Abstract
Purpose
An increasing number of long intergenic non-coding RNAs (lincRNAs) appear to play critical roles in cancer development and progression. To assess the association between SNPs that reside in regions of lincRNAs and breast cancer risk, we performed a large case-control study in China.
Methods
We carried out a two-stage case-control study including 2881 breast cancer cases and 3220 controls. In stage I, we genotyped 17 independent (r2 < 0.5) SNPs located in 6 tumor-related lincRNAs by using the TaqMan platform. In stage II, SNPs potentially associated with breast cancer risk were replicated in an independent population. Quantitative real-time PCR was used to measure H19 levels in tissues from 228 breast cancer patients with different genotypes.
Results
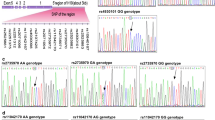
We identified 2 SNPs significantly associated with breast cancer risk in stage I (P < 0.05), but not significantly replicated in stage II. We combined the data from stage I and stage II, and found that, compared with the rs2071095 CC genotype, AA and CA + AA genotypes were associated with significantly decreased risk of breast cancer (adjusted OR 0.83, 95% CI 0.69–0.99; adjusted OR 0.88, 95% CI 0.80–0.98, respectively). Stratified analyses showed that rs2071095 was associated with breast cancer risk in estrogen receptor (ER)-positive patients (P = 0.002), but not in ER-negative ones (P = 0.332). Expression levels of H19 in breast cancer cases with AA genotype were significantly lower than those with CC genotype.
Conclusions
We identified that rs2071095 may contribute to the susceptibility of breast cancer in Chinese women via affecting H19 expression. The mechanisms underlying the association remain to be investigated.




Similar content being viewed by others
Data availability statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Chen W, Zheng R, Baade PD et al (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66:115–132
Pennisi E (2011) The Biology of Genomes. Disease risk links to gene regulation. Science 332:1031
Zheng J, Huang X, Tan W et al (2016) Pancreatic cancer risk variant in LINC00673 creates a miR-1231 binding site and interferes with PTPN11 degradation. Nat Genet 48:747–757
Kumar V, Westra HJ, Karjalainen J et al (2013) Human disease-associated genetic variation impacts large intergenic non-coding RNA expression. PLoS Genet 9:e1003201
Cabili MN, Trapnell C, Goff L et al (2011) Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25:1915–1927
Iyer MK, Niknafs YS, Malik R et al (2015) The landscape of long noncoding RNAs in the human transcriptome. Nat Genet 47:199–208
Geisler S, Coller J (2013) RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. Nat Rev Mol Cell Biol 14:699–712
Khalil AM, Guttman M, Huarte M et al (2009) Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc Natl Acad Sci USA 106:11667–11672
Rinn JL, Kertesz M, Wang JK et al (2007) Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129:1311–1323
Nagano T, Mitchell JA, Sanz LA et al (2008) The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322:1717–1720
Guttman M, Donaghey J, Carey BW et al (2011) lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477:295–300
Gupta RA, Shah N, Wang KC et al (2010) Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464:1071–1076
Bhan A, Soleimani M, Mandal SS (2017) Long noncoding RNA and cancer: a new paradigm. Cancer Res 77:3965–3981
Bhan A, Hussain I, Ansari KI et al (2013) Antisense transcript long noncoding RNA (lncRNA) HOTAIR is transcriptionally induced by estradiol. J Mol Biol 425:3707–3722
Huarte M, Guttman M, Feldser D et al (2010) A large intergenic noncoding RNA induced by p53 mediates global gene repression in the p53 response. Cell 142:409–419
Bayram S, Sumbul AT, Batmaci CY et al (2015) Effect of HOTAIR rs920778 polymorphism on breast cancer susceptibility and clinicopathologic features in a Turkish population. Tumour Biol 36:3863–3870
Peng R, Luo C, Guo Q et al (2018) Association analyses of genetic variants in long non-coding RNA MALAT1 with breast cancer susceptibility and mRNA expression of MALAT1 in Chinese Han population. Gene 642:241–248
Huang NS, Chi YY, Xue JY et al (2016) Long non-coding RNA metastasis associated in lung adenocarcinoma transcript 1 (MALAT1) interacts with estrogen receptor and predicted poor survival in breast cancer. Oncotarget 7:37957–37965
Timofeeva MN, Hung RJ, Rafnar T et al (2012) Influence of common genetic variation on lung cancer risk: meta-analysis of 14 900 cases and 29 485 controls. Hum Mol Genet 21:4980–4995
Royds JA, Pilbrow AP, Ahn A et al (2015) The rs11515 polymorphism is more frequent and associated with aggressive breast tumors with increased ANRIL and decreased p16 (INK4a) expression. Front Oncol 5:306
Riaz M, Berns EM, Sieuwerts AM et al (2012) Correlation of breast cancer susceptibility loci with patient characteristics, metastasis-free survival, and mRNA expression of the nearest genes. Breast Cancer Res Treat 133:843–851
Song F, Zheng H, Liu B et al (2009) An miR-502-binding site single-nucleotide polymorphism in the 3′-untranslated region of the SET8 gene is associated with early age of breast cancer onset. Clin Cancer Res 15:6292–6300
Bartolomei MS, Zemel S, Tilghman SM (1991) Parental imprinting of the mouse H19 gene. Nature 351:153–155
Seidl CI, Stricker SH, Barlow DP (2006) The imprinted Air ncRNA is an atypical RNAPII transcript that evades splicing and escapes nuclear export. EMBO J 25:3565–3575
Brannan CI, Dees EC, Ingram RS et al (1990) The product of the H19 gene may function as an RNA. Mol Cell Biol 10:28–36
Adriaenssens E, Dumont L, Lottin S et al (1998) H19 overexpression in breast adenocarcinoma stromal cells is associated with tumor values and steroid receptor status but independent of p53 and Ki-67 expression. Am J Pathol 153:1597–1607
Gabory A, Jammes H, Dandolo L (2010) The H19 locus: role of an imprinted non-coding RNA in growth and development. Bioessays 32:473–480
Cai X, Cullen BR (2007) The imprinted H19 noncoding RNA is a primary microRNA precursor. RNA 13:313–316
Kallen AN, Zhou XB, Xu J et al (2013) The imprinted H19 lncRNA antagonizes let-7 microRNAs. Mol Cell 52:101–112
Barsyte-Lovejoy D, Lau SK, Boutros PC et al (2006) The c-Myc oncogene directly induces the H19 noncoding RNA by allele-specific binding to potentiate tumorigenesis. Cancer Res 66:5330–5337
Ariel I, Sughayer M, Fellig Y et al (2000) The imprinted H19 gene is a marker of early recurrence in human bladder carcinoma. Mol Pathol 53:320–323
Sorin V, Ohana P, Gallula J et al (2012) H19-promoter-targeted therapy combined with gemcitabine in the treatment of pancreatic cancer. ISRN Oncol 2012: 351750
Adriaenssens E, Lottin S, Dugimont T et al (1999) Steroid hormones modulate H19 gene expression in both mammary gland and uterus. Oncogene 18:4460–4473
Basak P, Chatterjee S, Weger S et al (2015) Estrogen regulates luminal progenitor cell differentiation through H19 gene expression. Endocr Relat Cancer 22:505–517
Guo H, Ahmed M, Zhang F et al (2016) Modulation of long noncoding RNAs by risk SNPs underlying genetic predispositions to prostate cancer. Nat Genet 48:1142–1150
Maurano MT, Humbert R, Rynes E et al (2012) Systematic localization of common disease-associated variation in regulatory DNA. Science 337:1190–1195
Huang Q, Whitington T, Gao P et al (2014) A prostate cancer susceptibility allele at 6q22 increases RFX6 expression by modulating HOXB13 chromatin binding. Nat Genet 46:126–135
Zhang X, Cowper-Sal LR, Bailey SD et al (2012) Integrative functional genomics identifies an enhancer loo** to the SOX9 gene disrupted by the 17q24.3 prostate cancer risk locus. Genome Res 22:1437–1446
Cowper-Sal LR, Zhang X, Wright JB et al (2012) Breast cancer risk-associated SNPs modulate the affinity of chromatin for FOXA1 and alter gene expression. Nat Genet 44:1191–1198
Stacey SN, Sulem P, Masson G et al (2009) New common variants affecting susceptibility to basal cell carcinoma. Nat Genet 41:909–914
Iacobucci I, Sazzini M, Garagnani P et al (2011) A polymorphism in the chromosome 9p21 ANRIL locus is associated to Philadelphia positive acute lymphoblastic leukemia. Leuk Res 35:1052–1059
Healy J, Belanger H, Beaulieu P et al (2007) Promoter SNPs in G1/S checkpoint regulators and their impact on the susceptibility to childhood leukemia. Blood 109:683–692
Cunnington MS, Santibanez KM, Mayosi BM et al (2010) Chromosome 9p21 SNPs associated with multiple disease phenotypes correlate with ANRIL expression. PLoS Genet 6:e1000899
Motterle A, Pu X, Wood H et al (2012) Functional analyses of coronary artery disease associated variation on chromosome 9p21 in vascular smooth muscle cells. Hum Mol Genet 21:4021–4029
Holdt LM, Beutner F, Scholz M et al (2010) ANRIL expression is associated with atherosclerosis risk at chromosome 9p21. Arterioscler Thromb Vasc Biol 30:620–627
Pasmant E, Sabbagh A, Vidaud M et al (2011) ANRIL, a long, noncoding RNA, is an unexpected major hotspot in GWAS. FASEB J 25:444–448
Aguilo F, Zhou MM, Walsh MJ (2011) Long noncoding RNA, polycomb, and the ghosts haunting INK4b-ARF-INK4a expression. Cancer Res 71:5365–5369
Kotake Y, Nakagawa T, Kitagawa K et al (2011) Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. Oncogene 30:1956–1962
Yap KL, Li S, Munoz-Cabello AM et al (2010) Molecular interplay of the noncoding RNA ANRIL and methylated histone H3 lysine 27 by polycomb CBX7 in transcriptional silencing of INK4a. Mol Cell 38:662–674
Yu W, Gius D, Onyango P et al (2008) Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. Nature 451:202–206
Lan WG, Xu DH, Xu C et al (2016) Silencing of long non-coding RNA ANRIL inhibits the development of multidrug resistance in gastric cancer cells. Oncol Rep 36:263–270
Zhang EB, Kong R, Yin DD et al (2014) Long noncoding RNA ANRIL indicates a poor prognosis of gastric cancer and promotes tumor growth by epigenetically silencing of miR-99a/miR-449a. Oncotarget 5:2276–2292
Sun Y, Zheng ZP, Li H et al (2016) ANRIL is associated with the survival rate of patients with colorectal cancer, and affects cell migration and invasion in vitro. Mol Med Rep 14:1714–1720
Acknowledgements
This study was supported by the National Natural Science Foundation of China (No.81473039), and the Science & Technology Development Fund of Tian** Education Commission for Higher Education (No.20140141).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
We declare that all experiments that we have performed comply with the current laws of our country. All procedures performed in our study involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Rights and permissions
About this article
Cite this article
Cui, P., Zhao, Y., Chu, X. et al. SNP rs2071095 in LincRNA H19 is associated with breast cancer risk. Breast Cancer Res Treat 171, 161–171 (2018). https://doi.org/10.1007/s10549-018-4814-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-018-4814-y