Abstract
Komagataella phaffii, a nonconventional yeast, is increasingly attractive to researchers owing to its posttranslational modification ability, strict methanol regulatory mechanism, and lack of Crabtree effect. Although CRISPR-based gene editing systems have been established in K. phaffii, there are still some inadequacies compared to the model organism Saccharomyces cerevisiae. In this study, a redesigned gRNA plasmid carrying red and green fluorescent proteins facilitated plasmid construction and marker recycling, respectively, making marker recycling more convenient and reliable. Subsequently, based on the knockdown of Ku70 and DNA ligase IV, we experimented with integrating multiple DNA fragments at a single locus. A 26.5-kb-long DNA fragment divided into 11 expression cassettes for lycopene synthesis could be successfully integrated into a single locus at one time with a success rate of 57%. A 27-kb-long DNA fragment could also be precisely knocked out with a 50% positive rate in K. phaffii by introducing two DSBs simultaneously. Finally, to explore the feasibility of rapidly balancing the expression intensity of multiple genes in a metabolic pathway, a yeast combinatorial library was successfully constructed in K. phaffii using lycopene as an indicator, and an optimal combination of the metabolic pathway was identified by screening, with a yield titer of up to 182.73 mg/L in shake flask fermentation.
Key points
• Rapid marker recycling based on the visualization of a green fluorescent protein
• One-step multifragment integration and large fragment knockout in the genome
• A random assembly of multiple DNA elements to create yeast libraries in K. phaffii
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With the advantages of mild reaction conditions and high reaction specificity, a vast number of microbial cell factories have been constructed in recent years to synthesize a variety of bulk chemicals through synthetic biology and metabolic engineering. Yeast, as a typical eukaryotic microorganism, plays an important role in the microbial cell factory; for example, Liu et al. (2020) achieved the compartmentalized synthesis of squalene in the peroxisome organelle of Saccharomyces cerevisiae. Among various yeasts, K. phaffii is a well-established GRAS (generally recognized as safe) eukaryotic host that is typically used for the expression and production of heterologous proteins due to its excellent posttranslational modification ability and tight methanol regulation mechanism (Ahmad et al. 2014; Yang and Zhang 2018). In contrast to S. cerevisiae, K. phaffii is a methylotrophic Crabtree-negative yeast and accordingly produces almost no ethanol as a byproduct during glucose fermentation. Additionally, K. phaffii could achieve high-density fermentation in basic media with methanol as the only carbon source to maintain cell growth and synthesize products. And the peroxisome organelles of K. phaffii could proliferate and expand under methanol-induced conditions (Bernauer et al. 2021; Ohsawa et al. 2022). Therefore, an increasing number of researchers have focused on yeast engineering to construct cellular factories of K. phaffii (Guo et al. S5).
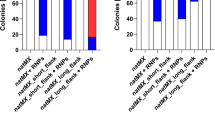
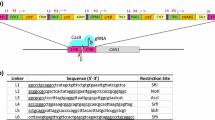
Design and validation of a yeast combinatorial library. a Schematic illustration of a yeast combinatorial library. The library consists of three promoters, four genes, and four terminators. The scissor means the cleavage site of CRISPR/Cas9. UHA and DHA are the upstream and downstream homologous arms of ADH900 loci, respectively, and their lengths were both approximately 1000 bp. b The screening process for yeast libraries. c Fermentation results of multiple clones picked from yeast libraries. Results of fermentation show that the maximum yield was 25 times higher than the minimum yield. All samples were run in triplicate
Next, 288 clones were transferred into 96 deep-well plates and cultured for 2–3 days (Fig. 6b). Then, 14 clones with different color grades were picked for sequencing and fermentation, with zw107 as a control. As expected, the results of both sequencing and HPLC analysis demonstrated that different combinations were generated in this yeast library and exhibited various phenotypes, and the lycopene yields varied from 7.28 to 182.73 mg/L (Fig. 6c). The maximum production was 25 times higher than the minimum. The sequencing results showed that strain TX-L3, in which crtI, crtYBW61R, tHMG1, and crtE were controlled by PGAP, PTEF1, PPGI1, and PTEF1, respectively, achieved maximum yield. The strain TX-L7, in which crtI, crtYBW61R, tHMG1, and crtE were controlled by PPGI1, PGAP, PPGI1, and PGAP, respectively, achieved minimum production. Moreover, we found that in higher-yielding strains, crtE (encoding GGPP synthases) and crtI (encoding phytoene desaturase), which catalyze the synthesis of GGPP from FPP and lycopene from phytoene, respectively, were under the control of the stronger promoter PGAP or PTEF1, suggesting that GGPP synthases and phytoene desaturase are two of the key enzymes in lycopene synthesis, which is consistent with previous reports (Zhang et al. 2020). Thus, it is practical and rapid to coordinate the expression levels of individual genes in the metabolic pathway to achieve a better combination.
Discussion
The methylotrophic yeast, K. phaffii, has been increasingly engineered into cell factories for the synthesis of various secondary metabolites in recent years (Cai et al. 2022; Guo et al. 2021; Zuo et al. 2022). To enable the construction of cell factories, several CRISPR-based gene editing systems have been established in K. phaffii (Cai et al. 2021; Liu et al. 2019; Nishi et al. 2022; Weninger et al. 2015, 2016; Zhang et al. 2021); however, compared to S. cerevisiae, K. phaffii still has potential for improvement in marker recycling, multifragment integration, etc. In this research, Kp6 (CBS7435ΔKu70) and Kp9 (CBS7435ΔKu70ΔDNL4) were constructed, and the KpHis4 locus was selected as the target to evaluate the homologous recombination efficiency of the two strains. The result showed better homologous recombination efficiency in Kp9, which is consistent with previously reported research results (Ito et al. 2018; Nishi et al. 2022). In addition, our previous experiments showed that the growth of the K. phaffii ΔURA3 strain was impaired, which is consistent with previously reported research results (Lin Cereghino et al. 2001); therefore, rapid marker recycling could not be performed similarly to that in S. cerevisiae on screening plates supplemented with 5-fluoroorotic acid (Kotaka et al. 2009; Moon et al. 2022). To simplify the marker recycling process, we redesigned the gRNA plasmid to determine whether plasmid elimination had been achieved through the visualization of green fluorescent protein. Compared to plasmid elimination methods in other systems (Liao et al. 2021; Liu et al. 2019; Weninger et al. 2018; Yang et al. 2020), the method we designed takes less time and may be more conducive to iterative gene editing. In addition, the GFP in the plasmid could assist in screening positive transformants.
When constructing cell factories, the number of genes integrated in one step is one of the factors that affect the efficiency of the construction. Although Nishi et al. (2022) realized the integration of four heterologous gene expression cassettes totaling 7.8 kb, which were divided into 10 fragments, in a single transformation in K. phaffii, there is still a gap compared to S. cerevisiae in which up to 15 fragments can be integrated in a one-step transformation at a single locus (Jakociunas et al. 2015). In this study, the one-step integration of 11 expression cassettes for a complete lycopene synthesis pathway with a total length of 26.5 kb at a single locus was achieved in K. phaffii for the first time. To the best of our knowledge, both the number of fragments and the total length of the fragments are by far the highest integrated in K. phaffii to date. Previously, NHEJ-based knockout of large fragments was performed by introducing DSBs via CRISPR/Cas9 in K. phaffii, and a positive rate of ~ 40% was achieved for a 4000-bp DNA fragment, but only ~ 2% of transformants were cleanly eliminated (Schusterbauer et al. 2022). Additionally, although Schusterbauer et al. (2022) achieved a “nonclean knockout” of a 100-kb DNA fragment in K. phaffii, the transformants lost the complete 3′ end of a chromosome rather than showing precise knockout of a large fragment inside a chromosome, as in our work. In addition to CRISPR/Cas9-based large fragment knockout, Zhang et al. (2021) realized CRISPR/Cpf1-based large fragment knockout with 11% efficiency for 20 kb. In this study, we first precisely knocked out a 27-kb DNA fragment inside a chromosome by introducing two DSBs at both ends of the fragment to be knocked out, thereby completely eliminating the lycopene synthesis pathway with 50% efficiency, which represented superior efficiency and length to those of the knockout achieved in the CRISPR/Cpf1 system (Zhang et al. 2021). Subsequently, three neutral targets, II-4, II-5, and II-6, were selected for testing larger DNA fragment knockouts in this study. However, in several repetitions of the experiment, no transformants successfully grew. The results of sequence alignment of proteins in the NCBI database indicated that the genes encoding the ribosomal biosynthesis protein RLP24 and the large ribosomal subunit biogenesis protein JIP5 were included in the region between II-4 and II-5, both of which are homologous analogs that are essential in S. cerevisiae for biogenesis of the large ribosomal subunit. Therefore, we speculated that the lack of transformants might be due to gene loss preventing the growth of cells. Further studies may be needed to determine the deletion of larger fragments. These results show that our tool could be used to efficiently and rapidly edit genomes, accelerating metabolic network rearrangements and facilitating the construction of cell factories.
We first constructed yeast combinatorial libraries in K. phaffii for the fast balancing of metabolic pathways, which has been demonstrated previously in S. cerevisiae (EauClaire et al. 2016). Through the yeast combinatorial library, we successfully obtained different lycopene synthesis strains with yields up to 182.73 mg/L in shake flask fermentation. This result suggests that in constructing metabolic pathways, not all genes need to be under the control of a strong promoter. Instead, the metabolic pathway should be optimized by combining the demands of the metabolic flux and the catalytic efficiency (specific activity) of the enzyme itself. In previous reports, the ratio of multiple enzymes associated with product synthesis was determined by individually overexpressing each protein and measuring the resulting catalytic capacity in vitro, which in turn informed the optimization of the metabolic pathway in vivo (Liu et al. 2017). This is an effective strategy to improve product synthesis. However, in many cases, the catalytic capacity of each individual enzyme is not easily determined. For example, some enzymes may be difficult to heterologously express or purify, or their substrates/products may not be easily detected, which makes the implementation of this strategy difficult. Other researchers optimized the copy number of each gene to determine the rate-limiting step and enhance the yield of the product (Lv et al. 2019), which involves an extensive and time-consuming workload when the metabolic pathway involves a large number of genes. In contrast, yeast combinatorial libraries could effectively facilitate the regulation and assembly of metabolic pathways. Additionally, this strategy could be extended to the free assembly of other elements, such as different genes, terminators, and transcriptional regulators, making the construction of cell factories more efficient and more time-saving.
In conclusion, this work further expands the application of CRISPR/Cas9 in K. phaffii; provides newly developed rapid, efficient, and continuous genome editing tools; and establishes a flexible and concise genome editing method. In this research, marker recycling, multifragment integration, and large fragment knockout were optimized, and yeast combinatorial libraries were introduced for the first time. In summary, the CRISPR/Cas9 system described in this study provides a more efficient tool for the construction of K. phaffii cell factories.
Data availability
All data generated or analyzed during this study are included in this published article (and its supplementary information files).
References
Ahmad M, Hirz M, Pichler H, Schwab H (2014) Protein expression in Pichia pastoris: recent achievements and perspectives for heterologous protein production. Appl Microbiol Biotechnol 98(12):5301–5317. https://doi.org/10.1007/s00253-014-5732-5
Bernauer L, Radkohl A, Lehmayer LGK, Emmerstorfer-Augustin A (2021) Komagataella phaffii as emerging model organism in fundamental research. Front Microbiol 11:607028. https://doi.org/10.3389/fmicb.2020.607028
Cai P, Duan X, Wu X, Gao L, Ye M, Zhou YJ (2021) Recombination machinery engineering facilitates metabolic engineering of the industrial yeast Pichia pastoris. Nucleic Acids Res 49(13):7791–7805. https://doi.org/10.1093/nar/gkab535
Cai P, Wu X, Deng J, Gao L, Shen Y, Yao L, Zhou YJ (2022) Methanol biotransformation toward high-level production of fatty acid derivatives by engineering the industrial yeast Pichia pastoris. Proc Natl Acad Sci U S A 119(29):e2201711119. https://doi.org/10.1073/pnas.2201711119
Che Z, Cao X, Chen G, Liang Z (2020) An effective combination of codon optimization, gene dosage, and process optimization for high-level production of fibrinolytic enzyme in Komagataella phaffii (Pichia pastoris). BMC Biotechnol 20(1):63. https://doi.org/10.1186/s12896-020-00654-7
Dalvie NC, Leal J, Whittaker CA, Yang Y, Brady JR, Love KR, Love JC (2020) Host-informed expression of CRISPR guide RNA for genomic engineering in Komagataella phaffii. ACS Synth Biol 9(1):26–35. https://doi.org/10.1021/acssynbio.9b00372
EauClaire SF, Zhang J, Rivera CG, Huang LL (2016) Combinatorial metabolic pathway assembly in the yeast genome with RNA-guided Cas9. J Ind Microbiol Biotechnol 43(7):1001–1015. https://doi.org/10.1007/s10295-016-1776-0
Guo F, Dai Z, Peng W, Zhang S, Zhou J, Ma J, Dong W, **n F, Zhang W, Jiang M (2021) Metabolic engineering of Pichia pastoris for malic acid production from methanol. Biotechnol Bioeng 118(1):357–371. https://doi.org/10.1002/bit.27575
Ito Y, Watanabe T, Aikawa S, Nishi T, Nishiyama T, Nakamura Y, Hasunuma T, Okubo Y, Ishii J, Kondo A (2018) Deletion of DNA ligase IV homolog confers higher gene targeting efficiency on homologous recombination in Komagataella phaffii. FEMS Yeast Res 18(7):foy074. https://doi.org/10.1093/femsyr/foy074
Jakociunas T, Rajkumar AS, Zhang J, Arsovska D, Rodriguez A, Jendresen CB, Skjodt ML, Nielsen AT, Borodina I, Jensen MK, Keasling JD (2015) CasEMBLR: Cas9-facilitated multiloci genomic integration of in vivo assembled DNA parts in Saccharomyces cerevisiae. ACS Synth Biol 4(11):1226–1234. https://doi.org/10.1021/acssynbio.5b00007
** X, Zhang W, Wang Y, Sheng J, Xu R, Li J, Du G, Kang Z (2021) Biosynthesis of non-animal chondroitin sulfate from methanol using genetically engineered Pichia pastoris. Green Chem 23(12):4365–4374. https://doi.org/10.1039/d1gc00260k
Kong S, Yu W, Gao N, Zhai X, Zhou YJ (2022) Expanding the neutral sites for integrated gene expression in Saccharomyces cerevisiae. Fems Microbiol Lett 369(1):fnac081. https://doi.org/10.1093/femsle/fnac081
Kotaka A, Sahara H, Kondo A, Ueda M, Hata Y (2009) Efficient generation of recessive traits in diploid sake yeast by targeted gene disruption and loss of heterozygosity. Appl Microbiol Biotechnol 82(2):387–395. https://doi.org/10.1007/s00253-008-1833-3
Leprince A, van Passel MW, dos Santos VA (2012) Streamlining genomes: toward the generation of simplified and stabilized microbial systems. Curr Opin Biotechnol 23(5):651–658. https://doi.org/10.1016/j.copbio.2012.05.001
Li ZH, Liu M, Lyu XM, Wang FQ, Wei DZ (2018) CRISPR/Cpf1 facilitated large fragment deletion in Saccharomyces cerevisiae. J Basic Microbiol 58(12):1100–1104. https://doi.org/10.1002/jobm.201800195
Liao X, Li L, Jameel A, **ng XH, Zhang C (2021) A versatile toolbox for CRISPR-based genome engineering in Pichia pastoris. Appl Microbiol Biotechnol 105(24):9211–9218. https://doi.org/10.1007/s00253-021-11688-y
Lin Cereghino GP, Lin Cereghino J, Sunga AJ, Johnson MA, Lim M, Gleeson MA, Cregg JM (2001) New selectable marker/auxotrophic host strain combinations for molecular genetic manipulation of Pichia pastoris. Gene 263(1–2):159–169. https://doi.org/10.1016/S0378-1119(00)00576-X
Liu GS, Li T, Zhou W, Jiang M, Tao XY, Liu M, Zhao M, Ren YH, Gao B, Wang FQ, Wei DZ (2020) The yeast peroxisome: a dynamic storage depot and subcellular factory for squalene overproduction. Metab Eng 57:151–161. https://doi.org/10.1016/j.ymben.2019.11.001
Liu Q, Shi X, Song L, Liu H, Zhou X, Wang Q, Zhang Y, Cai M (2019) CRISPR-Cas9-mediated genomic multiloci integration in Pichia pastoris. Microb Cell Fact 18(1):144. https://doi.org/10.1186/s12934-019-1194-x
Liu Z, Zhang Y, Jia X, Hu M, Deng Z, Xu Y, Liu T (2017) In vitro reconstitution and optimization of the entire pathway to convert glucose into fatty acid. ACS Synth Biol 6(4):701–709. https://doi.org/10.1021/acssynbio.6b00348
Lv Y, Marsafari M, Koffas M, Zhou J, Xu P (2019) Optimizing oleaginous yeast cell factories for flavonoids and hydroxylated flavonoids biosynthesis. ACS Synth Biol 8(11):2514–2523. https://doi.org/10.1021/acssynbio.9b00193
Ma T, Shi B, Ye Z, Li X, Liu M, Chen Y, **a J, Nielsen J, Deng Z, Liu T (2019) Lipid engineering combined with systematic metabolic engineering of Saccharomyces cerevisiae for high-yield production of lycopene. Metab Eng 52:134–142. https://doi.org/10.1016/j.ymben.2018.11.009
Moon HY, Sim GH, Kim HJ, Kim K, Kang HA (2022) Assessment of Cre-lox and CRISPR-Cas9 as tools for recycling of multiple-integrated selection markers in Saccharomyces cerevisiae. J Microbiol 60(1):18–30. https://doi.org/10.1007/s12275-022-1580-7
Näätsaari L, Mistlberger B, Ruth C, Hajek T, Hartner FS, Glieder A (2012) Deletion of the Pichia pastoris KU70 homologue facilitates platform strain generation for gene expression and synthetic biology. PLoS ONE 7(6):e39720. https://doi.org/10.1371/journal.pone.0039720
Nakamura Y, Nishi T, Noguchi R, Ito Y, Watanabe T, Nishiyama T, Aikawa S, Hasunuma T, Ishii J, Okubo Y, Kondo A (2018) A stable, autonomously replicating plasmid vector containing Pichia pastoris centromeric DNA. Appl Environ Microbiol 84(15):e02882-e2917. https://doi.org/10.1128/AEM.02882-17
Nishi T, Ito Y, Nakamura Y, Yamaji T, Hashiba N, Tamai M, Yasohara Y, Ishii J, Kondo A (2022) One-step in vivo assembly of multiple DNA fragments and genomic integration in Komagataella phaffii. ACS Synth Biol 11(2):644–654. https://doi.org/10.1021/acssynbio.1c00302
Ohsawa S, Oku M, Yurimoto H, Sakai Y (2022) Regulation of peroxisome homeostasis by post-translational modification in the methylotrophic yeast Komagataella phaffii. Front Cell Dev Biol 10:887806. https://doi.org/10.3389/fcell.2022.887806
Otto M, Skrekas C, Gossing M, Gustafsson J, Siewers V, David F (2021) Expansion of the yeast modular cloning toolkit for CRISPR-based applications, genomic integrations and combinatorial libraries. ACS Synth Biol 10(12):3461–3474. https://doi.org/10.1021/acssynbio.1c00408
Schusterbauer V, Fischer JE, Gangl S, Schenzle L, Rinnofner C, Geier M, Sailer C, Glieder A, Thallinger GG (2022) Whole genome sequencing analysis of effects of CRISPR/Cas9 in Komagataella phaffii: a budding yeast in distress. J Fungi (basel) 8(10):992. https://doi.org/10.3390/jof8100992
Shen Q, Yu Z, Zhou XT, Zhang SJ, Zou SP, **ong N, Xue YP, Liu ZQ, Zheng YG (2021) Identification of a novel promoter for driving antibiotic-resistant genes to reduce the metabolic burden during protein expression and effectively select multiple integrations in Pichia Pastoris. Appl Microbiol Biotechnol 105(8):3211–3223. https://doi.org/10.1007/s00253-021-11195-0
Tan Z, Li J, Wu M, Tang C, Zhang H, Wang J (2011) High-level heterologous expression of an alkaline lipase gene from Penicillium cyclopium PG37 in Pichia pastoris. World J Microbiol Biotechnol 27(12):2767–2774. https://doi.org/10.1007/s11274-011-0752-0
Weninger A, Fischer JE, Raschmanova H, Kniely C, Vogl T, Glieder A (2018) Expanding the CRISPR/Cas9 toolkit for Pichia pastoris with efficient donor integration and alternative resistance markers. J Cell Biochem 119(4):3183–3198. https://doi.org/10.1002/jcb.26474
Weninger A, Glieder A, Vogl T (2015) A toolbox of endogenous and heterologous nuclear localization sequences for the methylotrophic yeast Pichia pastoris. Fems Yeast Res 15(7):fov082 https://doi.org/10.1093/femsyr/fov082
Weninger A, Hatzl A-M, Schmid C, Vogl T, Glieder A (2016) Combinatorial optimization of CRISPR/Cas9 expression enables precision genome engineering in the methylotrophic yeast Pichia pastoris. J Biotechnol 235:139–149. https://doi.org/10.1016/j.jbiotec.2016.03.027
Yang Y, Liu G, Chen X, Liu M, Zhan C, Liu X, Bai Z (2020) High efficiency CRISPR/Cas9 genome editing system with an eliminable episomal sgRNA plasmid in Pichia pastoris. Enzyme Microb Technol 138:109556. https://doi.org/10.1016/j.enzmictec.2020.109556
Yang Z, Zhang Z (2018) Engineering strategies for enhanced production of protein and bio-products in Pichia pastoris: a review. Biotechnol Adv 36(1):182–195. https://doi.org/10.1016/j.biotechadv.2017.11.002
Zhang X, Gu S, Zheng X, Peng S, Li Y, Lin Y, Liang S (2021) A novel and efficient genome editing tool assisted by CRISPR-Cas12a/Cpf1 for Pichia pastoris. ACS Synth Biol 10(11):2927–2937. https://doi.org/10.1021/acssynbio.1c00172
Zhang X, Wang D, Duan Y, Zheng X, Lin Y, Liang S (2020) Production of lycopene by metabolically engineered Pichia pastoris. Biosci Biotechnol Biochem 84(3):463–470 https://doi.org/10.1080/09168451.2019.1693250
Zhang Y, Wang J, Wang Z, Zhang Y, Shi S, Nielsen J, Liu Z (2019) A gRNA-tRNA array for CRISPR-Cas9 based rapid multiplexed genome editing in Saccharomyces cerevisiae. Nat Commun 10(1):1053. https://doi.org/10.1038/s41467-019-09005-3
Zuo Y, **ao F, Gao J, Ye C, Jiang L, Dong C, Lian J (2022) Establishing Komagataella phaffii as a cell factory for efficient production of sesquiterpenoid alpha-santalene. J Agric Food Chem 70(26):8024–8031. https://doi.org/10.1021/acs.jafc.2c02353
Funding
This work was supported by the National Key Research and Development Program of China (No. 2023YFA0914100), and the Natural Science Foundation of Shanghai (No. 21ZR1416500).
Author information
Authors and Affiliations
Contributions
W.Z., F.W., and B.G. conceived and designed the research. W.Z., Y.L., and W.Q. conducted experiments. D.W. provided technical assistance. W.Z. and G.L. analyzed the data. W.Z. wrote the manuscript. All authors read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethical approval
This study does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, W., Li, Y., Liu, G. et al. CRISPR/Cas9-based toolkit for rapid marker recycling and combinatorial libraries in Komagataella phaffii. Appl Microbiol Biotechnol 108, 197 (2024). https://doi.org/10.1007/s00253-024-13037-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00253-024-13037-1