Abstract
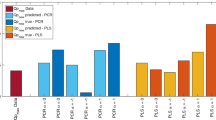
Various mechanistic and black-box models were applied for on-line estimations of viable cell concentrations in fed-batch cultivation processes for CHO cells. Data from six fed-batch cultivation experiments were used to identify the underlying models and further six independent data sets were used to determine the performance of the estimators. The performances were quantified by means of the root mean square error (RMSE) between the estimates and the corresponding off-line measured validation data sets. It is shown that even simple techniques based on empirical and linear model approaches provide a fairly good on-line estimation performance. Best results with respect to the validation data sets were obtained with hybrid models, multivariate linear regression technique and support vector regression. Hybrid models provide additional important information about the specific cellular growth rates during the cultivation.






Similar content being viewed by others
Abbreviations
- μ:
-
Specific growth rate (h−1)
- q P :
-
Specific product formation rate (g e9cells−1 h−1)
- X :
-
Cell concentration (e9cells L−1)
- X V :
-
Viable cell concentration (e9cells L−1)
- x :
-
Cells count (e9cells)
- m P :
-
Mass of product (g)
- W :
-
Culture weight (kg)
- ρ:
-
Culture density (kg L−1)
- t :
-
Process time (h)
- OUR:
-
Oxygen uptake rate (mmol kg−1 h−1)
- CPR:
-
Carbon dioxide production rate (mmol kg−1 h−1)
- tOUR:
-
Total oxygen uptake rate (mmol h−1)
- tCPR:
-
Total carbon dioxide production rate (mmol h−1)
- tcOUR:
-
Total cumulative O2 uptake rate (mmol)
- tcCPR:
-
Total cumulative CO2 production rate (mmol)
- Base:
-
Consumed Base (g, kg)
- Y OX :
-
Yield O2/Cells, Y OX = 10.34 (mmol e9cells−1)
- Y CX :
-
Yield CO2/Cells, Y CX = 11.37 (mmol e9cells−1)
- m O :
-
Maintenance for O2, m O = 0.001 (mmol e9cells−1 h−1)
- m C :
-
Maintenance for CO2, m C = 0.001 (mmol e9cells−1 h−1)
- Y bX :
-
Yield Base consumption/Cells Ybx = 9.0e-3 (kg e9cells−1)
- RMSE:
-
Root mean square error (e9cells kg−1)
- w.a.:
-
Weighted average of Base, tOUR, tCPR related models
References
Behrendt U, Koch S, Gooch DD, Steegmans U, Comer MJ (1994) Mass spectrometry: a tool for on-line monitoring of animal cell cultures. Cytotechnol 14:157–162
Bibila TA, Robinson DK (1995) In pursuit of the optimal fed-batch process for monoclonal antibody production. Biotechnol Progr 11:1–13
Brinkmann M, Lütkemeyer D, Gudermann F, Lehmann J (2002) New technologies for automated cell counting based on optical image analysis `The Cellscreen’. Cytotechnology 38:119–127
Cannizzaro C, Gügerli R, Marison I, Stockar Uv (2003) On-line biomass monitoring of CHO perfusion culture with scanning dielectric spectroscopy. Biotechnol Bioeng 84(5):597–610
Chang C-C, Lin CJ (2001) LIBSVM: a library for support vector machines, Software available at http://www.csie.ntu.edu.tw/~cjlin/libsvm
Chu L, Robinson DK (2001) Industrial choice for protein production by large-scale cell culture. Curr Op Biotechnol 12:180–187
Dorresteijn RC, Harbrink Numan K, de Gooijer CD, Tramper J, Beuvery EC (1996) On-line estimation of the biomass activity during animal-cell cultivations. Biotechnol Bioeng 51(2):206–214
Drucker H, Burges CJC, Kaufman L, Smola A, Vapnik V (1997) Support vector regression machines. Adv Neural Inf Process Syst 9:155–161
Ducommun P, Bolzonella I, Rhiel M, Pugeaud P, Uv Stockar, Marison IW (2001) On-line determination of animal cell concentration. Biotechnol Bioeng 72(5):515–522
Eyer K, Heinzle E (1996) On-line estimation of viable cells in a hybridoma culture at various DO levels using ATP balancing and redox potential measurement. Biotechnol Bioeng 49(3):277–283
FDA (2004) Guidance for industry: PAT—a framework for innovative pharmaceutical manufacturing and quality assurance, http://www.fda.gov/cvm/guidance/published.html
Gunther JC, Conner JS, Seborg SE (2008) PLS pattern matching in design of experiment, batch process data. Chemomet Intell Lab Syst 94(1):43–50
Hornik K, Stinchcombe M, White H (1989) Multilayer feed-forward networks are universal approximators. Neural Netw 2:359–366
Hossler P, Khattak SF, Li ZJ (2009) Optimal and consistent protein glycosylation in mammalian cell culture. Glycobiology 19(9):936–949
Jenzsch M, Simutis R, Eisbrenner G, Stückrath I, Lübbert A (2006) Estimation of biomass concentrations in fermentation processes for recombinant protein production. Bioproc Biosyst Eng 29(1):19–27
Joeris K, Frerichs J-G, Konstantinov K, Scheper T (2002) In-situ microscopy: on-line process monitoring of mammalian cell cultures. Cytotechnology 38:129–134
Junker BH, Wang HY (2006) Bioprocess monitoring and computer control: key roots of the current PAT initiative. Biotechnol Bioeng 95(2):226–261
Konstantinov KB, Pambayun R, Matanguihan R, Yoshida T, Perusich CM, Hu W-S (1992) On-line monitoring of hybridoma cell growth using a laser turbidity sensor. Biotechnol Bioeng 40:1337–1342
Luedeking R, Piret EL (1959) A kinetic study of lactic acid fermentation. Batch Process at controlled pH. J Biochem Microb Technol 1(4):393–412
Oeggerli A, Eyer K, Heinzle E (1994) On-line gas analysis in animal cell cultivation: I. Control of dissolved oxygen and pH. Biotechnol Bioeng 45:42–53
Schubert J, Simutis R, Dors M, Havlik I, Lübbert A (1994) Bioprocess optimization and control: application of hybrid modeling. J Biotechnol 35:51–68
Schügerl K (2001) Progress in monitoring, modeling and control of bioprocesses during the last 20 years. J Biotechnol 85(2):149–173
Shah D, Naciri M, Clee P, Al-Rubeai M (2006) NucleoCounter—an efficient technique for the determination of cell number and viability in animal cell culture processes. Cytotechnology 51:39–44
Simutis R, Oliveira R, Manikowski M, Feyo de Azevedo S, Lübbert A (1997) How to increase the performance of models for process optimization and control. J Biotechnol 59:73–89
Teixeira AP, Portugal CAM, Carinhas N, Dias JML, Crespo JP, Alves PM, Carrondo MJT, Oliveira R (2009) In situ 2D fluorometry and chemometric monitoring of mammalian cell cultures. Biotechnol Bioeng 102(4):1098–1106
Walsh G (2006) Biopharmaceutical benchmarks. Nat Biotechnol 24(7):769–776
Wlaschin KF, Hu W-S (2006) Fed-batch culture and dynamic nutrient feeding. Adv Biochem Eng/Biotechnol 101:43–74
Zabriskie DW, Humphrey AE (1978) Real-time estimation of aerobic batch fermentation biomass concentration by component balancing. AIChE J 24:138–146
Acknowledgments
The financial support from Ministry of Science and Education by means of the Excellence-Initiative Sachsen-Anhalt is gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Aehle, M., Simutis, R. & Lübbert, A. Comparison of viable cell concentration estimation methods for a mammalian cell cultivation process. Cytotechnology 62, 413–422 (2010). https://doi.org/10.1007/s10616-010-9291-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10616-010-9291-z