis inhibition (Fig. 4I).These results suggest that TEAD1-induced ferroptosis is SLC3A2 dependent.
TEAD1 Activates mTORC1 Signaling to Inhibit Ferroptosis and Sorafenib Sensitivity
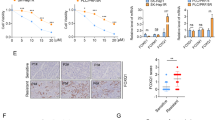
Previous studies have suggested that TEAD1 and SLC3A2 can activate mTORC1 signaling [18], which is associated with induction of ferroptosis [19]. Hence, we examined whether TEAD1/SLC3A2 axis regulates the mTORC1 signaling pathway in HCC. As displayed in Fig. 5A, overexpression of TEAD1 led to high levels of the phosphorylated mTOR (p-mTOR) and ribosomal protein S6 (p-RPS6) and increased the expression of GPX4 protein. Further, overexpression of TEAD1 could reverse the expression of p-mTOR, p-RPS6, and GPX4, restrained by sorafenib treatment. Moreover, knockdown of TEAD1 alone or in conjunction with sorafenib inhibited the expression of p-mTOR, p-RPS6, and GPX4 in Hep-3B cells (Fig. 5B).
TEAD1 activates mTORC1 signaling and increases GPX4 expression. A The expression of mTOR, RPS6, and GPX4 proteins in TEAD1-overexpressing BEL-7405 cells, examined by Western blot. B The expression of mTOR, RPS6, and GPX4 proteins in TEAD1-knockdown Hep-2B cells, examined by Western blot. After TEAD1-overexpressing BEL-7405 cells were co-treated with Sorafenib and Torin or Sorafenib alone, C Cell viability of CCK8. D Ferrous iron levels, E the level of lipid ROS, measured by C11-BODIPY staining coupled with flow cytometry, and F the levels of MDA, were analyzed. *p < 0.05; **p < 0.01; ***p < 0.001
Further, to confirm TEAD1 regulates ferroptosis via mTORC1 signaling, The TEAD1-overexpressing BEL-7405 cells were treated with sorafenib was applied to stimulate ferroptosis and then treated with mTORC1 inhibitor, Torin. We showed that TEAD1 overexpression recovered cell viability inhibited by sorafenib. However, Torin treatment reversed the effect of TEAD1 overexpression (Fig. 5C). Further, Analysis of ferroptosis markers showed that Torin treatment attenuated the decreased redox-active iron (Fig. 5D), lipid ROS (Fig. 5E), and MDA levels (Fig. 5F). Together, these results suggest that TEAD1 suppresses ferroptosis and enhances cell survival by activating mTORC1/GPX4 axis in HCC cells.
TEAD1 Depletion Suppresses Tumor Growth and Increased Ferroptosis In Vivo
The role of TEAD1 in HCC was further evaluated by xenograft model. TEAD1-knockdown Hep-3B cells or control cells were subcutaneously injected into the right flank of nude mice and treated with sorafenib for the first 7 days (Fig. 6A). As shown in Fig. 6B, both TEAD1 depletion and sorafenib treatment effectively reduced the volume of tumors, and the two treatments synergistically inhibited tumor growth. Tumor weight showed the same trend (Fig. 6C). Besides, both TEAD1 depletion and sorafenib treatment led to the inactivation of mTORC1 siganling and decreased GPX4 levels, and the two treatments work synergistically, shown by Western blot analysis (Fig. 6E). In vivo experiments confirmed that inhibition of TEAD1 enhanced the sensitive to sorafenib via mTORC1/GPX4 in HCC.
TEAD1 knockdown inhibits growth of xenograft in vivo. A Hep-3B cells with TEAD1 knockdown or control were injected subcutaneously into the flank of nude mice and then treated with sorafenib or PBS (six per group). B The growth curves of tumor volume in each group. Tumor volume: 0.5 × a × b2. C The images of tumors. D The tumor weight in each groups (n = 6). E The expression of TEAD1, mTOR, RPS6 and GPX4 in tumor tissues of each group. *p < 0.05; **p < 0.01; ***p < 0.001
Discussion
Although a large amount of evidence indicates that TEAD1 expression is related to the progression and drug resistance of malignant tumors, the role and molecular mechanism of TEAD1 in HCC are still obscure. In this study, we found that TEAD1 was upregulated in HCC, and upregulation of TEAD1 promoted cell proliferation, migration, and invasion in HCC cells. Further, upregulation of TEAD1 alleviated sorafenib-induced ferroptosis. Mechnistically, TEAD1 promotes the transcription of SLC3A2 and activates the mTORC1 signaling. These findings highlight the potential of TEAD1 as a possible therapeutic target for HCC treatment.
Many studies demonstrated the elevated expression and critical roles of TEAD1 in cancer development. It has been reported that TEAD1 enhanced tumor growth and proliferation via activating SP1 in colorectal cancer [20]. TEAD1 was involved in the oncogenic activities of Hippo-YAP1 signaling in osteosarcoma [21]. Previous studies revealed that high TEAD1 expression was observed in HCC tissues and associated with histological grade, tumor size, and serum a-fetoprotein (AFP) concentration of HCC patients [16, 22, 23]. Here, consistent with these findings, our data showed that the expression of TEAD1 was increased in HCC tissues. TEAD1 overexpression promoted malignant biological behaviors, while the depletion of TEAD1 repressed malignant phenotype of HCC cells. These data confirmed the oncogenic activities of TEAD1 in HCC.
Sorafenib is widely used as a first-line anticancer agent for advanced HCC. Recent studies have shown that sorafenib-treated HCC cells exhibit specific ferroptosis-related events, such as lipid peroxidation, glutathione consumption, and iron accumulation, which lead to cell death [24]. Therefore, inducing ferroptosis can enhance the anticancer effect of sorafenib and may be a new therapeutic strategy for HCC. TEAD1, together with its transcription co-activators YAP and TAZ, are the important participants of the Hippo signaling pathway. The hippo pathway is an effective tumor-suppressor, and its abnormality contributes to apoptosis evasion of cancer cells, cancer progression, metastasis, and chemoresistance. Recent studies identified YAP/TAZ and Hippo pathways as crucial determinants of ferroptosis in several cancers including ovarian cancer, lung adenocarcinoma., breast tumors, renal cell carcinoma [25,26,27,28,29,30,31]. For example, Gao et al. [32] revealed YAP/TAZ conferred the resistance of HCC cells to sorafenib via inhibiting ferroptosis. Yang et al. [33] reported that the Hippo pathway effector TAZ promoted sensitizes cells to ferroptosis. through the regulation of EMP1-NOX4 in renal cell carcinoma. However, the role of TEAD1 in ferroptosis regulation remains unclear. Here, our study showed that TEAD1 overexpression decreased sorafenib sensitivity and conferred the resistance of HCC cells to ferroptosis, while TEAD1 knockdown promoted ferroptosis, acting synergically with Sorafenib. Further studies showed that si-TEAD1-induced ferroptosis could be reversed by ferroptosis inhibitor. Collectively, these data suggest that TEAD1 is an important negative regulator of ferroptosis.
As a transcription factor, TEAD1 can directly activate transcription of many genes in physiological or pathological states [34]. For example, TEAD1 directly activates a large set of nuclear DNA-encoded mitochondrial genes in adult hearts [35]. TEAD1 upregulatd TWIST1 expression at the transcriptional level, and thereby promoting EMT in clear cell renal cell carcinoma [14]. In our study, through CHIP assay and luciferase report assay, we demonstrated that TEAD1 transcriptionally activated the SLC3A2 expression. SLC3A2 is subunits of the glutamate-cystine antiporter system xc(-). Upregularion of SLC3A2 inhibits ferroptosis, while downregulated SLC3A2 sensitized cells to ferroptosis activators [36]. We also confirmed that downregulated SLC3A2 attenuated the inhibitory effect of TEAD1 on ferroptosis.
mTOR is the central controller of protein synthesis, cell growth, proliferation, and survival. Abnormal activation of the mTOR pathway occurs at high frequency in human cancers and has multiple roles in tumor progression [37,38,39]. Recent studies identified mTORC1 as a key ferroptosis modulator [19]. Zhang et al. [40] revealed the crucial role of mTORC1 signaling in coordinating GPX4 protein synthesis and inhibition of mTORC1 decreased GPX4 expression and sensitize cancer cells to ferroptosis in renal cell carcinoma. Ni et al. revealed [41] that mTORC1 signaling was upregulated in Lapatinib resistant NSCLC cells, and inhibition of mTORC1 overcomes resistance to Lapatinib-induced ferroptosis via downregulating GPX4. Herein, we found that TEAD1 activated mTORC1 signaling and increased GPX4 protein levels, while Torin, mTORC1 inhibitor could reverse the inhibitory effect of TEAD1 on redox-active iron, lipid ROS, and MDA, as well as GPX4. These data indicate that TEAD1 confers the resistance of HCC to ferroptosis through activation of the mTORC1/GPX4 axis.
Conclusion
Our study disclosed that TEAD1 promotes the malignant behaviors and of HCC cells by resisting ferroptosis, and TEAD1 depletion enhances Sorafenib-induced ferroptosis. TEAD1 promotes the transcription of SLC3A2 and activates mTORC1 signaling, which might be the potential mechanism underlying TEAD1-regulated ferroptosis in HCC. Collectively, this study elucidated the molecular mechanism of ferroptosis-mediated HCC and provided targets for HCC treatments.
Data Availability
The datasets generated during and/or analysed during the current study are not publicly available, and are available from the corresponding author on reasonable request.
References
Hino, K., Iron and liver cancer: an inseparable connection. Adv Sci (Weinh), 2021.
Yanatori I, Hara Y, Nishina S, et al. Ferroptosis in the tumor microenvironment: perspectives for immunotherapy. Febs j 2021;27(9:856–867.
Yao, X., Li W, Fang D, et al., Emerging Roles of Energy Metabolism in Ferroptosis Regulation of Tumor Cells. 2021:e2100997.
Sun J, Zhou C, Zhao Y, et al. Quiescin sulfhydryl oxidase 1 promotes sorafenib-induced ferroptosis in hepatocellular carcinoma by driving EGFR endosomal trafficking and inhibiting NRF2 activation. Redox Biol 2021;41:101942.
Wang, Q, Bin C, Xue Q, et al., GSTZ1 sensitizes hepatocellular carcinoma cells to sorafenib-induced ferroptosis via inhibition of NRF2/GPX4 axis. 2021;12(5):426.
Asperti M, Bellini S, Grillo E, et al. H-ferritin suppression and pronounced mitochondrial respiration make Hepatocellular Carcinoma cells sensitive to RSL3-induced ferroptosis. Free Radic Biol Med 2021;169:294–303.
Wang Q, Guo Y, Wang W, et al. RNA binding protein DAZAP1 promotes HCC progression and regulates ferroptosis by interacting with SLC7A11 mRNA. Exp Cell Res 2021;399(1):112453.
Huang W, Chen K, Lu Y, et al. ABCC5 facilitates the acquired resistance of sorafenib through the inhibition of SLC7A11-induced ferroptosis in hepatocellular carcinoma. Neoplasia 2021;23(12):1227–1239.
Li B, Wei S, Yang L, et al. CISD2 Promotes Resistance to Sorafenib-Induced Ferroptosis by Regulating Autophagy in Hepatocellular Carcinoma. Front Oncol 2021;11:657723.
Lin KC, Park HW, Guan KL. Regulation of the Hippo Pathway Transcription Factor TEAD. J Med Chem 2017;42(11):862–872.
Heng BC, Zhang X, Aubel D, et al. An overview of signaling pathways regulating YAP/TAZ activity. 2021;78(2):497–512.
Cai X, Yu L, Chen Z et al. Arsenic trioxide-induced upregulation of miR-1294 suppresses tumor growth in hepatocellular carcinoma by targeting TEAD1 and PIM1. Cancer Biomark 2020;28(2):221–230.
Jiang, N, Zhao L, Zong D, et al. Long non-coding RNA LUADT1 promotes nasopharyngeal carcinoma cell proliferation and invasion by downregulating miR-1207–5p. Bioengineered, 2021.
Yin L, Li W, Xu A, et al. SH3BGRL2 inhibits growth and metastasis in clear cell renal cell carcinoma via activating hippo/TEAD1-Twist1 pathway. EBioMedicine 2020;51:102596.
Gibault F, Sturbaut M, Bailly F, et al. Targeting Transcriptional Enhanced Associate Domains (TEADs). Cell Mol Life Sci 2018;61(12):5057–5072.
Lin, W, Zhang T, Ding G, et al. Circular RNA circ CT3 promotes hepatocellular carcinoma progression by regulating the miR-1287-5p/TEAD1/PTCH1/LOX axis. Mol Med Rep, 2021;23(5).
Chandrashekar DS, Bashel B, Balasubramanya SAH, et al. UALCAN: A Portal for Facilitating Tumor Subgroup Gene Expression and Survival Analyses. Neoplasia 2017;19(8):649–658.
Osman I, He X, Liu J, et al. TEAD1 (TEA Domain Transcription Factor 1) Promotes Smooth Muscle Cell Proliferation Through Upregulating SLC1A5 (Solute Carrier Family 1 Member 5)-Mediated Glutamine Uptake. Circ Res 2019;124(9):1309–1322.
Lei G, Zhuang L, Gan B. mTORC1 and ferroptosis: Regulatory mechanisms and therapeutic potential. Bioessays 2021;43(8):e2100093.
Yu MH, Zhang W. TEAD1 enhances proliferation via activating SP1 in colorectal cancer. Biomed Pharmacother 2016;83:496–501.
Chai J, Xu S, Guo F. TEAD1 mediates the oncogenic activities of Hippo-YAP1 signaling in osteosarcoma. Biochem Biophys Res Commun 2017;488(2):297–302.
Hu X, He B, Zhou L, et al. Elevated TEAD1 Expression is not an Independent Prognosis Factor of Hepatocellular Carcinoma. Clin Lab 2018;64(5):743–748.
Lin Y, Huang G, ** H, et al. Circular RNA Gprc5a Promotes HCC Progression by Activating YAP1/TEAD1 Signalling Pathway by Sponging miR-1283. Int J Cancer 2020;13:4509–4521.
Liu H, Zhao L, Wang M, et al. FNDC5 Causes Resistance to Sorafenib by Activating the PI3K/Akt/Nrf2 Pathway in Hepatocellular Carcinoma Cells. Front Oncol 2022;12:852095.
Fang K, Du S, Shen D, et al. SUFU suppresses ferroptosis sensitivity in breast cancer cells via Hippo/YAP pathway. iScience 2022;25(7):104618.
Wu J, Minikes AM, Gao M, et al. Intercellular interaction dictates cancer cell ferroptosis via NF2-YAP signalling. Nature 2019;572(7769):402–406.
Yang WH, Huang Z, Wu J, et al. A TAZ-ANGPTL4-NOX2 Axis Regulates Ferroptotic Cell Death and Chemoresistance in Epithelial Ovarian Cancer. Mol Cancer Res 2020;18(1):79–90.
Yang, W.H. and C.C. Lin, The Hippo Pathway Effector YAP Promotes Ferroptosis via the E3 Ligase SKP2. 2021;19(6):1005–1014.
Yang Y, Li X, Wang T, et al. Emerging agents that target signaling pathways in cancer stem cells. 2020;13(1):60.
Zhang X, Yu K, Ma L, et al. Endogenous glutamate determines ferroptosis sensitivity via ADCY10-dependent YAP suppression in lung adenocarcinoma. Oncogene 2021;11(12):5650–5674.
Zheng L, Lin CC. DDR2 upregulation confers ferroptosis susceptibility of recurrent breast tumors through the Hippo pathway. J Hematol Oncol 2021;40(11):2018–2034.
Gao, R,Kalathur RKR, Coto-Llerena M, et al., YAP/TAZ and ATF4 drive resistance to Sorafenib in hepatocellular carcinoma by preventing ferroptosis. 2021;e14351.
Yang WH, Ding CC, Sun T, et al. The Hippo Pathway Effector TAZ Regulates Ferroptosis in Renal Cell Carcinoma. Cell Rep 2019;28(10):2501–2508.e4.
Wang, Y. and L. Liu, Unveiling E2F4, TEAD1 and AP-1 as regulatory transcription factors of the replicative senescence program by multi-omics analysis. 2022.
Liu, J, Wen T, Dong K, et al., TEAD1 protects against necroptosis in postmitotic cardiomyocytes through regulation of nuclear DNA-encoded mitochondrial genes. 2021;28(7):2045–2059.
Wu F, **ong G, Chen Z, et al. SLC3A2 inhibits ferroptosis in laryngeal carcinoma via mTOR pathway. Hereditas 2022;159(1):6.
Wei S, Dai M, Zhang C, et al. KIF2C: a novel link between Wnt/β-catenin and mTORC1 signaling in the pathogenesis of hepatocellular carcinoma. Protein Cell 2021;12(10):788–809.
Zhang, S, Y.F. Zhou, and J. Cao, mTORC1 Promotes ARID1A Degradation and Oncogenic Chromatin Remodeling in Hepatocellular Carcinoma. 2021;81(22):5652–5665.
Zheng, XFS, Chao X, Wang S, et al., Hepatocytic p62 suppresses ductular reaction and tumorigenesis in mouse livers with mTORC1 activation and defective autophagy. Cancer Res, 2021.
Zhang, Y, R.V. Swanda, and L. Nie, mTORC1 couples cyst(e)ine availability with GPX4 protein synthesis and ferroptosis regulation. 2021;12(1):1589.
Ni J, Chen K, Zhang J, et al. Inhibition of GPX4 or mTOR overcomes resistance to Lapatinib via promoting ferroptosis in NSCLC cells. Biochem Biophys Res Commun 2021;567:154–160.
Funding
This study was approved by Applied Basic Research Program of Science and Technology Department of Sichuan Province (No. 2019YJ0588) and Wu Jie** Medical Foundation (No. 320.6750.2020-01-42).
Author information
Authors and Affiliations
Contributions
HL and CY contributed to the study conception and design. Material preparation, data collection and analysis were performed by HL, MZ, FZ and NA. The first draft of the manuscript was written by HL and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interest
The authors have no relevant financial or non-financial interests to disclose.
Consent to participate
Informed consent was obtained from all individual participants included in the study (No. 20210519).
Ethical approval
This study was performed in line with the principles of the Declaration of Helsinki. This study was approved by the Research Ethics Committee of the Sichuan Provincial People's Hospital (No. 2021363).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
10620_2023_7824_MOESM1_ESM.jpg
Figure S1 The expression of TEAD1 after transfection with TEAD1-overexpressing plasmids or si-TEAD1. (A) The mRNA expression of TEAD1 in Hep3B after transfection with si-TEAD1. (B) The protein expression of TEAD1 in Hep3B after transfection with si-TEAD1. (A) The mRNA expression of TEAD1 in BEL-7405 after transfection with TEAD1-overexpressing plasmids. (B) The protein expression of TEAD1 in BEL-7405 after transfection with TEAD1-overexpressing plasmids. ***p<0.001. Supplementary file1 (JPG 376 kb)
10620_2023_7824_MOESM2_ESM.tif
Figure S2 Effect of TEAD1 on mitochondrial membrane potential. After BEL-7405/TEAD1_OE and BEL-7405/vector cells were treated with Sorafenib, JC-1 fluorescent probe was used to detect mitochondrial membrane potential. Supplementary file2 (TIF 20235 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License, which permits any non-commercial use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc/4.0/.
About this article
Cite this article
Li, H., Lan, H., Zhang, M. et al. TEA Domain Transcription Factor 1 Inhibits Ferroptosis and Sorafenib Sensitivity of Hepatocellular Carcinoma Cells. Dig Dis Sci 68, 3070–3082 (2023). https://doi.org/10.1007/s10620-023-07824-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10620-023-07824-5