Abstract
Purpose
Recent advances in sequencing technologies have enabled radical and rapid progress in the genetic diagnosis of inherited retinal disorders (IRDs). Although the list of gene variations continues to grow, it lacks the genetic etiology of ethnic groups like South Asians. Differences in racial backgrounds and consanguinity add to genetic heterogeneity and phenotypic overlaps.
Methods
This retrospective study includes documented data from the Gen-Eye clinic from years 2014 to 2019. Medical records and pedigrees of 591 IRD patients of Indian origin and genetic reports of 117 probands were reviewed. Genotype–phenotype correlations were performed to classify as correlating, non-correlating and unsolved cases.
Results
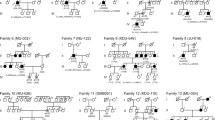
Among the 591 patients, we observed a higher prevalence of clinically diagnosed retinitis pigmentosa (38.9%) followed by unspecified diagnoses (28.5%). Consanguinity was reported to be high (55.6%) in this cohort. Among the variants identified in 117 probands, 36.4% of variants were pathogenic, 19.2% were likely pathogenic, and 44.4% were of uncertain significance. Among the pathogenic and likely pathogenic variants, autosomal recessive inheritance showed higher prevalence. About 35% (41/117) of cases showed genotype–phenotype correlation. Within the correlating cases, retinitis pigmentosa and Stargardt disease were predominant. Novel variants identified in RP, Stargardt, and LCA are reported here.
Conclusion
This first-of-a-kind report on an Indian cohort contributes to existing knowledge and expansion of variant databases, presenting relevant and plausible novel variants. Phenotypic overlap and variability lead to a differential diagnosis and hence a clear genotype–phenotype correlation helps in precise clinical confirmation. The study also emphasizes the importance of genetic counselling and testing for personalized vision care in a tertiary eye hospital.








Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article [and its supplementary information files].
References
Hanany M, Rivolta C, Sharon D (2020) Worldwide carrier frequency and genetic prevalence of autosomal recessive inherited retinal diseases. J Proc Natl Acad Sci 117:2710–2716. https://doi.org/10.1073/pnas.1913179117%
Tatour Y, Ben-Yosef T (2020) Syndromic Inherited Retinal Diseases: Genetic, Clinical and Diagnostic Aspects. Diagnostics (Basel, Switzerland) 10. https://doi.org/10.3390/diagnostics10100779
Talib M, Boon CJF (2020) Retinal Dystrophies and the Road to Treatment: Clinical Requirements and Considerations. Asia Pac J Ophthalmol (Phila) 9:159–179. https://doi.org/10.1097/apo.0000000000000290
Berger W, Kloeckener-Gruissem B, Neidhardt J (2010) The molecular basis of human retinal and vitreoretinal diseases. Prog Retin Eye Res 29:335–375. https://doi.org/10.1016/j.preteyeres.2010.03.004
Ferrari S, Di Iorio E, Barbaro V, Ponzin D, Sorrentino FS, Parmeggiani F (2011) Retinitis pigmentosa: genes and disease mechanisms. Curr Genomics 12:238–249. https://doi.org/10.2174/138920211795860107
Huang L, Zhang Q, Huang X, Qu C, Ma S, Mao Y, Yang J, Li Y, Li Y, Tan C, Zhao P, Yang Z (2017) Mutation screening in genes known to be responsible for Retinitis Pigmentosa in 98 Small Han Chinese Families. Sci Rep 7:1948. https://doi.org/10.1038/s41598-017-00963-6
Sen P, Bhargava A, George R, Ve Ramesh S, Hemamalini A, Prema R, Kumaramanickavel G, Vijaya L (2008) Prevalence of retinitis pigmentosa in South Indian population aged above 40 years. Ophthalmic Epidemiol 15:279–281. https://doi.org/10.1080/09286580802105814
Boughman JA, Conneally PM, Nance WE (1980) Population genetic studies of retinitis pigmentosa. Am J Hum Genet 32:223–235
Kumaran N, Moore AT, Weleber RG, Michaelides M (2017) Leber congenital amaurosis/early-onset severe retinal dystrophy: clinical features, molecular genetics and therapeutic interventions. Br J Ophthalmol 101:1147–1154. https://doi.org/10.1136/bjophthalmol-2016-309975
Mata NL, Weng J, Travis GH (2000) Biosynthesis of a major lipofuscin fluorophore in mice and humans with ABCR-mediated retinal and macular degeneration. Proc Natl Acad Sci USA 97:7154–7159. https://doi.org/10.1073/pnas.130110497
Chiang JP, Lamey T, McLaren T, Thompson JA, Montgomery H, De Roach J (2015) Progress and prospects of next-generation sequencing testing for inherited retinal dystrophy. Expert Rev Mol Diagn 15:1269–1275. https://doi.org/10.1586/14737159.2015.1081057
Consugar MB, Navarro-Gomez D, Place EM, Bujakowska KM, Sousa ME, Fonseca-Kelly ZD, Taub DG, Janessian M, Wang DY, Au ED, Sims KB, Sweetser DA, Fulton AB, Liu Q, Wiggs JL, Gai X, Pierce EA (2015) Panel-based genetic diagnostic testing for inherited eye diseases is highly accurate and reproducible, and more sensitive for variant detection, than exome sequencing. Genet Med 17:253–261. https://doi.org/10.1038/gim.2014.172
González-del Pozo M, Méndez-Vidal C, Bravo-Gil N, Vela-Boza A, Dopazo J, Borrego S, Antiñolo G (2014) Exome sequencing reveals novel and recurrent mutations with clinical significance in inherited retinal dystrophies. PLoS ONE 9:e116176. https://doi.org/10.1371/journal.pone.0116176
Moore AT (2017) Genetic Testing for Inherited Retinal Disease. Ophthalmology 124:1254–1255. https://doi.org/10.1016/j.ophtha.2017.06.018
Green RC, Berg JS, Grody WW, Kalia SS, Korf BR, Martin CL, McGuire AL, Nussbaum RL, O’Daniel JM, Ormond KE, Rehm HL, Watson MS, Williams MS, Biesecker LG (2013) ACMG recommendations for reporting of incidental findings in clinical exome and genome sequencing. Genet Med 15:565–574. https://doi.org/10.1038/gim.2013.73
Thompson DA, Iannaccone A, Ali RR, Arshavsky VY, Audo I, Bainbridge JWB, Besirli CG, Birch DG, Branham KE, Cideciyan AV, Daiger SP, Dalkara D, Duncan JL, Fahim AT, Flannery JG, Gattegna R, Heckenlively JR, Heon E, Jayasundera KT, Khan NW, Klassen H, Leroy BP, Molday RS, Musch DC, Pennesi ME, Petersen-Jones SM, Pierce EA, Rao RC, Reh TA, Sahel JA, Sharon D, Sieving PA, Strettoi E, Yang P, Zacks DN, Consortium TM (2020) Advancing Clinical Trials for Inherited Retinal Diseases: Recommendations from the Second Monaciano Symposium. J Translational Vision Science & Technology 9:2–2. https://doi.org/10.1167/tvst.9.7.2%
Russell S, Bennett J, Wellman JA, Chung DC, Yu ZF, Tillman A, Wittes J, Pappas J, Elci O, McCague S, Cross D, Marshall KA, Walshire J, Kehoe TL, Reichert H, Davis M, Raffini L, George LA, Hudson FP, Dingfield L, Zhu X, Haller JA, Sohn EH, Mahajan VB, Pfeifer W, Weckmann M, Johnson C, Gewaily D, Drack A, Stone E, Wachtel K, Simonelli F, Leroy BP, Wright JF, High KA, Maguire AM (2017) Efficacy and safety of voretigene neparvovec (AAV2-hRPE65v2) in patients with RPE65-mediated inherited retinal dystrophy: a randomised, controlled, open-label, phase 3 trial. Lancet (London, England) 390:849–860. https://doi.org/10.1016/s0140-6736(17)31868-8
Yohe S, Sivasankar M, Ghosh A, Ghosh A, Holle J, Murugan S, Gupta R, Schimmenti LA, Vedam R, Thyagarajan B (2020) Prevalence of mutations in inherited retinal diseases: A comparison between the United States and India. Mol Genet Genomic Med 8:e1081. https://doi.org/10.1002/mgg3.1081
Landrum MJ, Lee JM, Benson M, Brown GR, Chao C, Chitipiralla S, Gu B, Hart J, Hoffman D, Jang W, Karapetyan K, Katz K, Liu C, Maddipatla Z, Malheiro A, McDaniel K, Ovetsky M, Riley G, Zhou G, Holmes JB, Kattman BL, Maglott DR (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46:D1062-d1067. https://doi.org/10.1093/nar/gkx1153
Vaser R, Adusumalli S, Leng SN, Sikic M, Ng PC (2016) SIFT missense predictions for genomes. Nat Protoc 11:1–9. https://doi.org/10.1038/nprot.2015.123
Schwarz JM, Cooper DN, Schuelke M, Seelow D (2014) MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 11:361–362. https://doi.org/10.1038/nmeth.2890
García Bohórquez B, Aller E, Rodríguez Muñoz A, Jaijo T, García García G, Millán JM (2021) Updating the Genetic Landscape of Inherited Retinal Dystrophies. Front Cell Dev Biol 9:645600. https://doi.org/10.3389/fcell.2021.645600
Prokisch H, Hartig M, Hellinger R, Meitinger T, Rosenberg T (2007) A Population-Based Epidemiological and Genetic Study of X-Linked Retinitis Pigmentosa. J Invest Ophthalmol Vis Sci 48:4012–4018. https://doi.org/10.1167/iovs.07-0071%
Kikawa E, Nakazawa M, Chida Y, Shiono T, Tamai M (1994) A novel mutation (Asn244Lys) in the peripherin/RDS gene causing autosomal dominant retinitis pigmentosa associated with bull’s-eye maculopathy detected by nonradioisotopic SSCP. Genomics 20:137–139. https://doi.org/10.1006/geno.1994.1142
Wang H, den Hollander AI, Moayedi Y, Abulimiti A, Li Y, Collin RWJ, Hoyng CB, Lopez I, Abboud EB, Al-Rajhi AA, Bray M, Lewis RA, Lupski JR, Mardon G, Koenekoop RK, Chen R (2009) Mutations in SPATA7 cause Leber congenital amaurosis and juvenile retinitis pigmentosa. Am J Hum Genet 84:380–387. https://doi.org/10.1016/j.ajhg.2009.02.005
Foxman SG, Heckenlively JR, Bateman JB, Wirtschafter JD (1985) Classification of congenital and early onset retinitis pigmentosa. Arch Ophthalmol (Chicago, Ill: 1960) 103:1502–1506. https://doi.org/10.1001/archopht.1985.01050100078023
Serra R, Floris M, Pinna A, Boscia F, Cucca F, Angius A (2019) Novel mutations in c2orf71 causing an early onset form of cone-rod dystrophy: A molecular diagnosis after 20 years of clinical follow-up. Mol Vis 25:814–820
Birtel J, Eisenberger T, Gliem M, Müller PL, Herrmann P, Betz C, Zahnleiter D, Neuhaus C, Lenzner S, Holz FG, Mangold E, Bolz HJ, Charbel Issa P (2018) Clinical and genetic characteristics of 251 consecutive patients with macular and cone/cone-rod dystrophy. Sci Rep 8:4824. https://doi.org/10.1038/s41598-018-22096-0
Sohocki MM, Bowne SJ, Sullivan LS, Blackshaw S, Cepko CL, Payne AM, Bhattacharya SS, Khaliq S, Qasim Mehdi S, Birch DG, Harrison WR, Elder FF, Heckenlively JR, Daiger SP (2000) Mutations in a new photoreceptor-pineal gene on 17p cause Leber congenital amaurosis. Nat Genet 24:79–83. https://doi.org/10.1038/71732
Michaelides M, Chen LL, Brantley MA Jr, Andorf JL, Isaak EM, Jenkins SA, Holder GE, Bird AC, Stone EM, Webster AR (2007) ABCA4 mutations and discordant ABCA4 alleles in patients and siblings with bull’s-eye maculopathy. Br J Ophthalmol 91:1650–1655. https://doi.org/10.1136/bjo.2007.118356
Sutherland JE, Day MA (2011) Advantages and disadvantages of molecular testing in ophthalmology. Expert Rev Ophthalmol 6:221–245. https://doi.org/10.1586/eop.11.2
Wolock CJ, Stong N, Ma CJ, Nagasaki T, Lee W, Tsang SH, Kamalakaran S, Goldstein DB, Allikmets R (2019) A case-control collapsing analysis identifies retinal dystrophy genes associated with ophthalmic disease in patients with no pathogenic ABCA4 variants. Genet Med 21:2336–2344. https://doi.org/10.1038/s41436-019-0495-0
Wall JD, Stawiski EW, Ratan A, Kim HL, Kim C, Gupta R, Suryamohan K, Gusareva ES, Purbojati RW, Bhangale T, Stepanov V, Kharkov V, Schröder MS, Ramprasad V, Tom J, Durinck S, Bei Q, Li J, Guillory J, Phalke S, Basu A, Stinson J, Nair S, Malaichamy S, Biswas NK, Chambers JC, Cheng KC, George JT, Khor SS, Kim J-I, Cho B, Menon R, Sattibabu T, Bassi A, Deshmukh M, Verma A, Gopalan V, Shin J-Y, Pratapneni M, Santhosh S, Tokunaga K, Md-Zain BM, Chan KG, Parani M, Natarajan P, Hauser M, Allingham RR, Santiago-Turla C, Ghosh A, Gadde SGK, Fuchsberger C, Forer L, Schoenherr S, Sudoyo H, Lansing JS, Friedlaender J, Koki G, Cox MP, Hammer M, Karafet T, Ang KC, Mehdi SQ, Radha V, Mohan V, Majumder PP, Seshagiri S, Seo J-S, Schuster SC, Peterson AS, GenomeAsia KC (2019) The GenomeAsia 100K Project enables genetic discoveries across Asia. Nature 576:106–111. https://doi.org/10.1038/s41586-019-1793-z
Joseph B, Srinivasan A, Soumittra N, Vidhya A, Shetty NS, Uthra S, Kumaramanickavel G (2002) RPE65 gene: multiplex PCR and mutation screening in patients from India with retinal degenerative diseases. J Genet 81:19–23. https://doi.org/10.1007/bf02715866
Sundaresan P, Vijayalakshmi P, Thompson S, Ko AC, Fingert JH, Stone EM (2009) Mutations that are a common cause of Leber congenital amaurosis in northern America are rare in southern India. Mol Vis 15:1781–1787
Roca I, González-Castro L, Maynou J, Palacios L, Fernández H, Couce ML, Fernández-Marmiesse A (2020) PattRec: An easy-to-use CNV detection tool optimized for targeted NGS assays with diagnostic purposes. Genomics 112:1245–1256. https://doi.org/10.1016/j.ygeno.2019.07.011
Lalonde E, Rentas S, Lin F, Dulik MC, Skraban CM, Spinner NB (2020) Genomic Diagnosis for Pediatric Disorders: Revolution and Evolution. Front Pediatr 8:373–373. https://doi.org/10.3389/fped.2020.00373
Michie S, Marteau TM, Bobrow M (1997) Genetic counselling: the psychological impact of meeting patients’ expectations. J Med Genet 34:237–241. https://doi.org/10.1136/jmg.34.3.237
Murray RF Jr (1976) Psychosocial aspects of genetic counseling. Soc Work Health Care 2:13–23. https://doi.org/10.1300/J010v02n01_03
Huang JT, Heckenlively JR, Jayasundera KT, Branham KE (2014) The ophthalmic experience: unanticipated primary findings in the era of next generation sequencing. J Genet Couns 23:588–593. https://doi.org/10.1007/s10897-013-9679-y
Acknowledgements
Patients and their families visiting the Gen-Eye clinic and Narayana Nethralaya. Gen-Eye clinic coordinators—Naina Kumari and Reshma Varamballi for their efforts in patient management. Dr. Swaminathan Sethu and Dr. Arkasubhra Ghosh for their critical review and support. The Medgenome team for providing commercial next generation sequencing services and Anbu Kayalvizhi for variant data analysis.
Author information
Authors and Affiliations
Contributions
AG conceptualized the study design, CG and RR are co-first authors, CG, GPM, and AG wrote the manuscript, RR prepared images and tables, RP maintained patient records. AV, PB, TBM, and SB referred patients to the Gen-Eye Clinic based on clinical diagnosis for clinical and genetic correlations. We thank KBS for providing facilities and GKM for the genetics expertise.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Ethical clearance (EC Ref No: C/2021/08/03) for this retrospective study has been obtained by the Narayana Nethralaya Ethics Review Board.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Chitra Gopinath and Ramya Rompicherla are co-first authors
Supplementary Information

Supplementary Figure 1.
Decision of IRD diagnosis and patient inclusion criteria in the study. (PNG 865 kb)
Supplementary table 1.
Patient details, clinical diagnoses, genetic test results, clinical and genetic correlation statuses of 117 patients who underwent genetic testing. (XLSX 37 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gopinath, C., Rompicherla, R., Mathias, G.P. et al. Inherited retinal disorders: a genotype–phenotype correlation in an Indian cohort and the importance of genetic testing and genetic counselling. Graefes Arch Clin Exp Ophthalmol 261, 2003–2017 (2023). https://doi.org/10.1007/s00417-022-05955-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00417-022-05955-5