Summary
Background
Colorectal cancer (CRC) is among the most widespread malignancies in the world. MicroRNA (miRNA) has been identified as an important modulator of the biological processes of the cells. This group of noncoding RNAs also has a pivotal function in the growth and development of human cancers, including CRC. Among these miRNAs, miR-196, miR-132, miR-146a, and miR-134 have fundamental impacts on the regulation of cancers. The current study aimed to investigate the involvement of these miRNAs in CRC patients.
Methods
In this study, 50 pairs of tumor and tumor margin samples of CRC patients were investigated to assess the expression levels of miR-196, miR-132, miR-146a, and miR-134 in this cancer. For this purpose, firstly, quantitative real-time PCR (qRT-PCR) was applied. Also, KRAS mutation and clinicopathological characteristics of the CRC patients were analyzed in the study groups.
Results
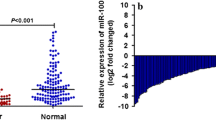
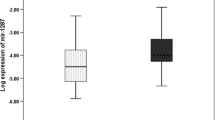
The findings demonstrated the overexpression of miR-196 (P-value = 0.0045) and miR-146a (P-value = 0.0033) in tumor tissues compared to controls. Conversely, the expression levels of miR-132 (P-value = 0.00032) and miR-134 (P-value < 0.0001) were downregulated in tumor tissues. Also, miR-146a was the only miRNA with significant expression change in the case of the KRAS gene mutation. Interestingly, the expression ratio of these miRNAs was significantly associated with some of the clinicopathological features of the patients, such as lymph node and distant metastases.
Conclusion
Our data demonstrated that these miRNAs appear to be promising novel biomarkers for early diagnosis of CRC and may pave the way for the future establishment of novel therapeutic options for CRC.





Similar content being viewed by others
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. Ca A Cancer J Clin. 2015;65:87–108.
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Piñeros M, et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144:1941–53.
Shirafkan N, Mansoori B, Mohammadi A, Shomali N, Ghasbi M, Baradaran B. MicroRNAs as novel biomarkers for colorectal cancer: new outlooks. Biomed Pharmacother. 2018;97:1319–30.
Esmailzadeh S, Mansoori B, Mohammadi A, Shanehbandi D, Baradaran B. siRNA-mediated silencing of HMGA2 induces apoptosis and cell cycle arrest in human colorectal carcinoma. J Gastrointest Cancer. 2017;48:156–63.
Corté H, Manceau G, Blons H, Laurent-Puig P. MicroRNA and colorectal cancer. Dig Liver Dis. 2012;44:195–200.
Gonzalez-Pons M, Cruz-Correa M. Colorectal cancer biomarkers: Where are we now? Biomed Res Int. 2015; https://doi.org/10.1155/2015/149014.
Wahid F, Shehzad A, Khan T, Kim YY. MicroRNAs: synthesis, mechanism, function, and recent clinical trials. Biochim Biophys Acta. 2010;1803:1231–43.
Ghasabi M, Mansoori B, Mohammadi A, Duijf PH, Shomali N, Shirafkan N, et al. MicroRNAs in cancer drug resistance: basic evidence and clinical applications. J Cell Physiol. 2019;234:2152–68.
Shenouda SK, Alahari SK. MicroRNA function in cancer: oncogene or a tumor suppressor? Cancer Metastasis Rev. 2009;28:369–78.
Mohammadi A, Mansoori B, Baradaran B. The role of microRNAs in colorectal cancer. Biomed Pharmacother. 2016;84:705–13.
Karami H, Baradaran B, Esfahani A, Estiar MA, Naghavi-Behzad M, Sakhinia M, et al. siRNA-mediated silencing of survivin inhibits proliferation and enhances etoposide chemosensitivity in acute myeloid leukemia cells. Asian Pac J Cancer Prev. 2013;14:7719–24.
Suzuki H, Takatsuka S, Akashi H, Yamamoto E, Nojima M, Maruyama R, et al. Genome-wide profiling of chromatin signatures reveals epigenetic regulation of microRNA genes in colorectal cancer. Cancer Res. 2011;71:5646–58.
Wang YX, Zhang XY, Zhang BF, Yang CQ, Chen XM, Gao HJ. Initial study of microRNA expression profiles of colonic cancer without lymph node metastasis. J Dig Dis. 2010;11:50–4.
Bin Zheng Y, Luo HP, Shi Q, Hao ZN, Ding Y, Wang QS, et al. miR-132 inhibits colorectal cancer invasion and metastasis via directly targeting ZEB2. World J Gastroenterol. 2014;20:6515–22.
Formosa A, Lena AM, Markert EK, Cortelli S, Miano R, Mauriello A, et al. DNA methylation silences miR-132 in prostate cancer. Oncogene. 2013;32:127–34.
Zhang S, Hao J, **e F, Hu X, Liu C, Tong J, et al. Downregulation of miR-132 by promoter methylation contributes to pancreatic cancer development. Carcinogenesis. 2011;32:1183–9.
Qin J, Ke J, Xu J, Wang F, Zhou Y, Jiang Y, et al. Downregulation of microRNA -132 by DNA hypermethylation is associated with cell invasion in colorectal cancer. Onco Targets Ther. 2015;8:3639–48.
Chae YS, Kim JG, Lee SJ, Kang BW, Lee YJ, Park JYJS, et al. A miR-146a polymorphism (rs2910164) predicts risk of and survival from colorectal cancer. Anticancer Res. 2013;33:3233–40.
Philippidou D, Schmitt M, Moser D, Margue C, Nazarov PV, Muller A, et al. Signatures of MicroRNAs and selected MicroRNA target genes in human melanoma. Cancer Res. 2010;70:4163–73.
Wang X, Tang S, Le SY, Lu R, Rader JS, Meyers C, et al. Aberrant expression of oncogenic and tumor-suppressive microRNAs in cervical cancer is required for cancer cell growth. Plos One. 2008;3(7):e2557.
Sun M, Fang S, Li W, Li C, Wang L, Wang F, et al. Associations of miR-146a and miR-146b expression and clinical characteristics in papillary thyroid carcinoma. Cancer Biomark. 2015;15:33–40.
Ye Q, Su L, Chen D, Zheng W, Liu Y. Astragaloside IV induced MIR-134 expression reduces EMT and increases chemotherapeutic sensitivity by suppressing CREB1 signaling in colorectal cancer cell line SW-480. Cell Physiol Biochem. 2017;43:1617–26.
Aghaee F, Pirayesh Islamian J, Baradaran B. Enhanced radiosensitivity and chemosensitivity of breast cancer cells by 2‑deoxy-D-glucose in combination therapy. J Breast Cancer. 2012;15:141.
Hajiasgharzadeh K, Tavangar SM, Javan M, Dehpour AR, Mani AR. Does hepatic vagus nerve modulate the progression of biliary fibrosis in rats? Auton Neurosci. 2014;185:67–75.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods. 2001;25:402–8.
Daryabari SS, Safaralizadeh R, Hosseinpourfeizi M, Moaddab Y, Shokouhi B. Overexpression of SSH1 in gastric adenocarcinoma and its correlation with clinicopathological features. J Gastrointest Oncol. 2018;9:728–33.
Madhavan D, Cuk K, Burwinkel B, Yang R. Cancer diagnosis and prognosis decoded by blood-based circulating micro RNA signatures. Front Genet. 2013;4:116.
Thome AD, Harms AS, Volpicelli-Daley LA, Standaert DG. MicroRNA-155 regulates alpha-synuclein-induced inflammatory responses in models of Parkinson disease. J Neurosci. 2016;36:2383–90.
Rayner KJ, Suarez Y, Davalos A, Parathath S, Fitzgerald ML, Tamehiro N, et al. MiR-33 contributes to the regulation of cholesterol homeostasis. Science. 2010;328(5985):1570–3.
Ahmed FE, Ahmed NC, Vos PW, Bonnerup C, Atkins JN, Casey M, et al. Diagnostic MicroRNA markers to screen for sporadic human colon cancer in stool: I. Proof of principle. Cancer Genomics Proteomics. 2013;10:93–113.
Iannone A, Losurdo G, Pricci M, Girardi B, Massaro A, Principi M, et al. Stool investigations for colorectal cancer screening: from occult blood test to DNA analysis. J Gastrointest Cancer. 2016;47:143–51.
Schimanski CC, Frerichs K, Rahman F, Berger M, Lang H, Galle PR, et al. High miR-196a levels promote the oncogenic phenotype of colorectal cancer cells. World J Gastroenterol. 2009;15:2089–96.
Ge J, Chen Z, Li R, Lu T, **ao G. Upregulation of microRNA-196a and microRNA-196b cooperatively correlate with aggressive progression and unfavorable prognosis in patients with colorectal cancer. Cancer Cell Int. 2014;14:128.
Chen CJ, Zhang Y, Zhang L, Weakley SM, Yao Q. MicroRNA-196: critical roles and clinical applications in development and cancer. J Cell Mol Med. 2011;15:14–23.
Shen S, Pan J, Lu X, Chi P. Role of miR-196 and its target gene HoxB8 in the development and proliferation of human colorectal cancer and the impact of neoadjuvant chemotherapy with FOLFOX4 on their expression. Oncol Lett. 2016;12:4041–7.
Geng F, Wu JL, Lu GF, Liang ZP, Duan ZL, Gu X. MicroRNA-132 targets PEA-15 and suppresses the progression of astrocytoma in vitro. J Neurooncol. 2016;129:211–20.
Zhang B, Lu L, Zhang X, Ye W, Wu J, ** Q, et al. Hsa-miR-132 regulates apoptosis in non-small cell lung cancer independent of acetylcholinesterase. J Mol Neurosci. 2014;53:335–44.
Alzahrani AM, Hanieh H, Ibrahim H‑IM, Mohafez O, Shehata T, Bani Ismail M, et al. Enhancing miR-132 expression by aryl hydrocarbon receptor attenuates tumorigenesis associated with chronic colitis. Int Immunopharmacol. 2017;52:342–51.
Xu G, Zhu H, Zhang M, Xu J. Histone deacetylase 3 is associated with gastric cancer cell growth via the miR-454-mediated targeting of CHD5. Int J Mol Med. 2018;41:155–63.
Wu J, He D, Yue B, Zhang C, Fang X, Chen H. miR-101‑1 expression pattern in Qinchuan cattle and its role in the regulation of cell differentiation. Gene. 2017;636:64–9.
Chen G, Umelo IA, Lv S, Teugels E, Fostier K, Kronenberger P, et al. miR-146a inhibits cell growth, cell migration and induces apoptosis in non-small cell lung cancer cells. PLoS One. 2013;8:e60317.
Yao Q, Cao Z, Tu C, Zhao Y, Liu H, Zhang S. MicroRNA-146a acts as a metastasis suppressor in gastric cancer by targeting WASF2. Cancer Lett. 2013;335:219–24.
Pizzini S, Bisognin A, Mandruzzato S, Biasiolo M, Facciolli A, Perilli L, et al. Impact of microRNAs on regulatory networks and pathways in human colorectal carcinogenesis and development of metastasis. BMC Genomics. 2013;14:589.
Mao Y, Li Y, **g F, Cai S, Zhang Z, Li Q, et al. Association of a genetic variant in microRNA-146a with risk of colorectal cancer: a population-based case-control study. Tumor Biol. 2014;35:6961–7.
Wang C, Guan S, Liu F, Chen X, Han L, Wang D, et al. Prognostic and diagnostic potential of miR-146a in oesophageal squamous cell carcinoma. Br J Cancer. 2016;114:290–7.
Niu CS, Yang Y, Cheng CD. MiR-134 regulates the proliferation and invasion of glioblastoma cells by reducing Nanog expression. Int J Oncol. 2013;42:1533–40.
Leivonen SK, Sahlberg KK, Mäkelä R, Due EU, Kallioniemi O, Børresen-Dale AL, et al. High-throughput screens identify microRNAs essential for HER2 positive breast cancer cell growth. Mol Oncol. 2014;8:93–104.
Liu CJ, Shen WG, Peng SY, Cheng HW, Kao SY, Lin SC, et al. MiR-134 induces oncogenicity and metastasis in head and neck carcinoma through targeting WWOX gene. Int J Cancer. 2014;134:811–21.
Acknowledgements
The authors would like to thank the Immunology Research Center, Tabriz University of Medical Sciences for providing facilities to carry out this work.
Author information
Authors and Affiliations
Contributions
MM, DS, MA, SH, and HMA performed the experiments. BB provided biological materials and reagents. MM and KH wrote the initial draft of the article. MM, DS, MA, and BB performed data analysis. BB and MP reviewed and edited the article. BB supervised the study.
Corresponding author
Ethics declarations
Conflict of interest
M. Maralani, D. Shanehbandi, M. Asadi, S. Hashemzadeh, K. Hajiasgharzadeh, H. Mashhadi Abdolahi, B. Baradaran and M. Peeters declare that they have no competing interests.
Ethical standards
All procedures performed in studies involving human participants or on human tissue were in accordance with the ethical standards of the institutional and/or national research committee (Ethics committee of Tabriz University of Medical Sciences, Tabriz, Iran) and with the 1975 Helsinki declaration and its later amendments or comparable ethical standards. Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Rights and permissions
About this article
Cite this article
Maralani, M., Shanehbandi, D., Asadi, M. et al. Expression profiles of miR-196, miR-132, miR-146a, and miR-134 in human colorectal cancer tissues in accordance with their clinical significance. Wien Klin Wochenschr 133, 1162–1170 (2021). https://doi.org/10.1007/s00508-021-01933-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00508-021-01933-9