Abstract
Both cultivated soybean and its wild relative Glycine soja exhibit strong photoperiodic sensitivity at different latitudes. Recent studies have demonstrated that the blue light-absorbing cryptochrome gene, CRY1a, is involved in the photoperiodic flowering of soybeans. However, no sequence variation was found in the cDNA among cultivars at different latitudinal clines. In the present study, we examined whether positive selection due to polymorphisms in the cryptochrome genes of G. soja occurs. Partial DNA sequences, mainly exons, of cryptochrome genes CRY1a-1d and CRY2a–2c were analyzed for 18 accessions in the Japanese archipelago. The neutral evolutionary pattern of the polymorphisms for all cryptochrome genes except for CRY1a was summarized using Tajima’s D test and low nucleotide diversity was shown for all genes. Although CRY1a did not show neutral evolution, balancing selection was recognized in the intron while not in the exon. No geographical pattern of polymorphisms was observed in the cryptochrome genes. These results reject the possibility of cryptochrome genes being involved in the photoperiodic flowering of wild soybeans along a latitudinal cline.



Similar content being viewed by others
References
Ahmad D, Cashmore AR (1993) HY4 gene of A. thaliana encodes a protein with characteristics of a blue-light photoreceptor. Nature 366:162–166
Balasubramanian S, Sureshkumar S, Agrawal M, Michael TP, Wessínger C, Maloof JN, Clark R, Warthmann N, Chory J, Weigel D (2006) The PHYTOCHROME C photoreceptor gene mediates natural variation in flowering and growth responses of Arabidopsis thaliana. Nat Genet 38:711–715
Chen M, Chory J, Fankhauser C (2004) Light signal transduction in higher plants. Annu Rev Genet 38:87–117
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Fukui J, Kaizuma N (1971) Morphological and ecological differentiation of various characters of the Japanese wild soybean, Glycine soja. J Faculty Agric Iwate Univ 10:195–208
García-Gil MR, Mikkonen M, Savolainen O (2003) Nucleotide diversity at two phytochrome loci along a latitudinal cline in Pinus sylvestris. Mol Ecol 12:1195–1206
Garner WW, Akkard HA (1920) Effect of the relative length of day and night and other factors of the environment on growth and reproduction in plants. J Agric Res 18:553–606
Gill N, Findley S, Walling JG, Hans C, Ma J, Doyle J, Stacey G, Jackson SA (2009) Molecular and chromosomal evidence for allopolyploidy in soybean. Plant Physiol 151:1167–1174
Guo H, Yang H, Mockler TC, Lin C (1998) Regulation of flowering time by Arabidopsis photoreceptors. Science 279:1360–1363
Halliday JK, Whitelam CG (2003) Changes in photoperiod or temperature alter the functional relationships between phytochromes and reveal roles for phyD and phyE. Plant Physiol 131:1913–1920
Heschel SM, Selby J, Butler C, Whitelam CG, Sharrock AR, Donohue K (2007) A new role for phytochromes in temperature-dependent germination. New Phytol 174:735–741
Hymowitz T (1970) On the domestication of soybean. Econ Bot 24:408–421
Hyten DL, Song Q, Zhu Y, Choi I, Nelson RL, Costa JM, Specht JE, Shoemaker RC, Cregan PB (2006) Impacts of genetic bottlenecks on soybean genome diversity. Proc Natl Acad Sci USA 103:16666–16671
Ikeda H, Fujii N, Setoguchi H (2009) Molecular evolution of phytochromes in Cardamine nipponica (Brassicaceae) suggests the involvement of PHYE in local adaptation. Genetics 182:603–614
Ikeda H, Fujii N, Setoguchi H (2011) Molecular evolution of cryptochrome genes and the evolutionary manner of photoreceptor genes in Cardamine nipponica (Brassicaceae). J Plant Res 124:85–92
Ingvarsson PK, Garcia MV, Hall VD, Luquez V, Jansson S (2006) Clinal variation in phyB2, a candidate gene for day-length-induced growth cessation and bud set, across a latitudinal gradient in European aspen (Populus tremula). Genetics 172:1845–1853
Ingvarsson PK, Garcia MV, Luquez V, Hall D, Jansson S (2008) Nucleotide polymorphisms and phenotypic associations within and around the phytochrome B2 locus in European aspen (Populus tremula, Salicaceae). Genetics 178:2217–2226
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Mathews S, McBreen K (2008) Phylogenetic relationships of B-related phytochromes in the Brassicaceae: redundancy and the persistence of phytochrome D. Mol Phylogenet Evol 49:411–423
Matsumura H, Kitajima H, Akada S, Abe J, Minaka N, Takahashi R (2009) Molecular cloning and linkage map** of cryptochrome multigene family in soybean. Plant Genome 2:271–281
Salvi S, Sponza G, Morgante M, Tomes D, Niu X, Fengler AK, Meeley R, Ananiev VE, Svitashev S, Bruggemann E, Li B, Hainey FC, Radovic S, Zaina G, Rafalski J-A, Tingey VS, Miao G, Phillips LR, Tuberosa R (2007) Conserved noncoding genomic sequences associated with a flowering-time quantitative trait locus in maize. Proc Natl Acad Sci USA 104:11376–11381
Setoguchi H, Ohba H (1995) Phylogenetic relationships in Crossostylis (Rhizophoraceae) inferred from restriction site variation of chloroplast DNA. J Plant Res 108:87–92
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
White GM, Hamblin MT, Kresovich S (2004) Molecular evolution of the phytochrome gene family in sorghum: changing rates of synonymous and replacement evolution. Mol Biol Evol 21:716–723
Zhang Q, Li H, Li R, Hu R, Fan C, Chen F, Wang Z, Liu X, Fu Y, Lin C (2008) Association of the circadian rhythmic expression of GmCRY1a with a latitudinal cline in photoperiodic flowering of soybean. Proc Natl Acad Sci USA 105:21028–21033
Acknowledgments
We thank Prof. A. Nagatani (Kyoto University) for helpful comments about photoreceptors, Dr. J. Abe (Hokkaido University) for providing the date of flowering of G. soja, and Drs. H. Ikeda (National Museum of Nature and Science) and Y. Mitsui (Tokyo University of Agriculture) for advice on technical aspects. We also thank Legume Base at the National BioResource Project (NBRP) at Miyazaki University, Japan, for providing the seed collections. This study was supported by a Grant-in-Aid for Scientific Research (#21370036) from the Japan Society for the Promotion of Science to H.S.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
10265_2011_470_MOESM1_ESM.eps
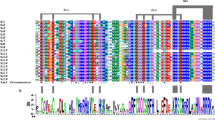
Fig. S1 Binding sites of PCR primers used to amplify the cryptochrome genes of Glycine soja. Closed and open triangles indicate forward and reverse primers, respectively. The type and number of primers are noted adjacent to the triangles (EPS 283 kb)
Rights and permissions
About this article
Cite this article
Ishibashi, N., Setoguchi, H. Polymorphism of DNA sequences of cryptochrome genes is not associated with the photoperiodic flowering of wild soybean along a latitudinal cline. J Plant Res 125, 483–488 (2012). https://doi.org/10.1007/s10265-011-0470-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10265-011-0470-6