Abstract
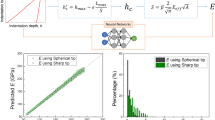
The Hertz contact mechanics model is commonly used to extract the elastic modulus of the cell, but the basic assumptions of the model are often not met in cell indentation experiments, which can lead to errors in the obtained elastic modulus of cell. The establishment of theoretical formulas or modification of the Hertz formulas has been proposed to reduce the errors introduced by indentation depth and cell thickness, but errors from cell radius and probe radius are largely neglected. Herein, we build a neural network model in machine learning to extract the elastic modulus of cell, which takes into account of four variables: indentation depth, cell thickness, cell radius, and probe radius. The validity of the model is demonstrated by the indentation experiment. The introduction of machine learning methods provides an alternative solution for extracting the elastic modulus of the cell and has potential for application.









Similar content being viewed by others
References
Butt HJ, Jaschke M (1995) Calculation of thermal noise in atomic force microscopy. Nanotechnology 6(1):1. https://doi.org/10.1088/0957-4484/6/1/001
Chen J, Lu G (2012) Finite element modelling of nanoindentation based methods for mechanical properties of cells. J Biomech 45(16):2810–2816. https://doi.org/10.1016/j.jbiomech.2012.08.037
Cross SE, ** YS, Rao J, Gimzewski JK (2007) Nanomechanical analysis of cells from cancer patients. Nat Nanotechnol 2(12):780–783. https://doi.org/10.1038/nnano.2007.388
Dimitriadis EK, Horkay F, Maresca J, Kachar B, Chadwick RC (2002) Determination of elastic moduli of thin layers of soft material using the atomic force microscope. Biophys J 82(5):2798–2810. https://doi.org/10.1016/S0006-3495(02)75620-8
Ding Y, Xu GK, Wang GF (2017) On the determination of elastic moduli of cells by AFM based indentation. Sci Rep-Uk 7(1):1–8. https://doi.org/10.1038/srep45575
Garcia PD, Ricardo G (2018) Determination of the elastic moduli of a single cell cultured on a rigid support by force microscopy. Biophys J 114(12):2923–2932. https://doi.org/10.1016/j.bpj.2018.05.012
Gavara N, Chadwick RS (2012) Determination of the elastic moduli of thin samples and adherent cells using conical atomic force microscope tips. Nat Nanotechnol 7(11):733–736. https://doi.org/10.1038/nnano.2012.163
Hertz H (1882) ber Die Berührung Fester Elastischer Krper. J Für Die Reine Angew Math (crelles Journal) 1882:156–171. https://doi.org/10.1515/crll.1882.92.156
Hornik K, Stinchcombe M, White H (1991) Approximation capabilities of multilayer feedforward networks. Neural Netw 4(2):251–257. https://doi.org/10.1016/0893-6080(91)90009-T
Kingma D, Ba J (2014) Adam: a method for stochastic optimization. J Comput Sci. Preprint http://arxiv.org/abs/1412.6980
Kiyohara S, Oda H, Miyata T, Mizoguchi T (2016) Prediction of interface structures and energies via virtual screening. Sci Adv 2(11):e1600746. https://doi.org/10.1126/sciadv.1600746
Lekka M, Fornal M, Pyka-Fościak G, Lebed K, Wizner B, Grodzicki T, Styczen J (2005) Erythrocyte stiffness probed using atomic force microscope. Biorheology 42(4):307–317. https://doi.org/10.1002/bip.20128
Li QS, Lee G, Ong CN, Lim CT (2008) AFM indentation study of breast cancer cells. Biochem Bioph Res Co 374(4):609–613. https://doi.org/10.1016/j.bbrc.2008.07.078
Liu X, Athanasiou CE, Padture NP, Sheldon BW, Gao H (2020) A machine learning approach to fracture mechanics problems. Acta Mater 190:105–112. https://doi.org/10.1016/j.actamat.2020.03.016
Long R, Hall MS, Wu M, Hui CY (2011) Effects of gel thickness on microscopic indentation measurements of gel modulus. Biophys J 101(3):643–650. https://doi.org/10.1016/j.bpj.2011.06.049
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, Killeen T, Lin Z, Gimelshein N, Antig L, Desmaison A, Köpf A, Yang E, DeVito Z, DeVito M, Tejani A, Chilamkurthy S, Steiner B, Fang L, Bai J, Chintala S (2019) PyTorch: an imperative style, high-performance deep learning library. N I P S 32:8026–8037. Preprint http://arxiv.org/abs/1912.01703
Rumelhart DE, Hinton GE, Williams RJ (1986) Learning representations by back propagating errors. Nature 323(6088):533–536. https://doi.org/10.1038/323533a0
Roan E, Wilhelm K, Bada A, Makena PS, Gorantla VK, Sinclair SE, Waters CM (2012) Hyperoxia alters the mechanical properties of alveolar epithelial cells. Am J Physiol-Lung C 302(12):L1235–L1241. https://doi.org/10.1152/ajplung.00223.2011
Sen S, Subramanian S, Discher DE (2005) Indentation and adhesive probing of a cell membrane with AFM: theoretical model and experiments. Biophys J 89(5):3203–3213. https://doi.org/10.1529/biophysj.105.063826
Sun W, Ma J, Wang C, Li H, Zhang W (2021a) Precise determination of elastic modulus of cell using conical AFM probe. J Biomech 118(60):110277. https://doi.org/10.1016/j.jbiomech.2021.110277
Sun W, Yin P, Wang C, Ren Y, Han X, Wu C, Zhang W (2021b) Determination of the elastic modulus of adherent cells using spherical atomic force microscope probe. J Mater Sci 56(32):18210–18218. https://doi.org/10.1007/s10853-021-06445-5
Titushkin I, Cho M (2007) Modulation of cellular mechanics during osteogenic differentiation of human mesenchymal stem cells. Biophys J 93(10):3693–3702. https://doi.org/10.1529/biophysj.107.107797
Titushkin I, Cho M (2009) Regulation of cell cytoskeleton and membrane mechanics by electric field: role of linker proteins. Biophys J 96(2):717–728. https://doi.org/10.1016/j.bpj.2008.09.035
Volle CB, Ferguson MA, Aidala KE, Spain EM, Núez ME (2008) Quantitative changes in the elasticity and adhesive properties of Escherichia coli ZK1056 prey cells during predation by Bdellovibrio bacteriovorus 109J. Langmuir 24(15):8102–8110. https://doi.org/10.1021/la8009354
Wang L, Tian L, Wang Y, Zhang W, Wang Z, Liu X (2020) Determination of viscohyperelastic properties of tubule epithelial cells by an approach combined with AFM nanoindentation and finite element analysis. Micron 129(2020):102779. https://doi.org/10.1016/j.micron.2019.102779
Wang Y, Riordon J, Kong T, Xu Y, Nguyen B, Zhong J, You JB, Lagunov A, Hannam TG, Jarvi K, Sinton D (2019) Prediction of DNA integrity from morphological parameters using a single-sperm DNA fragmentation index assay. Adv Sci 6(15):1900712. https://doi.org/10.1002/advs.201900712
Wildling L, Rankl C, Haselgrübler T, Gruber HJ, Holy M, Newman AH, Zou MF, Zhu R, Freissmuth M, Sitte HH, Hinterdorfer P (2012) Probing binding pocket of serotonin transporter by single molecular force spectroscopy on living cells. J Biol Chem 287(1):105–113. https://doi.org/10.1074/jbc.M111.304873
Xu B, Wang N, Chen T, Li M (2015) Empirical evaluation of rectified activations in convolutional network. J Comput Ence. Preprint http://arxiv.org/abs/1505.00853
Xu XS, Zhong WX, Zhang HW (1997) The Saint-Venant problem and principle in elasticity. Int J Solids Struct 34(22):2815–2827. https://doi.org/10.1016/S0020-7683(96)00198-9
Zhang CY, Zhang YW (2008) Computational analysis of adhesion force in the indentation of cells using atomic force microscopy. Phys Rev E 77(2):021912. https://doi.org/10.1103/PhysRevE.77.021912
Acknowledgements
This work was supported by National Key R&D Project of China (2018YFA0704103, 2018YFA0704104), and Fundamental Research Funds for the Central Universities (DUT22YG123, DUT21TD105, DUT21YG109).
Author information
Authors and Affiliations
Contributions
GZ was involved in conceptualization, formal analysis, investigation, methodology, visualization, writing—original draft, writing—review and editing. MC contributed to investigation, methodology. CW was involved in methodology, writing—original draft. XH and CW contributed to writing—review and editing. WZ was involved in conceptualization, funding acquisition, project administration, resources, supervision, validation, writing—original draft, writing—review and editing. It is hereby declared that all authors have read and agreed to the contents of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhou, G., Chen, M., Wang, C. et al. Machine learning method for extracting elastic modulus of cells. Biomech Model Mechanobiol 21, 1603–1612 (2022). https://doi.org/10.1007/s10237-022-01609-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10237-022-01609-x