Abstract
Background
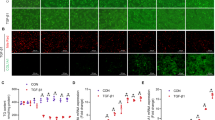
Multiple mechanisms contribute to the liver fibrosis following cholestasis. Recent research has focused on the role of transforming growth factor β-1 (TGF-β1) in the progression of fibrosis. The aim of our study is to examine the role of epigenetic chromatin marks, such as histone H3 lysine methylation (H3Kme), in bile duct ligation (BDL)-induced TGF-β1 gene expression in rat liver.
Methods
Time course of methylated-histone H3 and SET7/9 recruitment were determined by chromatin immunoprecipitation in livers from BDL rats on days 1, 4, 9 and 14. Levels of TGF-β1 and SET7/9 were determined by western blots. The effect of SET7/9 knockdown on BDL-induced expression of TGF-β1, serum enzymes and liver collagen content was studied in vivo.
Results
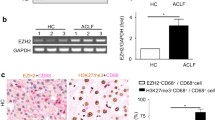
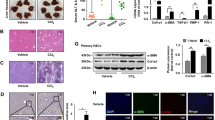
Results showed that BDL increased the expression of the TGF β-1. Increased levels of active chromatin marks (H3K4me1, H3K4me2, and H3K4me3) and decreased levels of repressive marks (H3K9me2 and H3K9me3) in TGF-β1 promoter accompanied the changes in expression of the TGF β-1. BDL also increased expression of the H3K4 methyltransferase SET7/9 and recruitment to the promoter. SET7/9 gene knockdown with siRNAs significantly attenuated BDL-induced TGF-β1 gene expression, serum enzymes and liver collagen content.
Conclusions
Taken together, these results show the functional role of epigenetic chromatin histone H3Kme in BDL-induced TGF-β1 expression. Pharmacologic and other therapies that reverse these modifications could have potential hepatoprotective effects for BDL-induced cirrhosis.








Similar content being viewed by others
Abbreviations
- α-SMA:
-
α-Smooth muscle actin
- BDL:
-
Bile duct ligation
- ChIP:
-
Chromatin immunoprecipitation
- ECM:
-
Extracellular matrix
- HATs:
-
Histone acetyl transferases
- HDACs:
-
Histone deacetylases
- HDMs:
-
Histone demethylases
- HSC:
-
Hepatic stellate cell
- H3KAc:
-
Acetylation of histone H3 lysines
- H3Kme:
-
Histone H3 lysine methylation
- qRT-PCR:
-
Quantitative reverse-transcription polymerase chain reaction
- TB:
-
Total bilirubin
- TGF-β1:
-
Transforming growth factor β-1
References
Scott-Conner CE, Grogan JB. The pathophysiology of biliary obstruction and its effect on phagocytic and immune function. J Surg Res. 1994;57(2):316–36 (Epub 1994/08/01).
Sheen-Chen SM, Ho HT, Chen WJ, Eng HL. Effect of ZVAD-fmk on hepatocyte apoptosis after bile duct ligation in rat. World J Gastroenterol WJG. 2005;11(15):2330–3 (Epub 2005/04/09).
Sheen-Chen SM, Ho HT, Chen WJ, Eng HL, Wu CH. Obstructive jaundice alters CD44 expression in rat small intestine. Digestion. 2002;65(2):112–7 (Epub 2002/05/22).
Tanaka A, Tsuneyama K, Mikami M, Uegaki S, Aiso M, Takikawa H. Gene expression profiling in whole liver of bile duct ligated rats: VEGF-A expression is up-regulated in hepatocytes adjacent to the portal tracts. J Gastroenterol Hepatol. 2007;22(11):1993–2000 (Epub 2007/10/05).
Denk GU, Cai SY, Chen WS, Lin A, Soroka CJ, Boyer JL. A comparison of gene expression in mouse liver and kidney in obstructive cholestasis utilizing high-density oligonucleotide microarray technology. World J Gastroenterol WJG. 2006;12(16):2536–48 (Epub 2006/05/12).
Takiya S, Tagaya T, Takahashi K, Kawashima H, Kamiya M, Fukuzawa Y, et al. Role of transforming growth factor beta 1 on hepatic regeneration and apoptosis in liver diseases. J Clin Pathol. 1995;48(12):1093–7 (Epub 1995/12/01).
Yamano T, Hirai R, Hato S, Uemura T, Shimizu N. Delayed liver regeneration with negative regulation of hepatocyte growth factor and positive regulation of transforming growth factor-beta1 mRNA after portal branch ligation in biliary obstructed rats. Surgery. 2002;131(2):163–71 (Epub 2002/02/21).
Khaw PT, Schultz GS, MacKay SL, Chegini N, Rotatori DS, Adams JL, et al. Detection of transforming growth factor-alpha messenger RNA and protein in human corneal epithelial cells. Invest Ophthalmol Vis Sci. 1992;33(12):3302–6 (Epub 1992/11/01).
Asano Y, Czuwara J, Trojanowska M. Transforming growth factor-beta regulates DNA binding activity of transcription factor Fli1 by p300/CREB-binding protein-associated factor-dependent acetylation. J Biol Chem. 2007;282(48):34672–83 (Epub 2007/09/22).
Massague J. The transforming growth factor-beta family. Annu Rev Cell Biol. 1990;6:597–641 (Epub 1990/01/01).
Zarnegar R, Michalopoulos G. Purification and biological characterization of human hepatopoietin A, a polypeptide growth factor for hepatocytes. Cancer Res. 1989;49(12):3314–20 (Epub 1989/06/15).
Tomita Y, Nakamura T, Ichihara A. Control of DNA synthesis and ornithine decarboxylase activity by hormones and amino acids in primary cultures of adult rat hepatocytes. Exp Cell Res. 1981;135(2):363–71 (Epub 1981/10/01).
Russell WE, Coffey RJ Jr, Ouellette AJ, Moses HL. Type beta transforming growth factor reversibly inhibits the early proliferative response to partial hepatectomy in the rat. Proc Natl Acad Sci USA. 1988;85(14):5126–30 (Epub 1988/07/01).
Friedman SL. Liver fibrosis—from bench to bedside. J Hepatol. 2003;38(Suppl 1):S38–53 (Epub 2003/02/20).
Wilson AS, Power BE, Molloy PL. DNA hypomethylation and human diseases. Biochim Biophys Acta. 2007;1775(1):138–62 (Epub 2006/10/19).
Jones PA, The DNA. methylation paradox. Trends Genet TIG. 1999;15(1):34–7 (Epub 1999/03/24).
Rice JC, Briggs SD, Ueberheide B, Barber CM, Shabanowitz J, Hunt DF, et al. Histone methyltransferases direct different degrees of methylation to define distinct chromatin domains. Mol Cell. 2003;12(6):1591–8 (Epub 2003/12/24).
Berger SL. Histone modifications in transcriptional regulation. Curr Opin Genet Dev. 2002;12(2):142–8 (Epub 2002/03/15).
Shilatifard A. Chromatin modifications by methylation and ubiquitination: implications in the regulation of gene expression. Annu Rev Biochem. 2006;75:243–69 (Epub 2006/06/08).
Jenuwein T, Allis CD. Translating the histone code. Science. 2001;293(5532):1074–80 (Epub 2001/08/11).
Reddy GK, Enwemeka CS. A simplified method for the analysis of hydroxyproline in biological tissues. Clin Biochem. 1996;29(3):225–9.
Lin CR, Yang CH, Huang CE, Wu CH, Chen YS, Sheen-Chen SM, et al. GADD45A protects against cell death in dorsal root ganglion neurons following peripheral nerve injury. J Neurosci Res. 2011;89(5):689–99 (Epub 2011/02/22).
Cha JY, Repa JJ. The liver X receptor (LXR) and hepatic lipogenesis. The carbohydrate-response element-binding protein is a target gene of LXR. J Biol Chem. 2007;282(1):743–51 (Epub 2006/11/17).
Friedman SL. Mechanisms of hepatic fibrogenesis. Gastroenterology. 2008;134(6):1655–69.
Dooley S, ten Dijke P. TGF-beta in progression of liver disease. Cell Tissue Res. 2012;347(1):245–56.
Wasmuth HE, Tacke F, Trautwein C. Chemokines in liver inflammation and fibrosis. Semin Liver Dis. 2010;30(3):215–25 (Epub 2010/07/29).
Gressner AM, Weiskirchen R, Breitkopf K, Dooley S. Roles of TGF-beta in hepatic fibrosis. Front Biosci J Virtual Libr. 2002;7:d793–807 (Epub 2002/03/19).
Ueberham E, Low R, Ueberham U, Schonig K, Bujard H, Gebhardt R. Conditional tetracycline-regulated expression of TGF-beta1 in liver of transgenic mice leads to reversible intermediary fibrosis. Hepatology. 2003;37(5):1067–78 (Epub 2003/04/30).
Schnur J, Olah J, Szepesi A, Nagy P, Thorgeirsson SS. Thioacetamide-induced hepatic fibrosis in transforming growth factor beta-1 transgenic mice. Eur J Gastroenterol Hepatol. 2004;16(2):127–33 (Epub 2004/04/13).
Oberhammer FA, Pavelka M, Sharma S, Tiefenbacher R, Purchio AF, Bursch W, et al. Induction of apoptosis in cultured hepatocytes and in regressing liver by transforming growth factor beta 1. Proc Natl Acad Sci USA. 1992;89(12):5408–12 (Epub 1992/06/15).
Nakamura T, Sakata R, Ueno T, Sata M, Ueno H. Inhibition of transforming growth factor beta prevents progression of liver fibrosis and enhances hepatocyte regeneration in dimethylnitrosamine-treated rats. Hepatology. 2000;32(2):247–55 (Epub 2000/08/01).
Lang Q, Liu Q, Xu N, Qian KL, Qi JH, Sun YC, et al. The antifibrotic effects of TGF-beta1 siRNA on hepatic fibrosis in rats. Biochem Biophys Res Commun. 2011;409(3):448–53 (Epub 2011/05/24).
Mann DA, Mann J. Epigenetic regulation of hepatic stellate cell activation. J Gastroenterol Hepatol. 2008;23(Suppl 1):S108–11 (Epub 2008/04/11).
Mann J, Oakley F, Akiboye F, Elsharkawy A, Thorne AW, Mann DA. Regulation of myofibroblast transdifferentiation by DNA methylation and MeCP2: implications for wound healing and fibrogenesis. Cell Death Differ. 2007;14(2):275–85 (Epub 2006/06/10).
Niki T, Rombouts K, De Bleser P, De Smet K, Rogiers V, Schuppan D, et al. A histone deacetylase inhibitor, trichostatin A, suppresses myofibroblastic differentiation of rat hepatic stellate cells in primary culture. Hepatology. 1999;29(3):858–67 (Epub 1999/03/03).
Rombouts K, Knittel T, Machesky L, Braet F, Wielant A, Hellemans K, et al. Actin filament formation, reorganization and migration are impaired in hepatic stellate cells under influence of trichostatin A, a histone deacetylase inhibitor. J Hepatol. 2002;37(6):788–96 (Epub 2002/11/26).
Mann J, Chu DC, Maxwell A, Oakley F, Zhu NL, Tsukamoto H, et al. MeCP2 controls an epigenetic pathway that promotes myofibroblast transdifferentiation and fibrosis. Gastroenterology. 2010;138(2):705–14, 14 e1–4 (Epub 2009/10/22).
Roderburg C, Urban GW, Bettermann K, Vucur M, Zimmermann H, Schmidt S, et al. Micro-RNA profiling reveals a role for miR-29 in human and murine liver fibrosis. Hepatology. 2011;53(1):209–18 (Epub 2010/10/05).
Rice JC, Allis CD. Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr Opin Cell Biol. 2001;13(3):263–73 (Epub 2001/05/10).
Martin C, Zhang Y. The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol. 2005;6(11):838–49 (Epub 2005/11/02).
Corvol P. VEGF, anti-vEGF and diseases. Bulletin de l’Academie Nationale de Medecine. 2008;192(2):289–300 (discussion -2. Epub 2008/09/30. Conference invitee VEGF, anti-vEGF et pathologies).
Hampsey M, Reinberg D. Tails of intrigue: phosphorylation of RNA polymerase II mediates histone methylation. Cell. 2003;113(4):429–32 (Epub 2003/05/22).
Wang H, Cao R, **a L, Erdjument-Bromage H, Borchers C, Tempst P, et al. Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol Cell. 2001;8(6):1207–17 (Epub 2002/01/10).
Nishioka K, Chuikov S, Sarma K, Erdjument-Bromage H, Allis CD, Tempst P, et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev. 2002;16(4):479–89 (Epub 2002/02/19).
Kouskouti A, Scheer E, Staub A, Tora L, Talianidis I. Gene-specific modulation of TAF10 function by SET9-mediated methylation. Mol Cell. 2004;14(2):175–82 (Epub 2004/04/22).
Berger SL. The complex language of chromatin regulation during transcription. Nature. 2007;447(7143):407–12 (Epub 2007/05/25).
Esteve PO, Chin HG, Benner J, Feehery GR, Samaranayake M, Horwitz GA, et al. Regulation of DNMT1 stability through SET7-mediated lysine methylation in mammalian cells. Proc Natl Acad Sci USA. 2009;106(13):5076–81 (Epub 2009/03/14).
Yang XD, Huang B, Li M, Lamb A, Kelleher NL, Chen LF. Negative regulation of NF-kappaB action by Set9-mediated lysine methylation of the RelA subunit. EMBO J. 2009;28(8):1055–66 (Epub 2009/03/06).
Li Y, Reddy MA, Miao F, Shanmugam N, Yee JK, Hawkins D, et al. Role of the histone H3 lysine 4 methyltransferase, SET7/9, in the regulation of NF-kappaB-dependent inflammatory genes. Relevance to diabetes and inflammation. J Biol Chem. 2008;283(39):26771–81 (Epub 2008/07/25).
Acknowledgments
This work was supported in part by grant No. 8A1011 and 880893 from Chang Gung Memorial Hospital Research, Kaohsiung, Taiwan, and by grant No. 101-2314-B-182-021, 100-2314-B-182-068, 101-2314-B-182-090, 101-2314-B-182A-014-, and 101-2314-B-182A-068-MY3 from the Taiwan National Science Council Research, Taipei, Taiwan.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sheen-Chen, SM., Lin, CR., Chen, KH. et al. Epigenetic histone methylation regulates transforming growth factor β-1 expression following bile duct ligation in rats. J Gastroenterol 49, 1285–1297 (2014). https://doi.org/10.1007/s00535-013-0892-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-013-0892-0